Daniel de Marchi
University of North Carolina at Chapel Hill
Shapley Marginal Surplus for Strong Models
Aug 16, 2024Abstract:Shapley values have seen widespread use in machine learning as a way to explain model predictions and estimate the importance of covariates. Accurately explaining models is critical in real-world models to both aid in decision making and to infer the properties of the true data-generating process (DGP). In this paper, we demonstrate that while model-based Shapley values might be accurate explainers of model predictions, machine learning models themselves are often poor explainers of the DGP even if the model is highly accurate. Particularly in the presence of interrelated or noisy variables, the output of a highly predictive model may fail to account for these relationships. This implies explanations of a trained model's behavior may fail to provide meaningful insight into the DGP. In this paper we introduce a novel variable importance algorithm, Shapley Marginal Surplus for Strong Models, that samples the space of possible models to come up with an inferential measure of feature importance. We compare this method to other popular feature importance methods, both Shapley-based and non-Shapley based, and demonstrate significant outperformance in inferential capabilities relative to other methods.
Revisiting Bellman Errors for Offline Model Selection
Jan 31, 2023



Abstract:Offline model selection (OMS), that is, choosing the best policy from a set of many policies given only logged data, is crucial for applying offline RL in real-world settings. One idea that has been extensively explored is to select policies based on the mean squared Bellman error (MSBE) of the associated Q-functions. However, previous work has struggled to obtain adequate OMS performance with Bellman errors, leading many researchers to abandon the idea. Through theoretical and empirical analyses, we elucidate why previous work has seen pessimistic results with Bellman errors and identify conditions under which OMS algorithms based on Bellman errors will perform well. Moreover, we develop a new estimator of the MSBE that is more accurate than prior methods and obtains impressive OMS performance on diverse discrete control tasks, including Atari games. We open-source our data and code to enable researchers to conduct OMS experiments more easily.
A Robust Hypothesis Test for Tree Ensemble Pruning
Jan 25, 2023Abstract:Gradient boosted decision trees are some of the most popular algorithms in applied machine learning. They are a flexible and powerful tool that can robustly fit to any tabular dataset in a scalable and computationally efficient way. One of the most critical parameters to tune when fitting these models are the various penalty terms used to distinguish signal from noise in the current model. These penalties are effective in practice, but are lacking in robust theoretical justifications. In this paper we develop and present a novel theoretically justified hypothesis test of split quality for gradient boosted tree ensembles and demonstrate that using this method instead of the common penalty terms leads to a significant reduction in out of sample loss. Additionally, this method provides a theoretically well-justified stopping condition for the tree growing algorithm. We also present several innovative extensions to the method, opening the door for a wide variety of novel tree pruning algorithms.
Augmenting Molecular Images with Vector Representations as a Featurization Technique for Drug Classification
Aug 09, 2020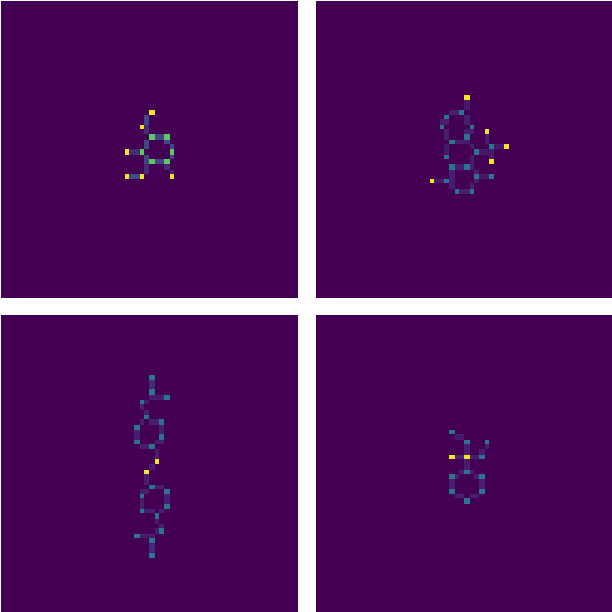
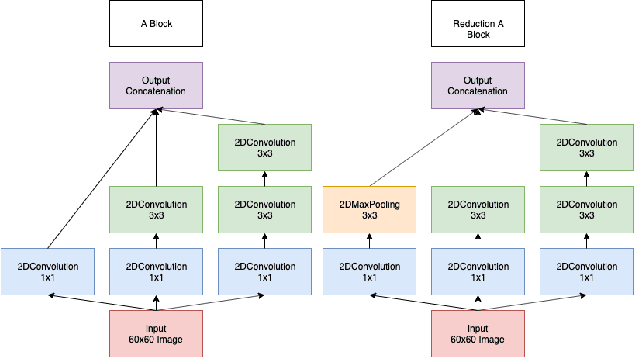
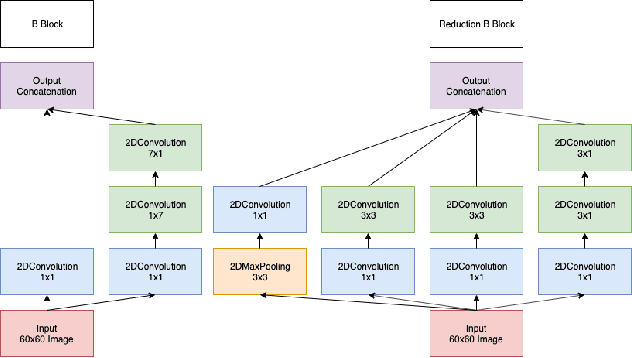
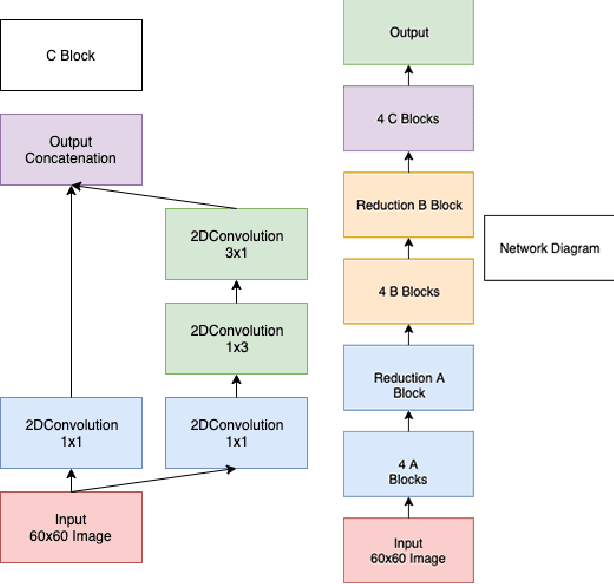
Abstract:One of the key steps in building deep learning systems for drug classification and generation is the choice of featurization for the molecules. Previous featurization methods have included molecular images, binary strings, graphs, and SMILES strings. This paper proposes the creation of molecular images captioned with binary vectors that encode information not contained in or easily understood from a molecular image alone. Specifically, we use Morgan fingerprints, which encode higher level structural information, and MACCS keys, which encode yes or no questions about a molecules properties and structure. We tested our method on the HIV dataset published by the Pande lab, which consists of 41,127 molecules labeled by if they inhibit the HIV virus. Our final model achieved a state of the art AUC ROC on the HIV dataset, outperforming all other methods. Moreover, the model converged significantly faster than most other methods, requiring dramatically less computational power than unaugmented images.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge