Fatemeh Bagheri
EOG Artifact Removal from Single and Multi-channel EEG Recordings through the combination of Long Short-Term Memory Networks and Independent Component Analysis
Aug 25, 2023



Abstract:Introduction: Electroencephalogram (EEG) signals have gained significant popularity in various applications due to their rich information content. However, these signals are prone to contamination from various sources of artifacts, notably the electrooculogram (EOG) artifacts caused by eye movements. The most effective approach to mitigate EOG artifacts involves recording EOG signals simultaneously with EEG and employing blind source separation techniques, such as independent component analysis (ICA). Nevertheless, the availability of EOG recordings is not always feasible, particularly in pre-recorded datasets. Objective: In this paper, we present a novel methodology that combines a long short-term memory (LSTM)-based neural network with ICA to address the challenge of EOG artifact removal from contaminated EEG signals. Approach: Our approach aims to accomplish two primary objectives: 1) estimate the horizontal and vertical EOG signals from the contaminated EEG data, and 2) employ ICA to eliminate the estimated EOG signals from the EEG, thereby producing an artifact-free EEG signal. Main results: To evaluate the performance of our proposed method, we conducted experiments on a publicly available dataset comprising recordings from 27 participants. We employed well-established metrics such as mean squared error, mean absolute error, and mean error to assess the quality of our artifact removal technique. Significance: Furthermore, we compared the performance of our approach with two state-of-the-art deep learning-based methods reported in the literature, demonstrating the superior performance of our proposed methodology.
Fast fully-reproducible serial/parallel Monte Carlo and MCMC simulations and visualizations via ParaMonte::Python library
Oct 01, 2020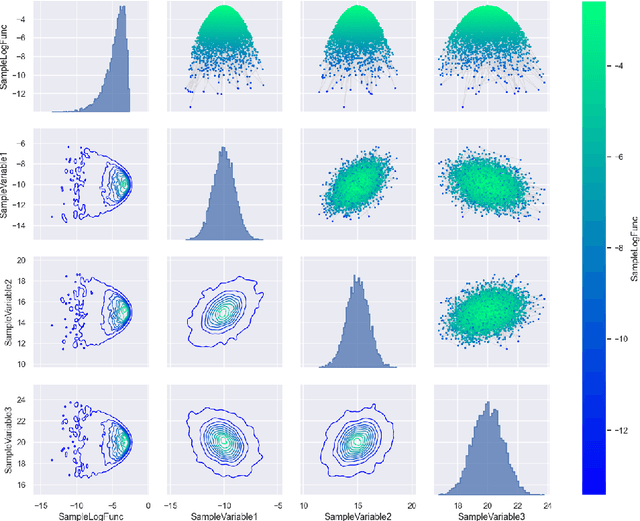
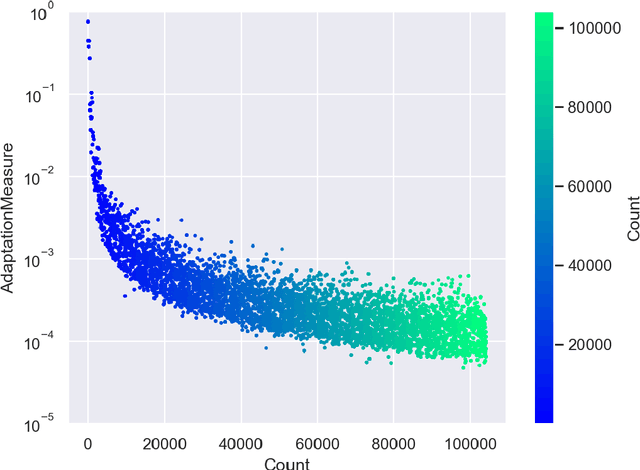
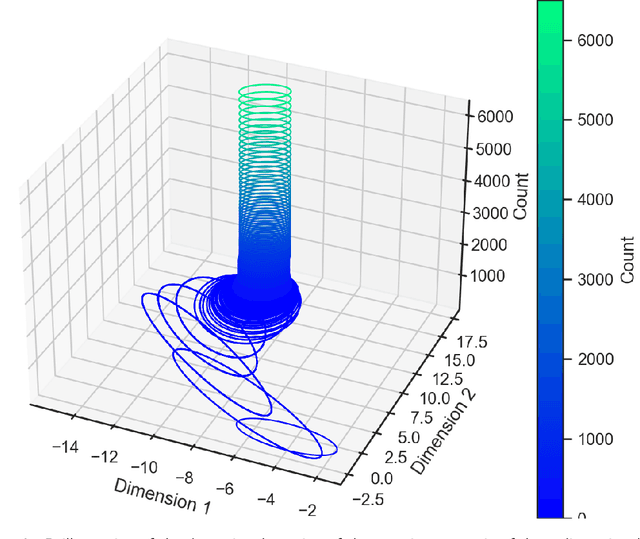
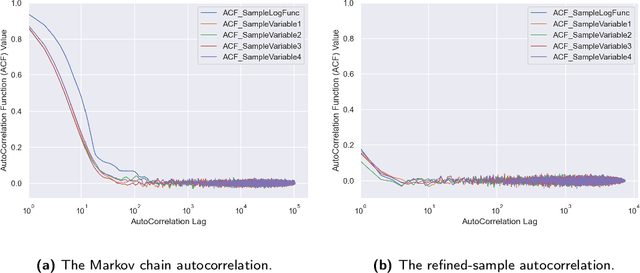
Abstract:ParaMonte::Python (standing for Parallel Monte Carlo in Python) is a serial and MPI-parallelized library of (Markov Chain) Monte Carlo (MCMC) routines for sampling mathematical objective functions, in particular, the posterior distributions of parameters in Bayesian modeling and analysis in data science, Machine Learning, and scientific inference in general. In addition to providing access to fast high-performance serial/parallel Monte Carlo and MCMC sampling routines, the ParaMonte::Python library provides extensive post-processing and visualization tools that aim to automate and streamline the process of model calibration and uncertainty quantification in Bayesian data analysis. Furthermore, the automatically-enabled restart functionality of ParaMonte::Python samplers ensure seamless fully-deterministic into-the-future restart of Monte Carlo simulations, should any interruptions happen. The ParaMonte::Python library is MIT-licensed and is permanently maintained on GitHub at https://github.com/cdslaborg/paramonte/tree/master/src/interface/Python.
ParaMonte: A high-performance serial/parallel Monte Carlo simulation library for C, C++, Fortran
Sep 29, 2020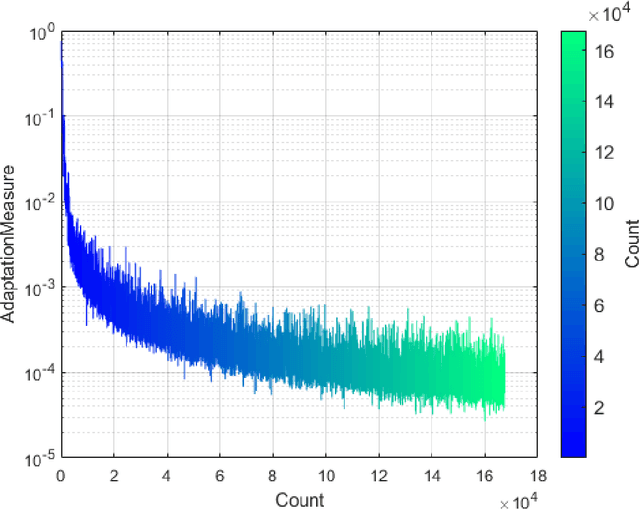
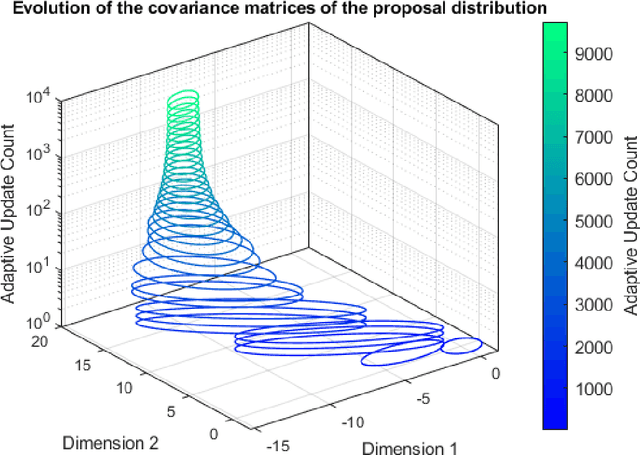
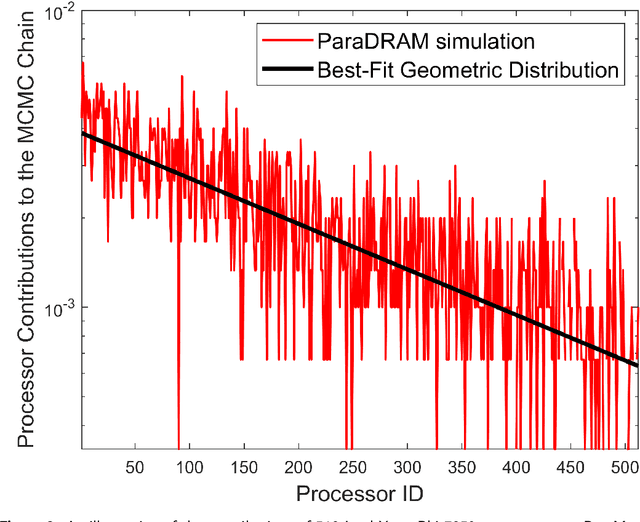
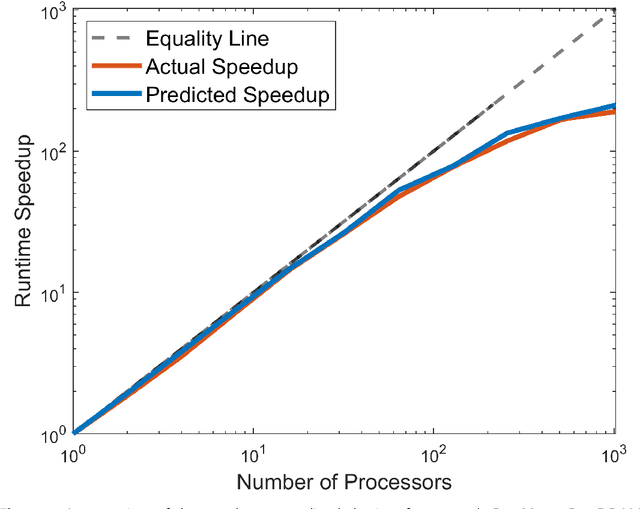
Abstract:ParaMonte (standing for Parallel Monte Carlo) is a serial and MPI/Coarray-parallelized library of Monte Carlo routines for sampling mathematical objective functions of arbitrary-dimensions, in particular, the posterior distributions of Bayesian models in data science, Machine Learning, and scientific inference. The ParaMonte library has been developed with the design goal of unifying the **automation**, **accessibility**, **high-performance**, **scalability**, and **reproducibility** of Monte Carlo simulations. The current implementation of the library includes **ParaDRAM**, a **Para**llel **D**elyaed-**R**ejection **A**daptive **M**etropolis Markov Chain Monte Carlo sampler, accessible from a wide range of programming languages including C, C++, Fortran, with a unified Application Programming Interface and simulation environment across all supported programming languages. The ParaMonte library is MIT-licensed and is permanently located and maintained at [https://github.com/cdslaborg/paramonte](https://github.com/cdslaborg/paramonte).
ParaDRAM: A Cross-Language Toolbox for Parallel High-Performance Delayed-Rejection Adaptive Metropolis Markov Chain Monte Carlo Simulations
Aug 21, 2020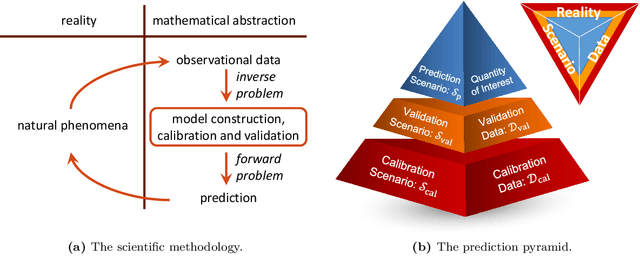
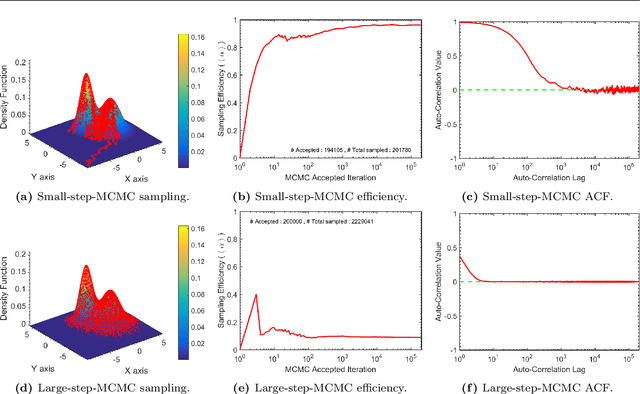
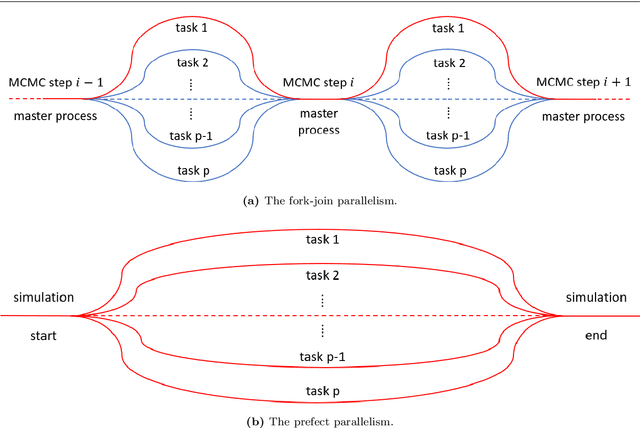
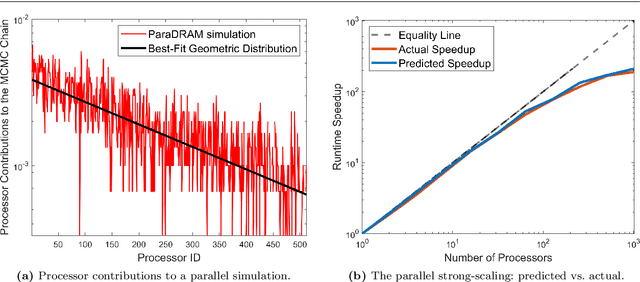
Abstract:We present ParaDRAM, a high-performance Parallel Delayed-Rejection Adaptive Metropolis Markov Chain Monte Carlo software for optimization, sampling, and integration of mathematical objective functions encountered in scientific inference. ParaDRAM is currently accessible from several popular programming languages including C/C++, Fortran, MATLAB, Python and is part of the ParaMonte open-source project with the following principal design goals: 1. full automation of Monte Carlo simulations, 2. interoperability of the core library with as many programming languages as possible, thus, providing a unified Application Programming Interface and Monte Carlo simulation environment across all programming languages, 3. high-performance 4. parallelizability and scalability of simulations from personal laptops to supercomputers, 5. virtually zero-dependence on external libraries, 6. fully-deterministic reproducibility of simulations, 7. automatic comprehensive reporting and post-processing of the simulation results. We present and discuss several novel techniques implemented in ParaDRAM to automatically and dynamically ensure the good-mixing and the diminishing-adaptation of the resulting pseudo-Markov chains from ParaDRAM. We also discuss the implementation of an efficient data storage method used in ParaDRAM that reduces the average memory and storage requirements of the algorithm by, a factor of 4 for simple simulation problems, to an order of magnitude and more for sampling complex high-dimensional mathematical objective functions. Finally, we discuss how the design goals of ParaDRAM can help users readily and efficiently solve a variety of machine learning and scientific inference problems on a wide range of computing platforms.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge