Eitam Sheetrit
Knowledge-Instruct: Effective Continual Pre-training from Limited Data using Instructions
Apr 08, 2025



Abstract:While Large Language Models (LLMs) acquire vast knowledge during pre-training, they often lack domain-specific, new, or niche information. Continual pre-training (CPT) attempts to address this gap but suffers from catastrophic forgetting and inefficiencies in low-data regimes. We introduce Knowledge-Instruct, a novel approach to efficiently inject knowledge from limited corpora through pure instruction-tuning. By generating information-dense synthetic instruction data, it effectively integrates new knowledge while preserving general reasoning and instruction-following abilities. Knowledge-Instruct demonstrates superior factual memorization, minimizes catastrophic forgetting, and remains scalable by leveraging synthetic data from relatively small language models. Additionally, it enhances contextual understanding, including complex multi-hop reasoning, facilitating integration with retrieval systems. We validate its effectiveness across diverse benchmarks, including Companies, a new dataset that we release to measure knowledge injection capabilities.
SECQUE: A Benchmark for Evaluating Real-World Financial Analysis Capabilities
Apr 06, 2025



Abstract:We introduce SECQUE, a comprehensive benchmark for evaluating large language models (LLMs) in financial analysis tasks. SECQUE comprises 565 expert-written questions covering SEC filings analysis across four key categories: comparison analysis, ratio calculation, risk assessment, and financial insight generation. To assess model performance, we develop SECQUE-Judge, an evaluation mechanism leveraging multiple LLM-based judges, which demonstrates strong alignment with human evaluations. Additionally, we provide an extensive analysis of various models' performance on our benchmark. By making SECQUE publicly available, we aim to facilitate further research and advancements in financial AI.
Mixing It Up: The Cocktail Effect of Multi-Task Fine-Tuning on LLM Performance -- A Case Study in Finance
Oct 01, 2024



Abstract:The application of large language models (LLMs) in domain-specific contexts, including finance, has expanded rapidly. Domain-specific LLMs are typically evaluated based on their performance in various downstream tasks relevant to the domain. In this work, we present a detailed analysis of fine-tuning LLMs for such tasks. Somewhat counterintuitively, we find that in domain-specific cases, fine-tuning exclusively on the target task is not always the most effective strategy. Instead, multi-task fine-tuning - where models are trained on a cocktail of related tasks - can significantly enhance performance. We demonstrate how this approach enables a small model, such as Phi-3-Mini, to achieve state-of-the-art results, even surpassing the much larger GPT-4-o model on financial benchmarks. Our study involves a large-scale experiment, training over 200 models using several widely adopted LLMs as baselines, and empirically confirms the benefits of multi-task fine-tuning. Additionally, we explore the use of general instruction data as a form of regularization, suggesting that it helps minimize performance degradation. We also investigate the inclusion of mathematical data, finding improvements in numerical reasoning that transfer effectively to financial tasks. Finally, we note that while fine-tuning for downstream tasks leads to targeted improvements in task performance, it does not necessarily result in broader gains in domain knowledge or complex domain reasoning abilities.
ReMatch: Retrieval Enhanced Schema Matching with LLMs
Mar 03, 2024Abstract:Schema matching is a crucial task in data integration, involving the alignment of a source database schema with a target schema to establish correspondence between their elements. This task is challenging due to textual and semantic heterogeneity, as well as differences in schema sizes. Although machine-learning-based solutions have been explored in numerous studies, they often suffer from low accuracy, require manual mapping of the schemas for model training, or need access to source schema data which might be unavailable due to privacy concerns. In this paper we present a novel method, named ReMatch, for matching schemas using retrieval-enhanced Large Language Models (LLMs). Our method avoids the need for predefined mapping, any model training, or access to data in the source database. In the ReMatch method the tables of the target schema and the attributes of the source schema are first represented as structured passage-based documents. For each source attribute document, we retrieve $J$ documents, representing target schema tables, according to their semantic relevance. Subsequently, we create a prompt for every source table, comprising all its attributes and their descriptions, alongside all attributes from the set of top $J$ target tables retrieved previously. We employ LLMs using this prompt for the matching task, yielding a ranked list of $K$ potential matches for each source attribute. Our experimental results on large real-world schemas demonstrate that ReMatch significantly improves matching capabilities and outperforms other machine learning approaches. By eliminating the requirement for training data, ReMatch becomes a viable solution for real-world scenarios.
Predicting Unplanned Readmissions in the Intensive Care Unit: A Multimodality Evaluation
May 14, 2023Abstract:A hospital readmission is when a patient who was discharged from the hospital is admitted again for the same or related care within a certain period. Hospital readmissions are a significant problem in the healthcare domain, as they lead to increased hospitalization costs, decreased patient satisfaction, and increased risk of adverse outcomes such as infections, medication errors, and even death. The problem of hospital readmissions is particularly acute in intensive care units (ICUs), due to the severity of the patients' conditions, and the substantial risk of complications. Predicting Unplanned Readmissions in ICUs is a challenging task, as it involves analyzing different data modalities, such as static data, unstructured free text, sequences of diagnoses and procedures, and multivariate time-series. Here, we investigate the effectiveness of each data modality separately, then alongside with others, using state-of-the-art machine learning approaches in time-series analysis and natural language processing. Using our evaluation process, we are able to determine the contribution of each data modality, and for the first time in the context of readmission, establish a hierarchy of their predictive value. Additionally, we demonstrate the impact of Temporal Abstractions in enhancing the performance of time-series approaches to readmission prediction. Due to conflicting definitions in the literature, we also provide a clear definition of the term Unplanned Readmission to enhance reproducibility and consistency of future research and to prevent any potential misunderstandings that could result from diverse interpretations of the term. Our experimental results on a large benchmark clinical data set show that Discharge Notes written by physicians, have better capabilities for readmission prediction than all other modalities.
Temporal Pattern Discovery for Accurate Sepsis Diagnosis in ICU Patients
Sep 06, 2017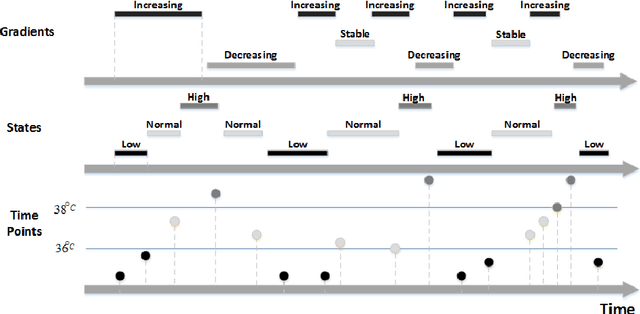
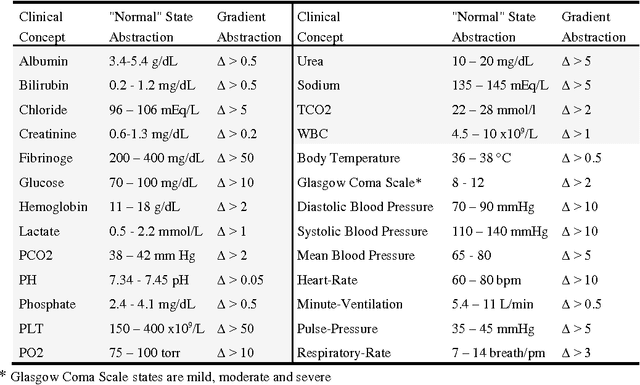


Abstract:Sepsis is a condition caused by the body's overwhelming and life-threatening response to infection, which can lead to tissue damage, organ failure, and finally death. Common signs and symptoms include fever, increased heart rate, increased breathing rate, and confusion. Sepsis is difficult to predict, diagnose, and treat. Patients who develop sepsis have an increased risk of complications and death and face higher health care costs and longer hospitalization. Today, sepsis is one of the leading causes of mortality among populations in intensive care units (ICUs). In this paper, we look at the problem of early detection of sepsis by using temporal data mining. We focus on the use of knowledge-based temporal abstraction to create meaningful interval-based abstractions, and on time-interval mining to discover frequent interval-based patterns. We used 2,560 cases derived from the MIMIC-III database. We found that the distribution of the temporal patterns whose frequency is above 10% discovered in the records of septic patients during the last 6 and 12 hours before onset of sepsis is significantly different from that distribution within a similar period, during an equivalent time window during hospitalization, in the records of non-septic patients. This discovery is encouraging for the purpose of performing an early diagnosis of sepsis using the discovered patterns as constructed features.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge