Eric de Sturler
ResGene-T: A Tensor-Based Residual Network Approach for Genomic Prediction
Feb 28, 2026Abstract:In this work, we propose a new deep learning model for Genomic Prediction (GP), which involves correlating genotypic data with phenotypic. The genotypes are typically fed as a sequence of characters to the 1D-Convolution Neural Network layer of the underlying deep learning model. Inspired by earlier work that represented genotype as a 2D-image for genotype-phenotype classification, we extend this idea to GP, which is a regression task. We use a ResNet-18 as the underlying architecture, and term this model as ResGene-2D. Although the 2D-image representation captures biological interactions well, it requires all the layers of the model to do so. This limits training efficiency. Thus, as seen in the earlier work that proposed a 2D-image representation, our ResGene-2D performs almost the same as other models (3% improvement). To overcome this, we propose a novel idea of converting the 2D-image into a 3D/ tensor and feed this to the ResNet-18 architecture, and term this model as ResGene-T. We evaluate our proposed models on three crop species having ten phenotypic traits and compare it with seven most popular models (two statistical, two machine learning, and three deep learning). ResGene-T performs the best among all these seven methods (gains from 14.51% to 41.51%).
Parametric Level-sets Enhanced To Improve Reconstruction (PaLEnTIR)
Apr 21, 2022
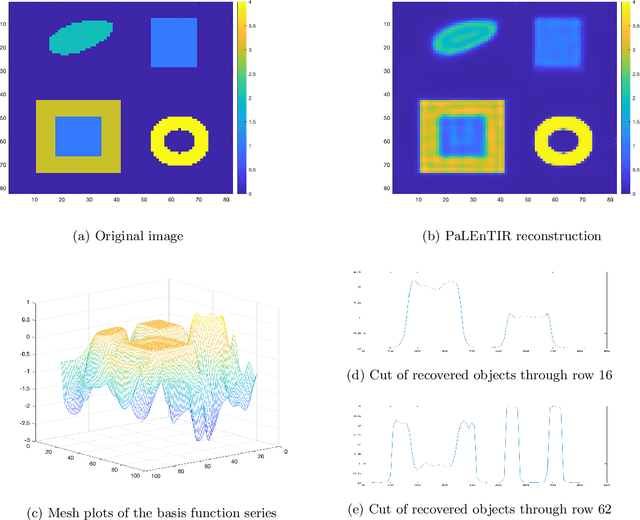
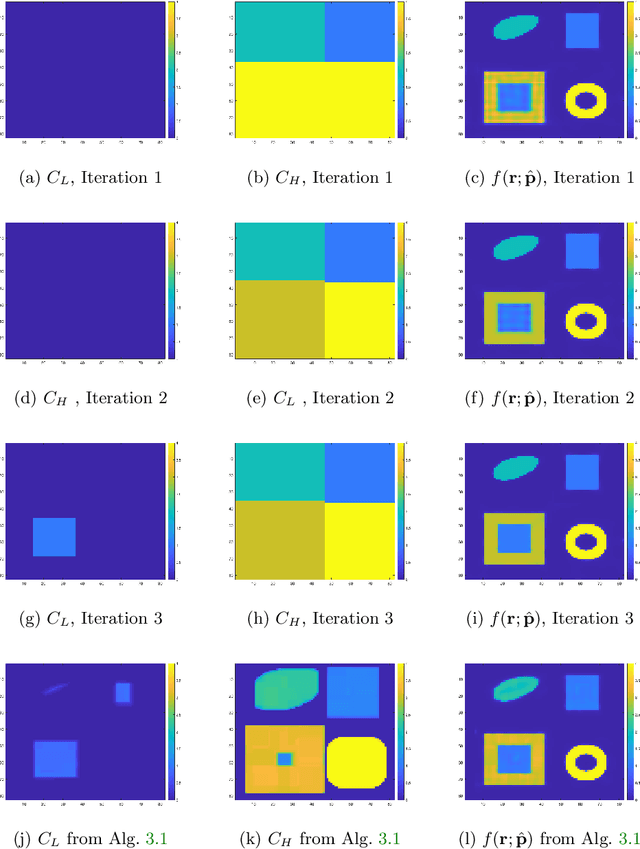
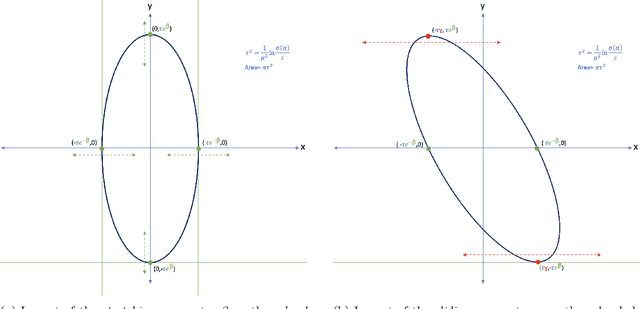
Abstract:In this paper, we consider the restoration and reconstruction of piecewise constant objects in two and three dimensions using PaLEnTIR, a significantly enhanced Parametric level set (PaLS) model relative to the current state-of-the-art. The primary contribution of this paper is a new PaLS formulation which requires only a single level set function to recover a scene with piecewise constant objects possessing multiple unknown contrasts. Our model offers distinct advantages over current approaches to the multi-contrast, multi-object problem, all of which require multiple level sets and explicit estimation of the contrast magnitudes. Given upper and lower bounds on the contrast, our approach is able to recover objects with any distribution of contrasts and eliminates the need to know either the number of contrasts in a given scene or their values. We provide an iterative process for finding these space-varying contrast limits. Relative to most PaLS methods which employ radial basis functions (RBFs), our model makes use of non-isotropic basis functions, thereby expanding the class of shapes that a PaLS model of a given complexity can approximate. Finally, PaLEnTIR improves the conditioning of the Jacobian matrix required as part of the parameter identification process and consequently accelerates the optimization methods by controlling the magnitude of the PaLS expansion coefficients, fixing the centers of the basis functions, and the uniqueness of parametric to image mappings provided by the new parameterization. We demonstrate the performance of the new approach using both 2D and 3D variants of X-ray computed tomography, diffuse optical tomography (DOT), denoising, deconvolution problems. Application to experimental sparse CT data and simulated data with different types of noise are performed to further validate the proposed method.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge