Darlington Ahiale Akogo
A Preliminary Exploration into an Alternative CellLineNet: An Evolutionary Approach
Jul 26, 2020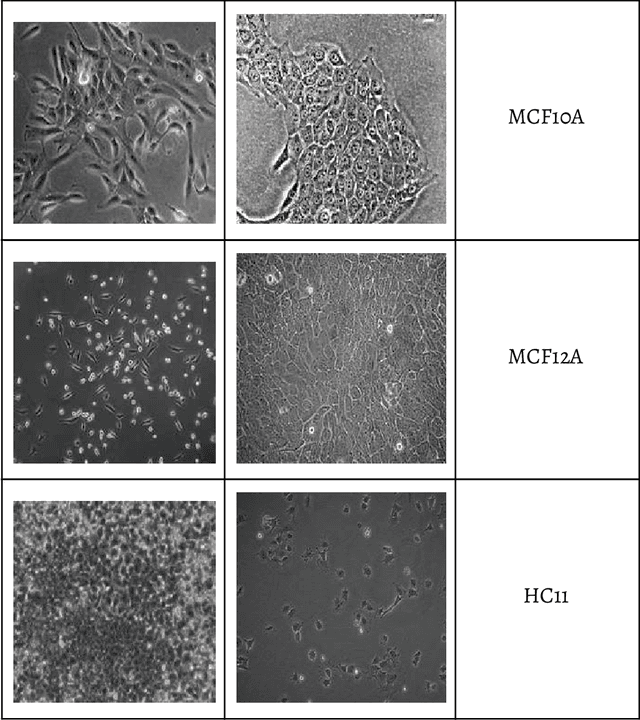
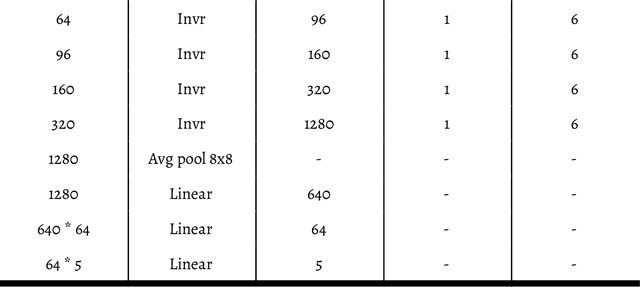
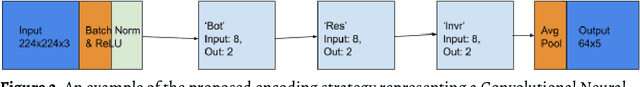
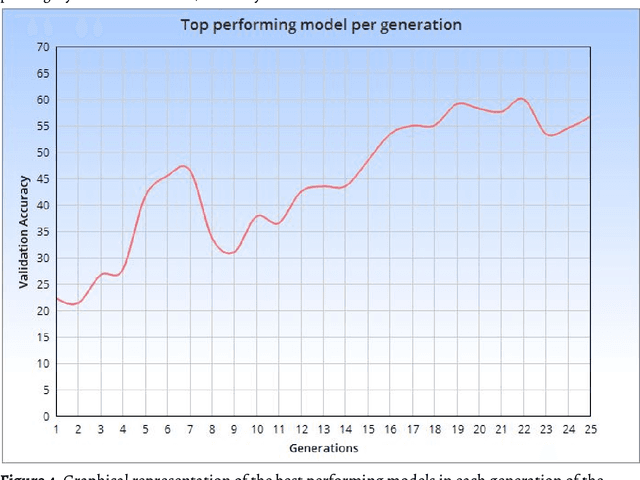
Abstract:Within this paper, the exploration of an evolutionary approach to an alternative CellLineNet: a convolutional neural network adept at the classification of epithelial breast cancer cell lines, is presented. This evolutionary algorithm introduces control variables that guide the search of architectures in the search space of inverted residual blocks, bottleneck blocks, residual blocks and a basic 2x2 convolutional block. The promise of EvoCELL is predicting what combination or arrangement of the feature extracting blocks that produce the best model architecture for a given task. Therein, the performance of how the fittest model evolved after each generation is shown. The final evolved model CellLineNet V2 classifies 5 types of epithelial breast cell lines consisting of two human cancer lines, 2 normal immortalized lines, and 1 immortalized mouse line (MDA-MB-468, MCF7, 10A, 12A and HC11). The Multiclass Cell Line Classification Convolutional Neural Network extends our earlier work on a Binary Breast Cancer Cell Line Classification model. This paper presents an on-going exploratory approach to neural network architecture design and is presented for further study.
CellLineNet: End-to-End Learning and Transfer Learning For Multiclass Epithelial Breast cell Line Classification via a Convolutional Neural Network
Aug 18, 2018



Abstract:Computer Vision for Analyzing and Classifying cells and tissues often require rigorous lab procedures and so automated Computer Vision solutions have been sought. Most work in such field usually requires Feature Extractions before the analysis of such features via Machine Learning and Machine Vision algorithms. We developed a Convolutional Neural Network that classifies 5 types of epithelial breast cell lines comprised of two human cancer lines, 2 normal immortalized lines, and 1 immortalized mouse line (MDA-MB-468, MCF7, 10A, 12A and HC11) without requiring feature extraction. The Multiclass Cell Line Classification Convolutional Neural Network extends our earlier work on a Binary Breast Cancer Cell Line Classification model. CellLineNet is 31-layer Convolutional Neural Network trained, validated and tested on a 3,252 image dataset of 5 types of Epithelial Breast cell Lines (MDA-MB-468, MCF7, 10A, 12A and HC11) in an end-to-end fashion. End-to-End Learning enables CellLineNet to identify and learn on its own, visual features and regularities most important to Breast Cancer Cell Line Classification from the dataset of images. Using Transfer Learning, the 28-layer MobileNet Convolutional Neural Network architecture with pre-trained ImageNet weights is extended and fine tuned to the Multiclass Epithelial Breast cell Line Classification problem. CellLineNet simply requires an imaged Cell Line as input and it outputs the type of breast epithelial cell line (MDA-MB-468, MCF7, 10A, 12A or HC11) as predicted probabilities for the 5 classes. CellLineNet scored a 96.67% Accuracy.
End-to-End Learning via a Convolutional Neural Network for Cancer Cell Line Classification
Jul 25, 2018



Abstract:Computer Vision for automated analysis of cells and tissues usually include extracting features from images before analyzing such features via various Machine Learning and Machine Vision algorithms. We developed a Convolutional Neural Network model that classifies MDA-MB-468 and MCF7 breast cancer cells via brightfield microscopy images without the need of any prior feature extraction. Our 6-layer Convolutional Neural Network is directly trained, validated and tested on 1,241 images of MDA-MB-468 and MCF7 breast cancer cell line in an end-to-end fashion, allowing a system to distinguish between different cancer cell types. The model takes in as input imaged breast cancer cell line and then outputs the cell line type (MDA-MB-468 or MCF7) as predicted probabilities between the two classes. Our model scored a 99% Accuracy.
ScaffoldNet: Detecting and Classifying Biomedical Polymer-Based Scaffolds via a Convolutional Neural Network
May 17, 2018



Abstract:We developed a Convolutional Neural Network model to identify and classify Airbrushed (alternatively known as Blow-spun), Electrospun and Steel Wire scaffolds. Our model ScaffoldNet is a 6-layer Convolutional Neural Network trained and tested on 3,043 images of Airbrushed, Electrospun and Steel Wire scaffolds. The model takes in as input an imaged scaffold and then outputs the scaffold type (Airbrushed, Electrospun or Steel Wire) as predicted probabilities for the 3 classes. Our model scored a 99.44% Accuracy, demonstrating potential for adaptation to investigating and solving complex machine learning problems aimed at abstract spatial contexts, or in screening complex, biological, fibrous structures seen in cortical bone and fibrous shells.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge