Francesco Saverio Pavone
Semantic Segmentation of Neuronal Bodies in Fluorescence Microscopy Using a 2D+3D CNN Training Strategy with Sparsely Annotated Data
Sep 02, 2020
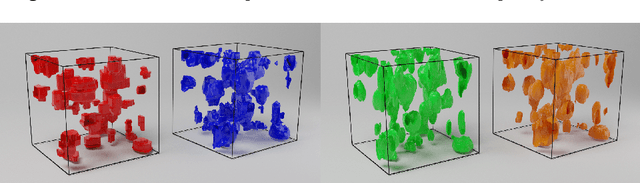
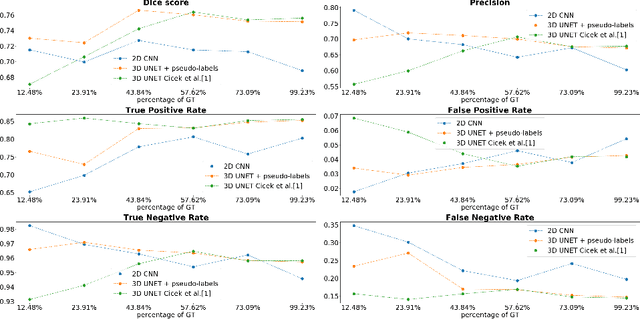

Abstract:Semantic segmentation of neuronal structures in 3D high-resolution fluorescence microscopy imaging of the human brain cortex can take advantage of bidimensional CNNs, which yield good results in neuron localization but lead to inaccurate surface reconstruction. 3D CNNs, on the other hand, would require manually annotated volumetric data on a large scale and hence considerable human effort. Semi-supervised alternative strategies which make use only of sparse annotations suffer from longer training times and achieved models tend to have increased capacity compared to 2D CNNs, needing more ground truth data to attain similar results. To overcome these issues we propose a two-phase strategy for training native 3D CNN models on sparse 2D annotations where missing labels are inferred by a 2D CNN model and combined with manual annotations in a weighted manner during loss calculation.
Cell identification in whole-brain multiview images of neural activation
Nov 04, 2015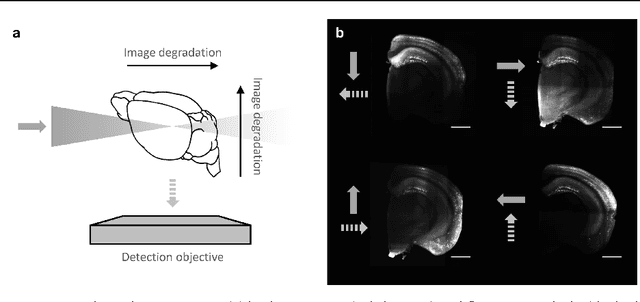

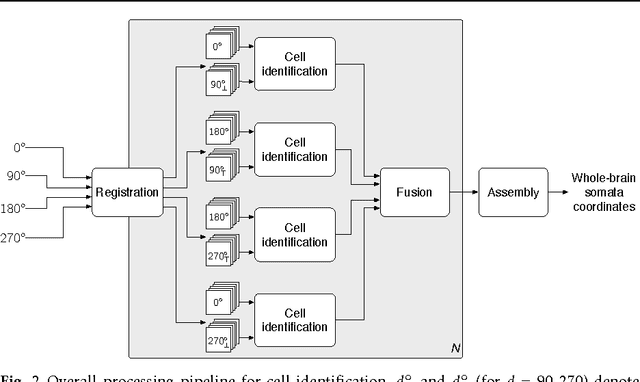
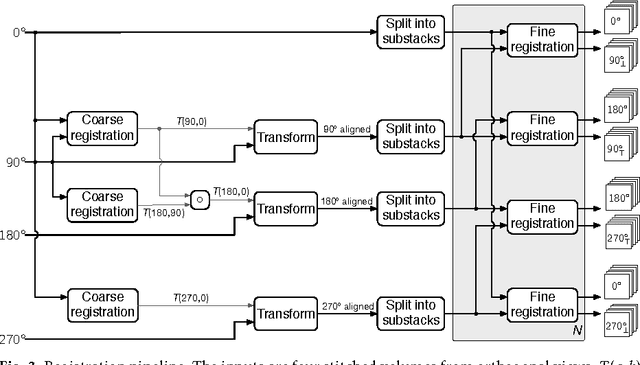
Abstract:We present a scalable method for brain cell identification in multiview confocal light sheet microscopy images. Our algorithmic pipeline includes a hierarchical registration approach and a novel multiview version of semantic deconvolution that simultaneously enhance visibility of fluorescent cell bodies, equalize their contrast, and fuses adjacent views into a single 3D images on which cell identification is performed with mean shift. We present empirical results on a whole-brain image of an adult Arc-dVenus mouse acquired at 4micron resolution. Based on an annotated test volume containing 3278 cells, our algorithm achieves an $F_1$ measure of 0.89.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge