Dalibor Stys
Cell segmentation from telecentric bright-field transmitted light microscopic images using a Residual Attention U-Net: a case study on HeLa line
Mar 23, 2022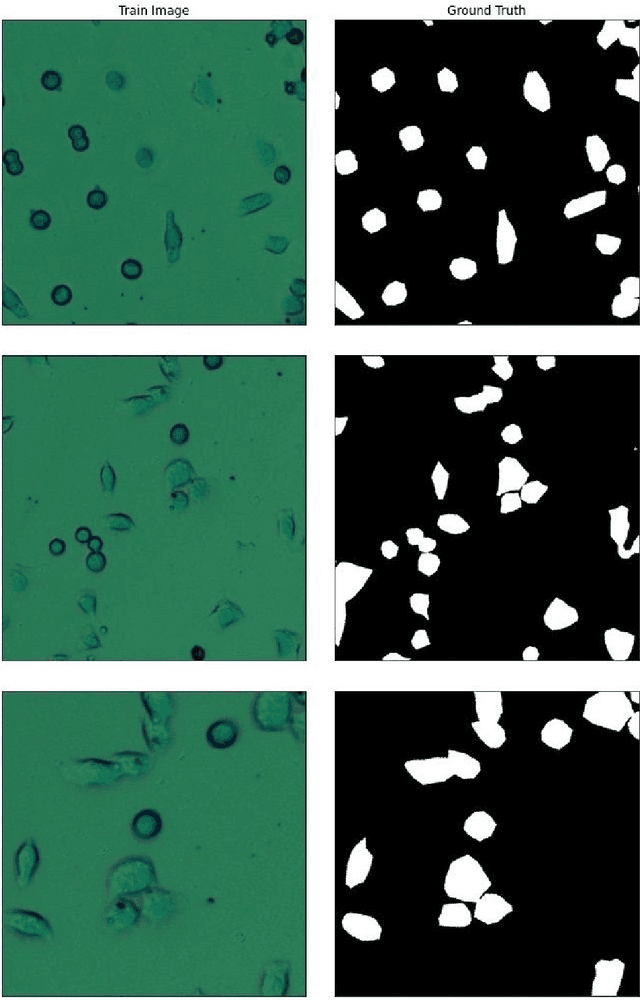

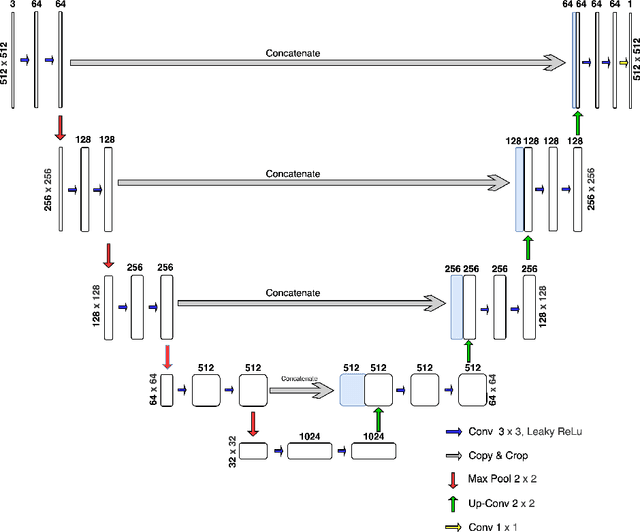
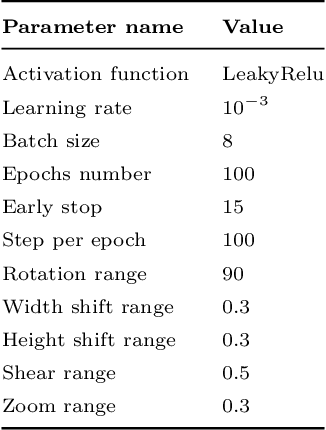
Abstract:Living cell segmentation from bright-field light microscopic images is challenging due to the image complexity and temporal changes in the living cells. Recently developed deep learning (DL)-based methods became popular in medical and microscopic image segmentation tasks due to their success and promising outcomes. The main objective of this paper is to develop a deep learning, UNet-based method to segment the living cells of the HeLa line in bright-field transmitted light microscopy. To find the most suitable architecture for our datasets, we have proposed a residual attention U-Net and compared it with an attention and a simple U-Net architecture. The attention mechanism highlights the remarkable features and suppresses activations in the irrelevant image regions. The residual mechanism overcomes with vanishing gradient problem. The Mean-IoU score for our datasets reaches 0.9505, 0.9524, and 0.9530 for the simple, attention, and residual attention U-Net, respectively. We achieved the most accurate semantic segmentation results in the Mean-IoU and Dice metrics by applying the residual and attention mechanisms together. The watershed method applied to this best - Residual Attention - semantic segmentation result gave the segmentation with the specific information for each cell.
Technically correct visualization of biological microscopic experiments
Mar 14, 2019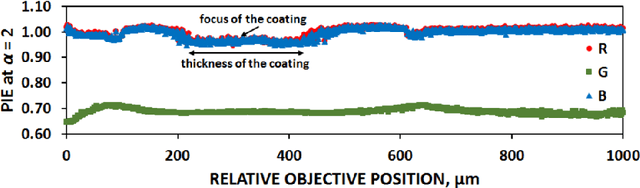
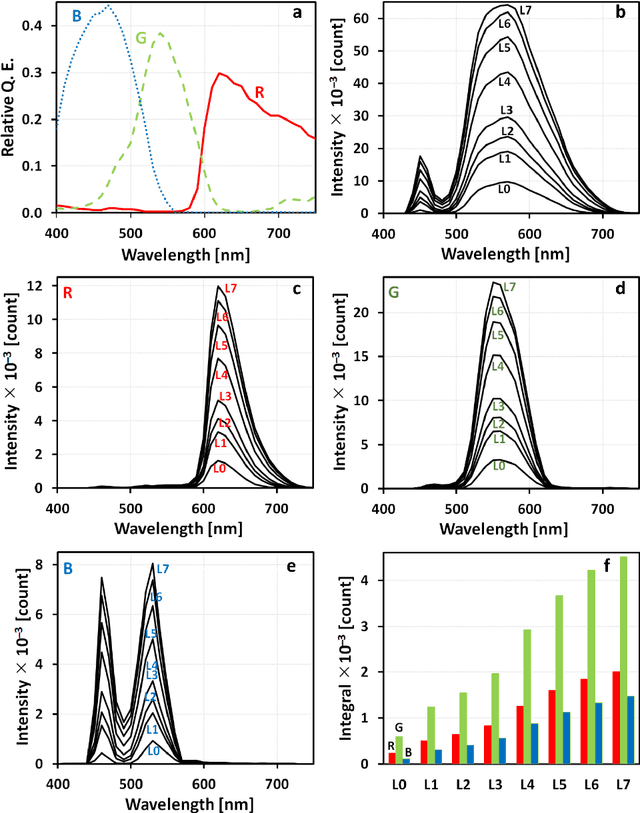
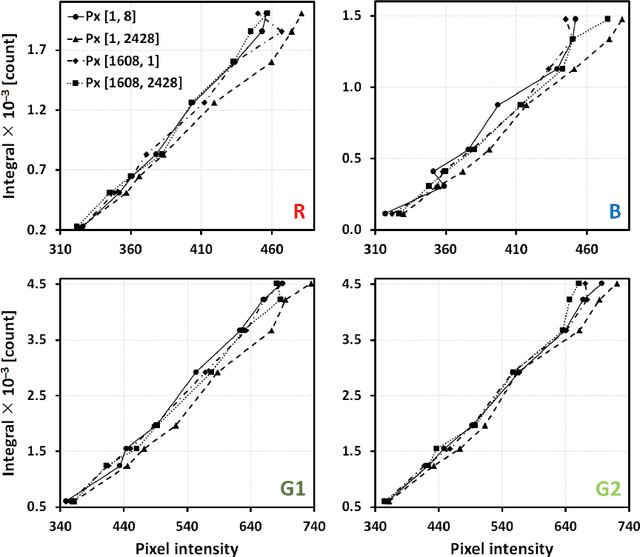
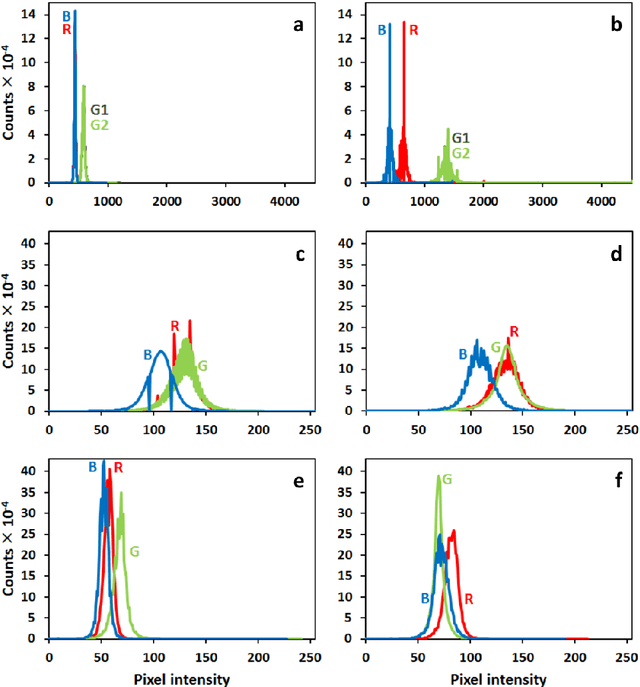
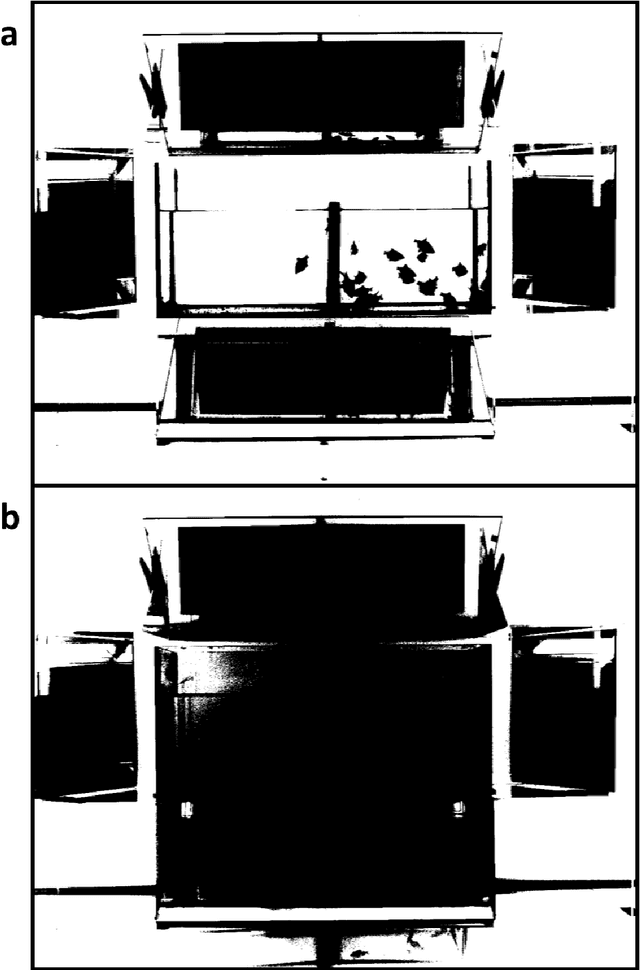
Abstract:The most realistic images that reflect native cellular and intracellular structure and behavior can be obtained only using brightfield microscopy. At high-intensity pulsing LED illumination, we captured the primary 12-bit-per-channel (bpc) signal from an observed sample using a brightfield microscope equipped with a high-resolution (4872x3248) camera. In order to suppress image distortions arising from light passing through the whole microscope optical path, from camera sensor defects and geometrical peculiarities of sensor sensitivity, these uncompressed 12-bpc images underwent a kind of correction after simultaneous calibration of all the parts of the experimental arrangement. Moreover, the final corrected images (from biological experiments) show the number of photons reaching each camera pixel and can be visualized in 8-bpc intensity depth after the Least Information Loss compression (Stys et al., Lect. Notes Bioinform. 9656, 2016).
Colorimetric Calibration of a Digital Camera
Aug 14, 2017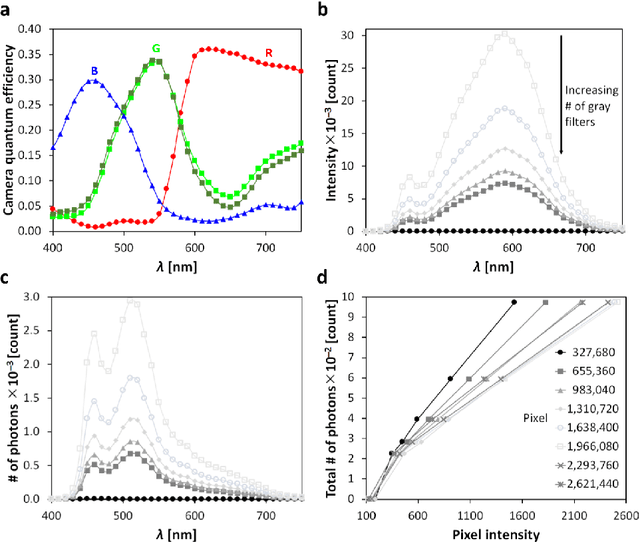
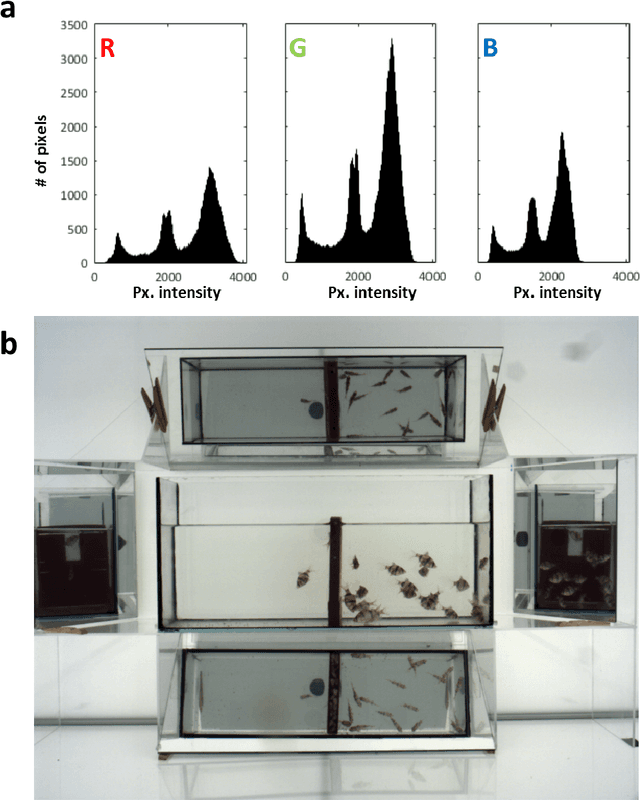
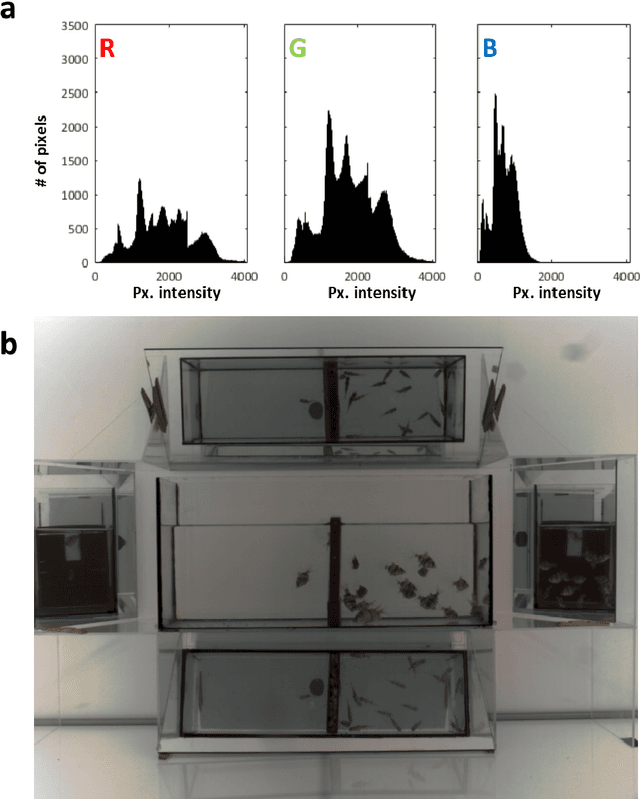

Abstract:In this paper, we introduce a novel - physico-chemical - approach for calibration of a digital camera chip. This approach utilizes results of measurement of incident light spectra of calibration films of different levels of gray for construction of calibration curve (number of incident photons vs. image pixel intensity) for each camera pixel. We show spectral characteristics of such corrected digital raw image files (a primary camera signal) and demonstrate their suitability for next image processing and analysis.
Observation of dynamics inside an unlabeled live cell using bright-field photon microscopy: Evaluation of organelles' trajectories
Dec 13, 2016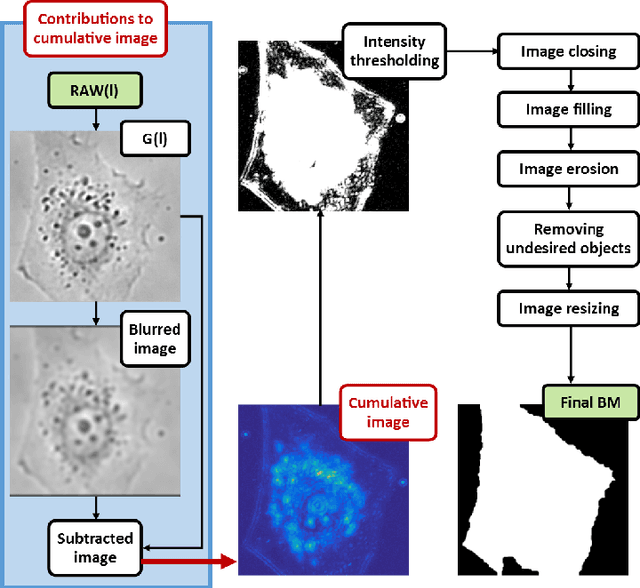
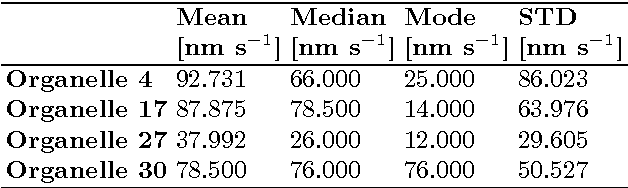
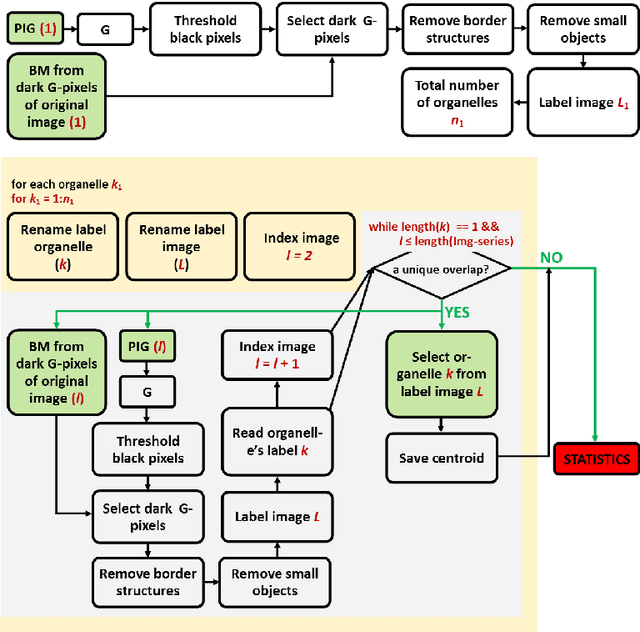
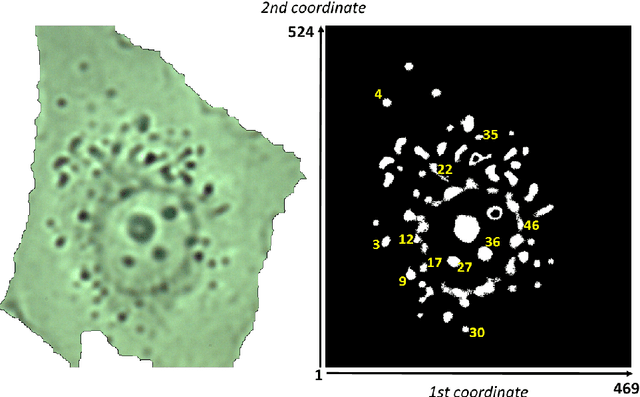
Abstract:This article presents an algorithm for the evaluation of organelles' movements inside of an unmodified live cell. We used a time-lapse image series obtained using wide-field bright-field photon transmission microscopy as an algorithm input. The benefit of the algorithm is the application of the R\'enyi information entropy, namely a variable called a point information gain, which enables to highlight the borders of the intracellular organelles and to localize the organelles' centers of mass with the precision of one pixel.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge