Dr Jeff Dalton
Consensus of state of the art mortality prediction models: From all-cause mortality to sudden death prediction
Aug 30, 2023Abstract:Worldwide, many millions of people die suddenly and unexpectedly each year, either with or without a prior history of cardiovascular disease. Such events are sparse (once in a lifetime), many victims will not have had prior investigations for cardiac disease and many different definitions of sudden death exist. Accordingly, sudden death is hard to predict. This analysis used NHS Electronic Health Records (EHRs) for people aged $\geq$50 years living in the Greater Glasgow and Clyde (GG\&C) region in 2010 (n = 380,000) to try to overcome these challenges. We investigated whether medical history, blood tests, prescription of medicines, and hospitalisations might, in combination, predict a heightened risk of sudden death. We compared the performance of models trained to predict either sudden death or all-cause mortality. We built six models for each outcome of interest: three taken from state-of-the-art research (BEHRT, Deepr and Deep Patient), and three of our own creation. We trained these using two different data representations: a language-based representation, and a sparse temporal matrix. We used global interpretability to understand the most important features of each model, and compare how much agreement there was amongst models using Rank Biased Overlap. It is challenging to account for correlated variables without increasing the complexity of the interpretability technique. We overcame this by clustering features into groups and comparing the most important groups for each model. We found the agreement between models to be much higher when accounting for correlated variables. Our analysis emphasises the challenge of predicting sudden death and emphasises the need for better understanding and interpretation of machine learning models applied to healthcare applications.
Improving ECG Classification Interpretability using Saliency Maps
Jan 10, 2022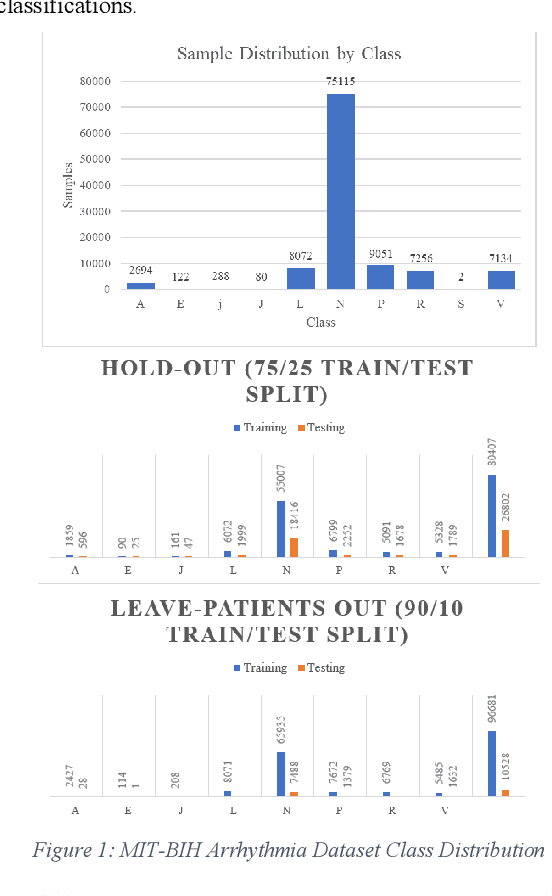
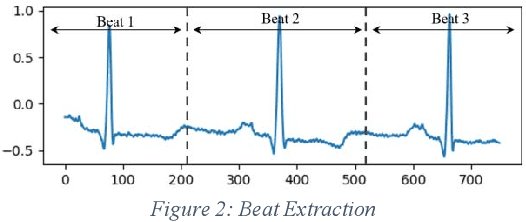
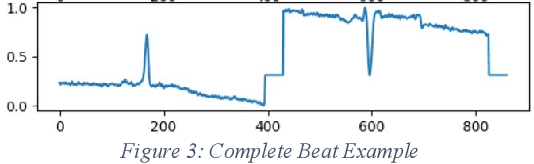
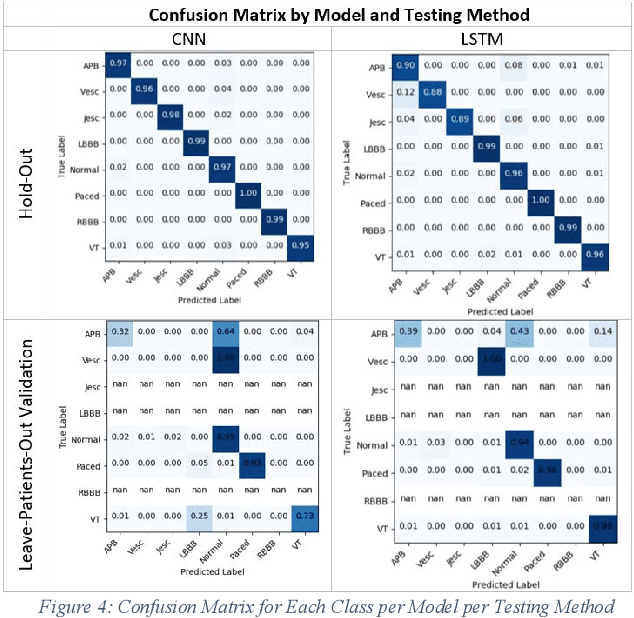
Abstract:Cardiovascular disease is a large worldwide healthcare issue; symptoms often present suddenly with minimal warning. The electrocardiogram (ECG) is a fast, simple and reliable method of evaluating the health of the heart, by measuring electrical activity recorded through electrodes placed on the skin. ECGs often need to be analyzed by a cardiologist, taking time which could be spent on improving patient care and outcomes. Because of this, automatic ECG classification systems using machine learning have been proposed, which can learn complex interactions between ECG features and use this to detect abnormalities. However, algorithms built for this purpose often fail to generalize well to unseen data, reporting initially impressive results which drop dramatically when applied to new environments. Additionally, machine learning algorithms suffer a "black-box" issue, in which it is difficult to determine how a decision has been made. This is vital for applications in healthcare, as clinicians need to be able to verify the process of evaluation in order to trust the algorithm. This paper proposes a method for visualizing model decisions across each class in the MIT-BIH arrhythmia dataset, using adapted saliency maps averaged across complete classes to determine what patterns are being learned. We do this by building two algorithms based on state-of-the-art models. This paper highlights how these maps can be used to find problems in the model which could be affecting generalizability and model performance. Comparing saliency maps across complete classes gives an overall impression of confounding variables or other biases in the model, unlike what would be highlighted when comparing saliency maps on an ECG-by-ECG basis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge