Francisco Galuppo Azevedo
Real-Time Explanations for Tabular Foundation Models
Mar 31, 2026Abstract:Interpretability is central for scientific machine learning, as understanding \emph{why} models make predictions enables hypothesis generation and validation. While tabular foundation models show strong performance, existing explanation methods like SHAP are computationally expensive, limiting interactive exploration. We introduce ShapPFN, a foundation model that integrates Shapley value regression directly into its architecture, producing both predictions and explanations in a single forward pass. On standard benchmarks, ShapPFN achieves competitive performance while producing high-fidelity explanations ($R^2$=0.96, cosine=0.99) over 1000\times faster than KernelSHAP (0.06s vs 610s). Our code is available at https://github.com/kunumi/ShapPFN
Task Scarcity and Label Leakage in Relational Transfer Learning
Mar 31, 2026Abstract:Training relational foundation models requires learning representations that transfer across tasks, yet available supervision is typically limited to a small number of prediction targets per database. This task scarcity causes learned representations to encode task-specific shortcuts that degrade transfer even within the same schema, a problem we call label leakage. We study this using K-Space, a modular architecture combining frozen pretrained tabular encoders with a lightweight message-passing core. To suppress leakage, we introduce a gradient projection method that removes label-predictive directions from representation updates. On RelBench, this improves within-dataset transfer by +0.145 AUROC on average, often recovering near single-task performance. Our results suggest that limited task diversity, not just limited data, constrains relational foundation models.
Evaluating the state-of-the-art in mapping research spaces: a Brazilian case study
Apr 07, 2021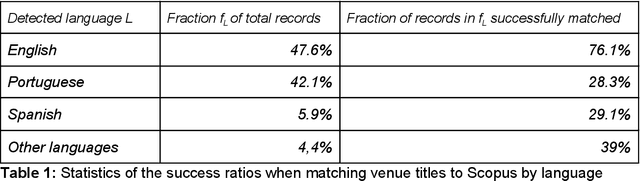
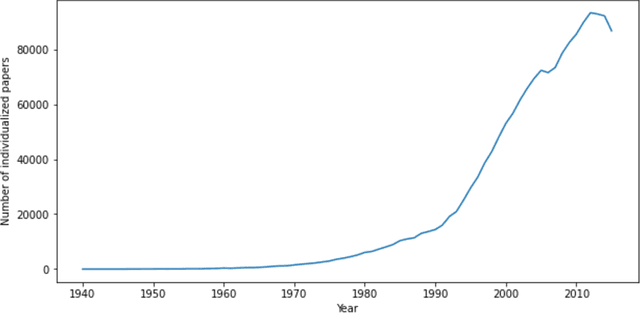

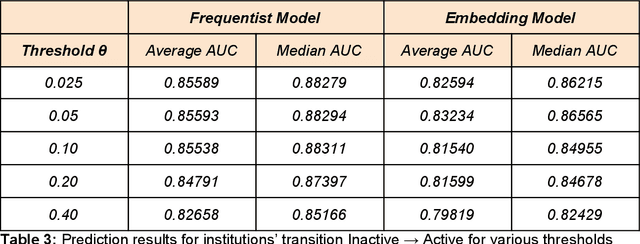
Abstract:Scientific knowledge cannot be seen as a set of isolated fields, but as a highly connected network. Understanding how research areas are connected is of paramount importance for adequately allocating funding and human resources (e.g., assembling teams to tackle multidisciplinary problems). The relationship between disciplines can be drawn from data on the trajectory of individual scientists, as researchers often make contributions in a small set of interrelated areas. Two recent works propose methods for creating research maps from scientists' publication records: by using a frequentist approach to create a transition probability matrix; and by learning embeddings (vector representations). Surprisingly, these models were evaluated on different datasets and have never been compared in the literature. In this work, we compare both models in a systematic way, using a large dataset of publication records from Brazilian researchers. We evaluate these models' ability to predict whether a given entity (scientist, institution or region) will enter a new field w.r.t. the area under the ROC curve. Moreover, we analyze how sensitive each method is to the number of publications and the number of fields associated to one entity. Last, we conduct a case study to showcase how these models can be used to characterize science dynamics in the context of Brazil.
* 28 pages, 11 figures
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge