Representation Power of Graph Convolutions : Neural Tangent Kernel Analysis
Oct 18, 2022Mahalakshmi Sabanayagam, Pascal Esser, Debarghya Ghoshdastidar
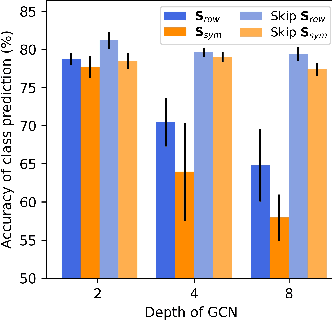
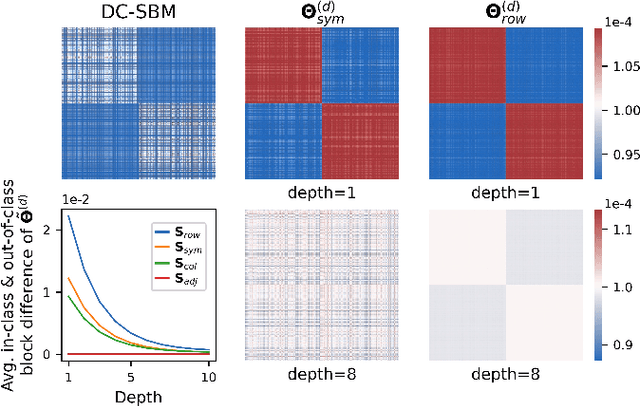
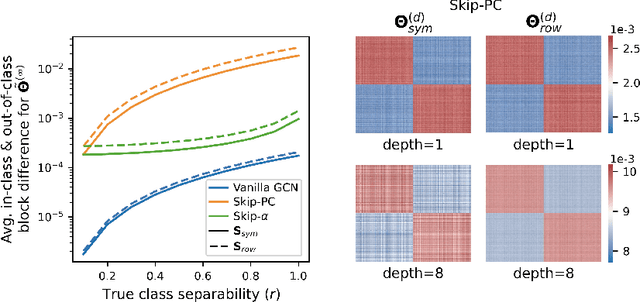
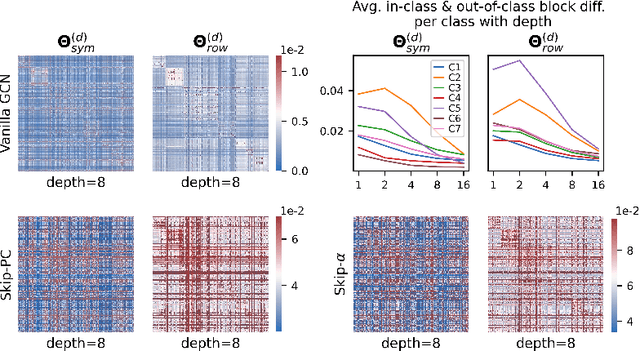
The fundamental principle of Graph Neural Networks (GNNs) is to exploit the structural information of the data by aggregating the neighboring nodes using a graph convolution. Therefore, understanding its influence on the network performance is crucial. Convolutions based on graph Laplacian have emerged as the dominant choice with the symmetric normalization of the adjacency matrix $A$, defined as $D^{-1/2}AD^{-1/2}$, being the most widely adopted one, where $D$ is the degree matrix. However, some empirical studies show that row normalization $D^{-1}A$ outperforms it in node classification. Despite the widespread use of GNNs, there is no rigorous theoretical study on the representation power of these convolution operators, that could explain this behavior. In this work, we analyze the influence of the graph convolutions theoretically using Graph Neural Tangent Kernel in a semi-supervised node classification setting. Under a Degree Corrected Stochastic Block Model, we prove that: (i) row normalization preserves the underlying class structure better than other convolutions; (ii) performance degrades with network depth due to over-smoothing, but the loss in class information is the slowest in row normalization; (iii) skip connections retain the class information even at infinite depth, thereby eliminating over-smoothing. We finally validate our theoretical findings on real datasets.
Interpolation and Regularization for Causal Learning
Feb 18, 2022Leena Chennuru Vankadara, Luca Rendsburg, Ulrike von Luxburg, Debarghya Ghoshdastidar
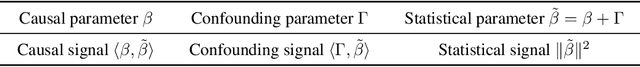
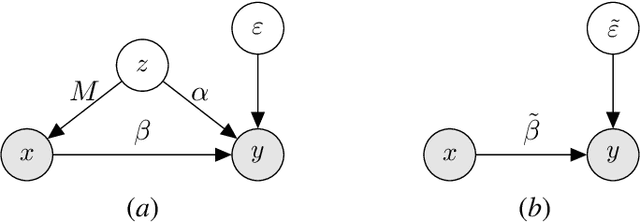

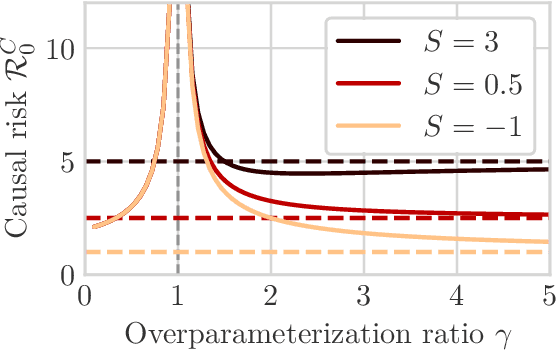
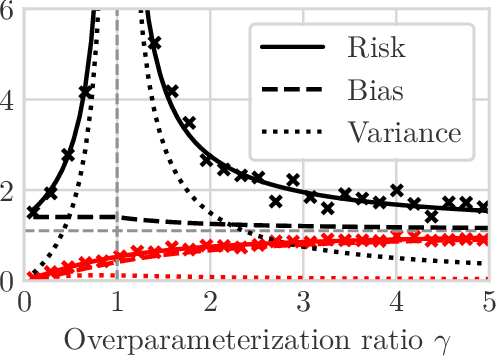
We study the problem of learning causal models from observational data through the lens of interpolation and its counterpart -- regularization. A large volume of recent theoretical, as well as empirical work, suggests that, in highly complex model classes, interpolating estimators can have good statistical generalization properties and can even be optimal for statistical learning. Motivated by an analogy between statistical and causal learning recently highlighted by Janzing (2019), we investigate whether interpolating estimators can also learn good causal models. To this end, we consider a simple linearly confounded model and derive precise asymptotics for the *causal risk* of the min-norm interpolator and ridge-regularized regressors in the high-dimensional regime. Under the principle of independent causal mechanisms, a standard assumption in causal learning, we find that interpolators cannot be optimal and causal learning requires stronger regularization than statistical learning. This resolves a recent conjecture in Janzing (2019). Beyond this assumption, we find a larger range of behavior that can be precisely characterized with a new measure of *confounding strength*. If the confounding strength is negative, causal learning requires weaker regularization than statistical learning, interpolators can be optimal, and the optimal regularization can even be negative. If the confounding strength is large, the optimal regularization is infinite, and learning from observational data is actively harmful.
Learning Theory Can (Sometimes) Explain Generalisation in Graph Neural Networks
Dec 07, 2021Pascal Mattia Esser, Leena Chennuru Vankadara, Debarghya Ghoshdastidar
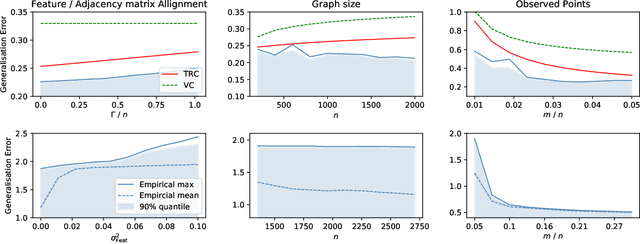
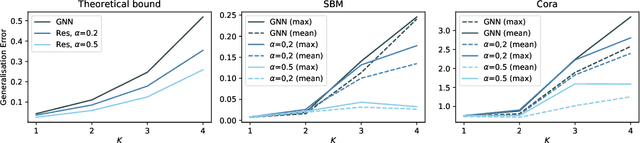
In recent years, several results in the supervised learning setting suggested that classical statistical learning-theoretic measures, such as VC dimension, do not adequately explain the performance of deep learning models which prompted a slew of work in the infinite-width and iteration regimes. However, there is little theoretical explanation for the success of neural networks beyond the supervised setting. In this paper we argue that, under some distributional assumptions, classical learning-theoretic measures can sufficiently explain generalization for graph neural networks in the transductive setting. In particular, we provide a rigorous analysis of the performance of neural networks in the context of transductive inference, specifically by analysing the generalisation properties of graph convolutional networks for the problem of node classification. While VC Dimension does result in trivial generalisation error bounds in this setting as well, we show that transductive Rademacher complexity can explain the generalisation properties of graph convolutional networks for stochastic block models. We further use the generalisation error bounds based on transductive Rademacher complexity to demonstrate the role of graph convolutions and network architectures in achieving smaller generalisation error and provide insights into when the graph structure can help in learning. The findings of this paper could re-new the interest in studying generalisation in neural networks in terms of learning-theoretic measures, albeit in specific problems.
Causal Forecasting:Generalization Bounds for Autoregressive Models
Nov 18, 2021Leena Chennuru Vankadara, Philipp Michael Faller, Lenon Minorics, Debarghya Ghoshdastidar, Dominik Janzing
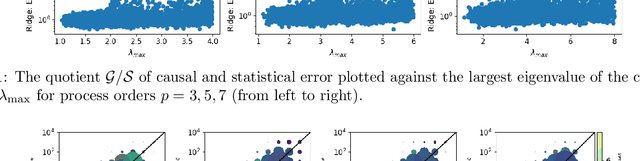
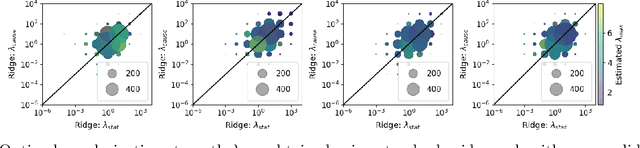
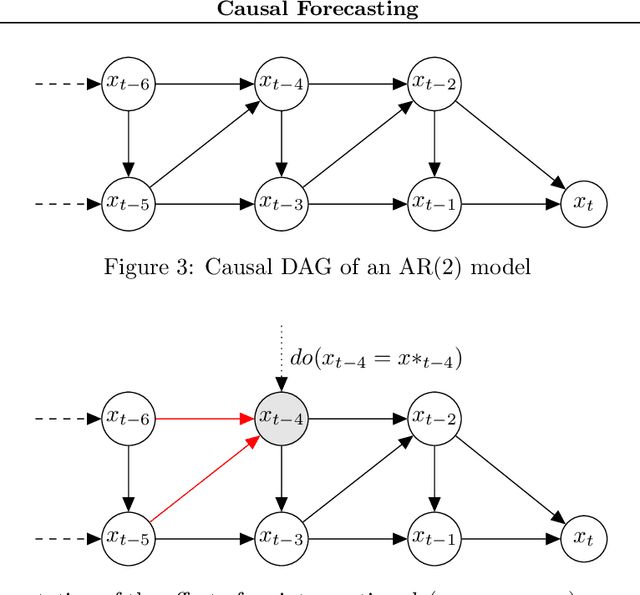
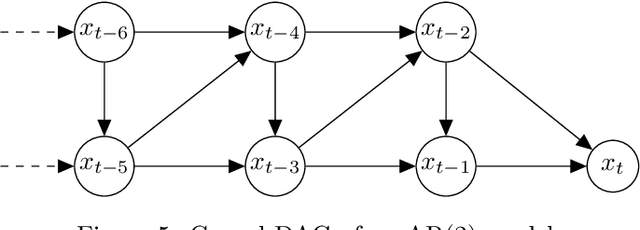
Despite the increasing relevance of forecasting methods, the causal implications of these algorithms remain largely unexplored. This is concerning considering that, even under simplifying assumptions such as causal sufficiency, the statistical risk of a model can differ significantly from its \textit{causal risk}. Here, we study the problem of *causal generalization* -- generalizing from the observational to interventional distributions -- in forecasting. Our goal is to find answers to the question: How does the efficacy of an autoregressive (VAR) model in predicting statistical associations compare with its ability to predict under interventions? To this end, we introduce the framework of *causal learning theory* for forecasting. Using this framework, we obtain a characterization of the difference between statistical and causal risks, which helps identify sources of divergence between them. Under causal sufficiency, the problem of causal generalization amounts to learning under covariate shifts albeit with additional structure (restriction to interventional distributions). This structure allows us to obtain uniform convergence bounds on causal generalizability for the class of VAR models. To the best of our knowledge, this is the first work that provides theoretical guarantees for causal generalization in the time-series setting.
Recovery Guarantees for Kernel-based Clustering under Non-parametric Mixture Models
Oct 18, 2021Leena Chennuru Vankadara, Sebastian Bordt, Ulrike von Luxburg, Debarghya Ghoshdastidar
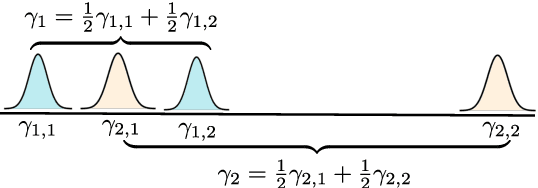
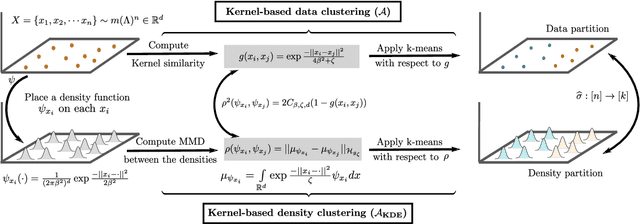
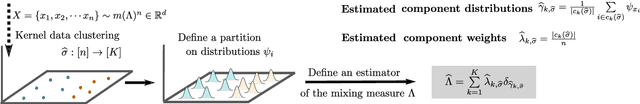
Despite the ubiquity of kernel-based clustering, surprisingly few statistical guarantees exist beyond settings that consider strong structural assumptions on the data generation process. In this work, we take a step towards bridging this gap by studying the statistical performance of kernel-based clustering algorithms under non-parametric mixture models. We provide necessary and sufficient separability conditions under which these algorithms can consistently recover the underlying true clustering. Our analysis provides guarantees for kernel clustering approaches without structural assumptions on the form of the component distributions. Additionally, we establish a key equivalence between kernel-based data-clustering and kernel density-based clustering. This enables us to provide consistency guarantees for kernel-based estimators of non-parametric mixture models. Along with theoretical implications, this connection could have practical implications, including in the systematic choice of the bandwidth of the Gaussian kernel in the context of clustering.
New Insights into Graph Convolutional Networks using Neural Tangent Kernels
Oct 08, 2021Mahalakshmi Sabanayagam, Pascal Esser, Debarghya Ghoshdastidar
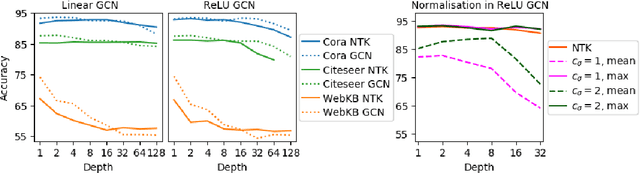
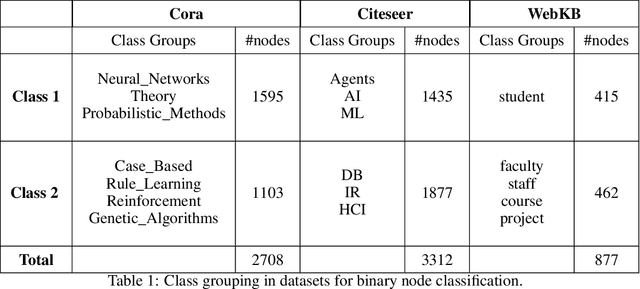
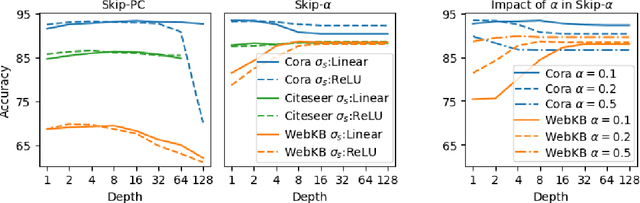
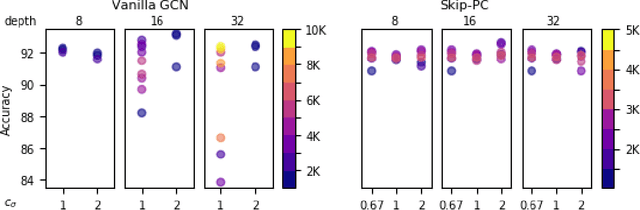
Graph Convolutional Networks (GCNs) have emerged as powerful tools for learning on network structured data. Although empirically successful, GCNs exhibit certain behaviour that has no rigorous explanation -- for instance, the performance of GCNs significantly degrades with increasing network depth, whereas it improves marginally with depth using skip connections. This paper focuses on semi-supervised learning on graphs, and explains the above observations through the lens of Neural Tangent Kernels (NTKs). We derive NTKs corresponding to infinitely wide GCNs (with and without skip connections). Subsequently, we use the derived NTKs to identify that, with suitable normalisation, network depth does not always drastically reduce the performance of GCNs -- a fact that we also validate through extensive simulation. Furthermore, we propose NTK as an efficient `surrogate model' for GCNs that does not suffer from performance fluctuations due to hyper-parameter tuning since it is a hyper-parameter free deterministic kernel. The efficacy of this idea is demonstrated through a comparison of different skip connections for GCNs using the surrogate NTKs.
Graphon based Clustering and Testing of Networks: Algorithms and Theory
Oct 06, 2021Mahalakshmi Sabanayagam, Leena Chennuru Vankadara, Debarghya Ghoshdastidar
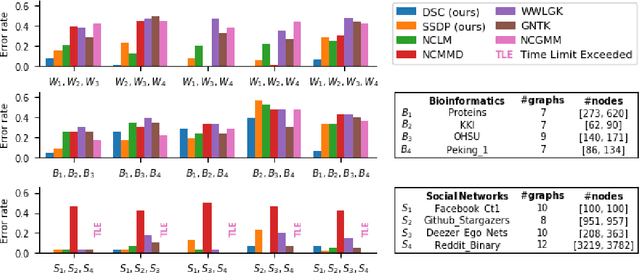
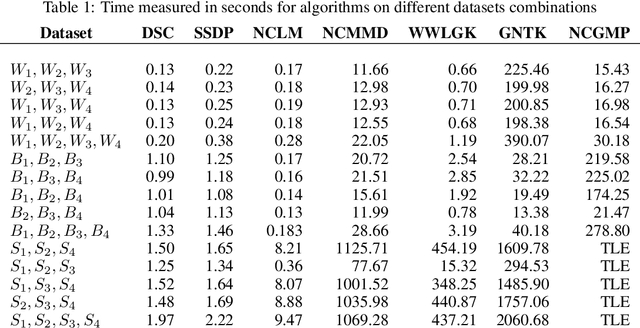
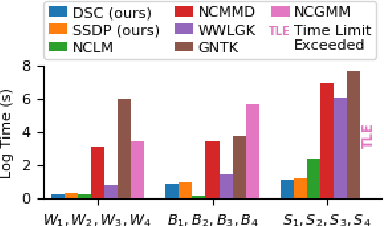
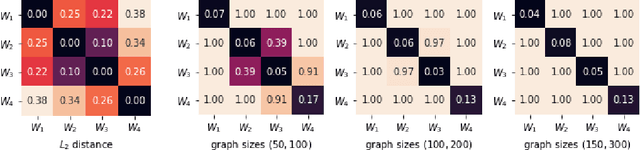
Network-valued data are encountered in a wide range of applications and pose challenges in learning due to their complex structure and absence of vertex correspondence. Typical examples of such problems include classification or grouping of protein structures and social networks. Various methods, ranging from graph kernels to graph neural networks, have been proposed that achieve some success in graph classification problems. However, most methods have limited theoretical justification, and their applicability beyond classification remains unexplored. In this work, we propose methods for clustering multiple graphs, without vertex correspondence, that are inspired by the recent literature on estimating graphons -- symmetric functions corresponding to infinite vertex limit of graphs. We propose a novel graph distance based on sorting-and-smoothing graphon estimators. Using the proposed graph distance, we present two clustering algorithms and show that they achieve state-of-the-art results. We prove the statistical consistency of both algorithms under Lipschitz assumptions on the graph degrees. We further study the applicability of the proposed distance for graph two-sample testing problems.
Near-Optimal Comparison Based Clustering
Oct 09, 2020Michaël Perrot, Pascal Mattia Esser, Debarghya Ghoshdastidar
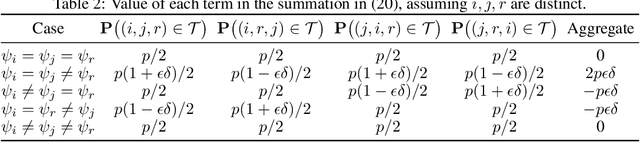
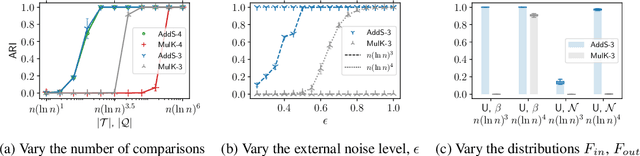
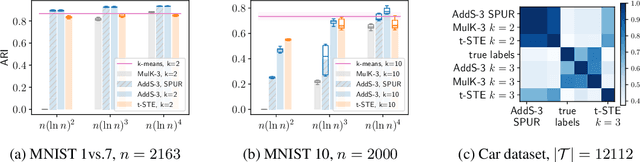

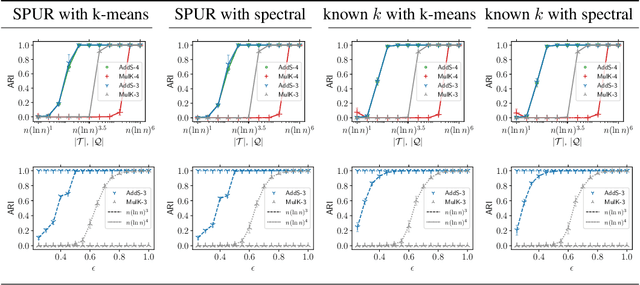
The goal of clustering is to group similar objects into meaningful partitions. This process is well understood when an explicit similarity measure between the objects is given. However, far less is known when this information is not readily available and, instead, one only observes ordinal comparisons such as "object i is more similar to j than to k." In this paper, we tackle this problem using a two-step procedure: we estimate a pairwise similarity matrix from the comparisons before using a clustering method based on semi-definite programming (SDP). We theoretically show that our approach can exactly recover a planted clustering using a near-optimal number of passive comparisons. We empirically validate our theoretical findings and demonstrate the good behaviour of our method on real data.
On the optimality of kernels for high-dimensional clustering
Dec 01, 2019Leena Chennuru Vankadara, Debarghya Ghoshdastidar
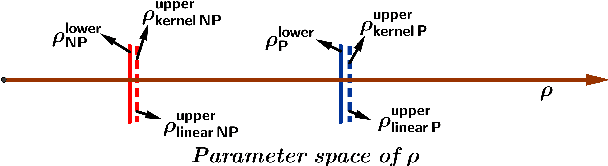
This paper studies the optimality of kernel methods in high-dimensional data clustering. Recent works have studied the large sample performance of kernel clustering in the high-dimensional regime, where Euclidean distance becomes less informative. However, it is unknown whether popular methods, such as kernel k-means, are optimal in this regime. We consider the problem of high-dimensional Gaussian clustering and show that, with the exponential kernel function, the sufficient conditions for partial recovery of clusters using the NP-hard kernel k-means objective matches the known information-theoretic limit up to a factor of $\sqrt{2}$ for large $k$. It also exactly matches the known upper bounds for the non-kernel setting. We also show that a semi-definite relaxation of the kernel k-means procedure matches up to constant factors, the spectral threshold, below which no polynomial-time algorithm is known to succeed. This is the first work that provides such optimality guarantees for the kernel k-means as well as its convex relaxation. Our proofs demonstrate the utility of the less known polynomial concentration results for random variables with exponentially decaying tails in a higher-order analysis of kernel methods.
Practical methods for graph two-sample testing
Nov 30, 2018Debarghya Ghoshdastidar, Ulrike von Luxburg
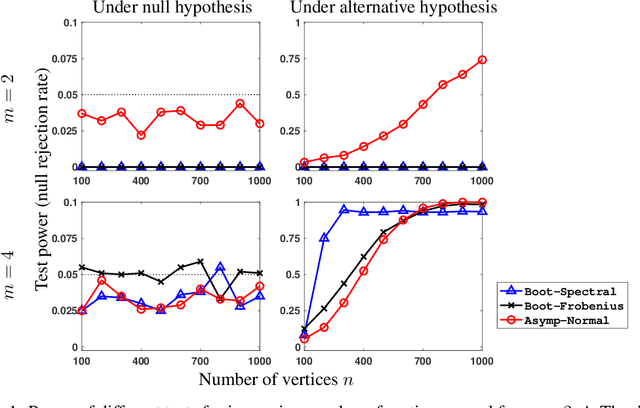
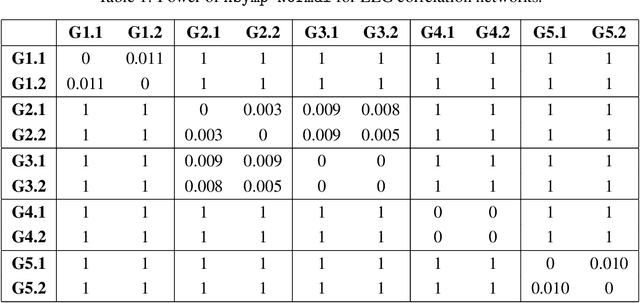
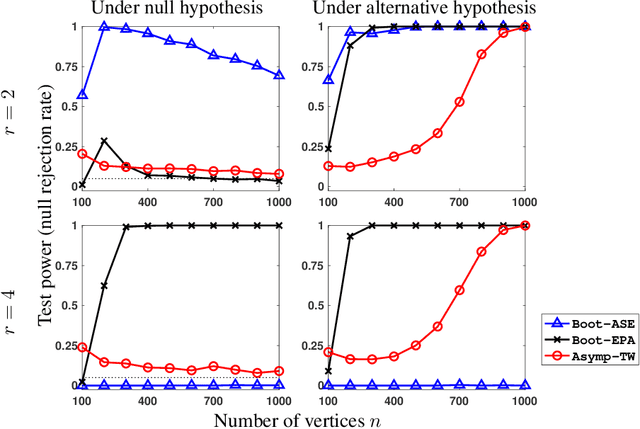
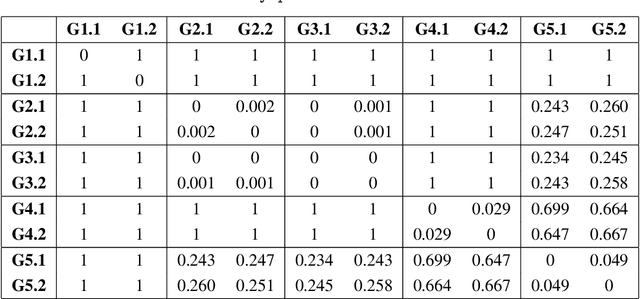
Hypothesis testing for graphs has been an important tool in applied research fields for more than two decades, and still remains a challenging problem as one often needs to draw inference from few replicates of large graphs. Recent studies in statistics and learning theory have provided some theoretical insights about such high-dimensional graph testing problems, but the practicality of the developed theoretical methods remains an open question. In this paper, we consider the problem of two-sample testing of large graphs. We demonstrate the practical merits and limitations of existing theoretical tests and their bootstrapped variants. We also propose two new tests based on asymptotic distributions. We show that these tests are computationally less expensive and, in some cases, more reliable than the existing methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge