Christopher Meek
Graphical Models and Exponential Families
Jan 30, 2013
Abstract:We provide a classification of graphical models according to their representation as subfamilies of exponential families. Undirected graphical models with no hidden variables are linear exponential families (LEFs), directed acyclic graphical models and chain graphs with no hidden variables, including Bayesian networks with several families of local distributions, are curved exponential families (CEFs) and graphical models with hidden variables are stratified exponential families (SEFs). An SEF is a finite union of CEFs satisfying a frontier condition. In addition, we illustrate how one can automatically generate independence and non-independence constraints on the distributions over the observable variables implied by a Bayesian network with hidden variables. The relevance of these results for model selection is examined.
Quantifier Elimination for Statistical Problems
Jan 23, 2013
Abstract:Recent improvement on Tarski's procedure for quantifier elimination in the first order theory of real numbers makes it feasible to solve small instances of the following problems completely automatically: 1. listing all equality and inequality constraints implied by a graphical model with hidden variables. 2. Comparing graphyical models with hidden variables (i.e., model equivalence, inclusion, and overlap). 3. Answering questions about the identification of a model or portion of a model, and about bounds on quantities derived from a model. 4. Determing whether a given set of independence assertions. We discuss the foundation of quantifier elimination and demonstrate its application to these problems.
Dependency Networks for Collaborative Filtering and Data Visualization
Jan 16, 2013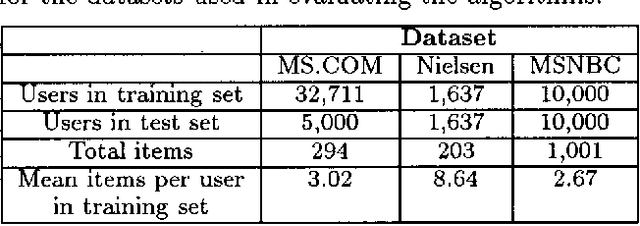
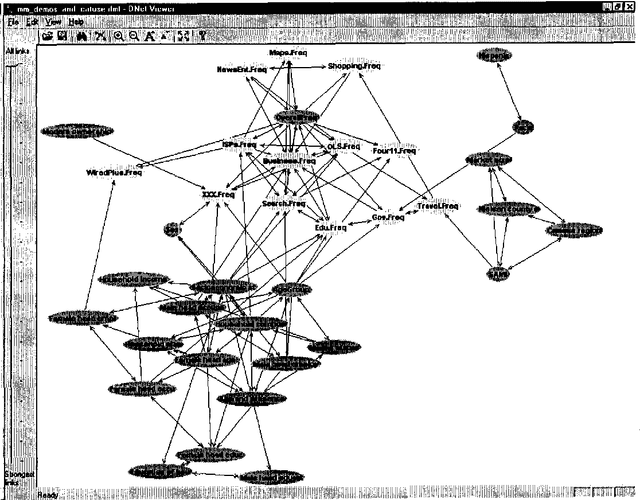
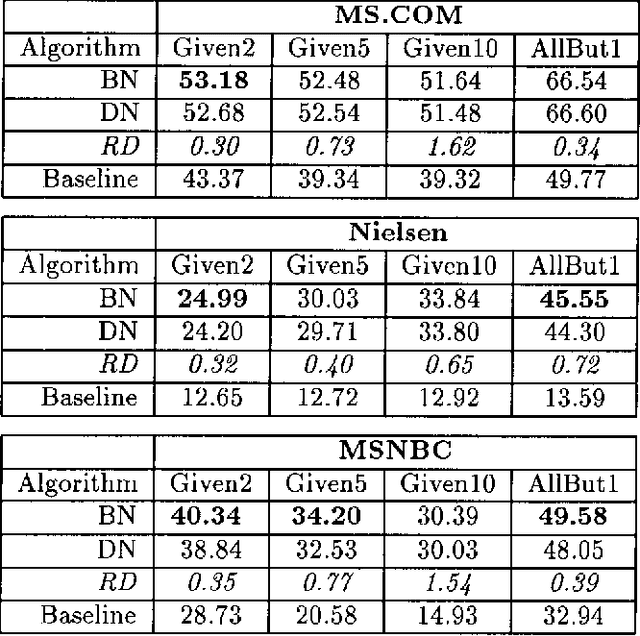
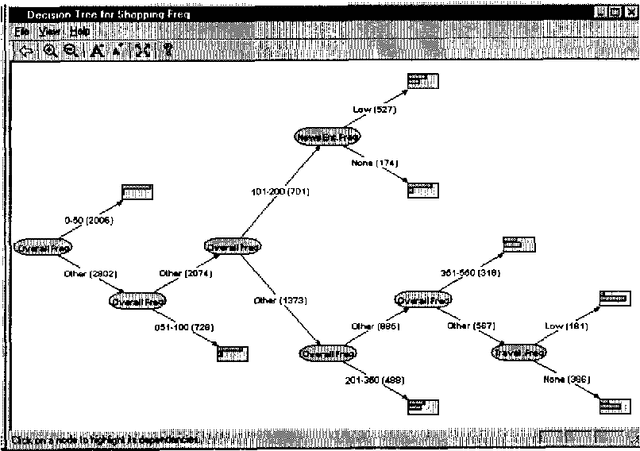
Abstract:We describe a graphical model for probabilistic relationships---an alternative to the Bayesian network---called a dependency network. The graph of a dependency network, unlike a Bayesian network, is potentially cyclic. The probability component of a dependency network, like a Bayesian network, is a set of conditional distributions, one for each node given its parents. We identify several basic properties of this representation and describe a computationally efficient procedure for learning the graph and probability components from data. We describe the application of this representation to probabilistic inference, collaborative filtering (the task of predicting preferences), and the visualization of acausal predictive relationships.
Perfect Tree-Like Markovian Distributions
Jan 16, 2013Abstract:We show that if a strictly positive joint probability distribution for a set of binary random variables factors according to a tree, then vertex separation represents all and only the independence relations enclosed in the distribution. The same result is shown to hold also for multivariate strictly positive normal distributions. Our proof uses a new property of conditional independence that holds for these two classes of probability distributions.
Using Temporal Data for Making Recommendations
Jan 10, 2013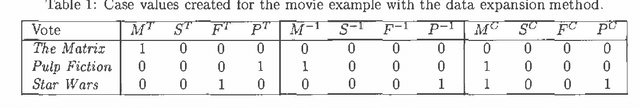
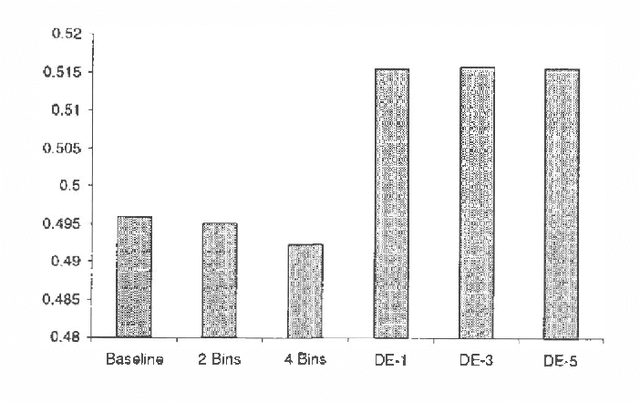
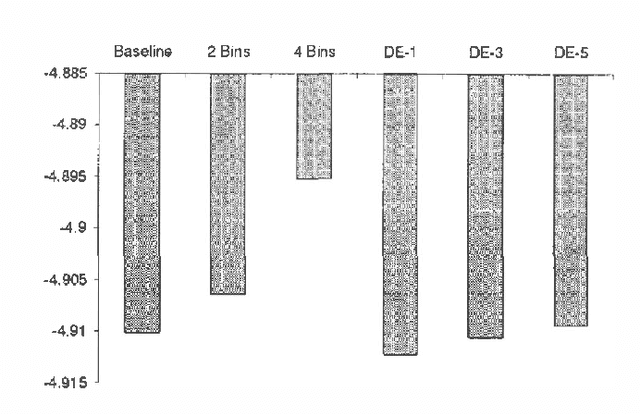
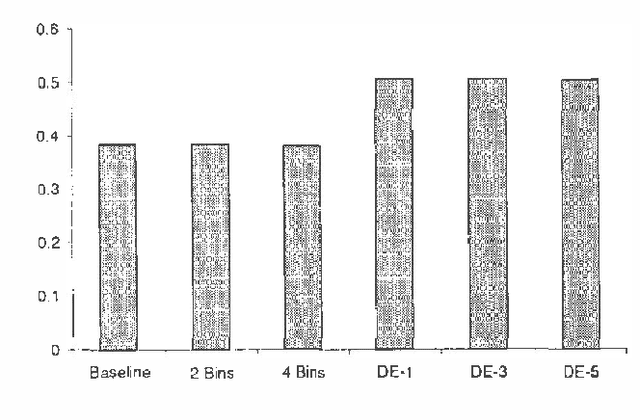
Abstract:We treat collaborative filtering as a univariate time series estimation problem: given a user's previous votes, predict the next vote. We describe two families of methods for transforming data to encode time order in ways amenable to off-the-shelf classification and density estimation tools, and examine the results of using these approaches on several real-world data sets. The improvements in predictive accuracy we realize recommend the use of other predictive algorithms that exploit the temporal order of data.
Staged Mixture Modelling and Boosting
Dec 12, 2012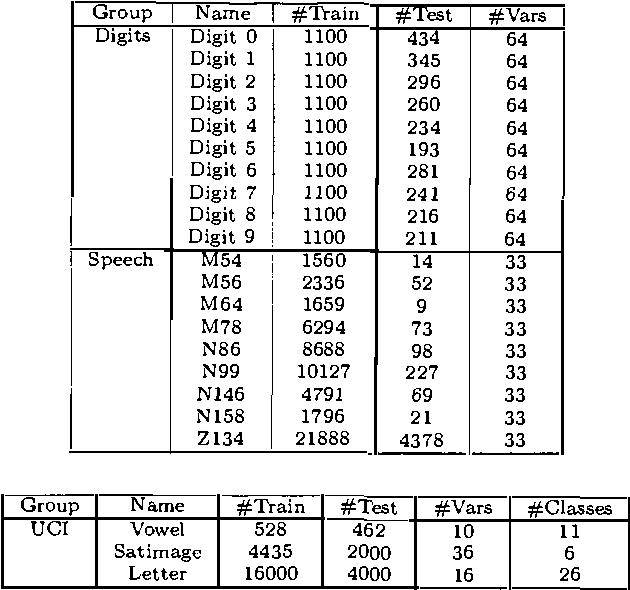
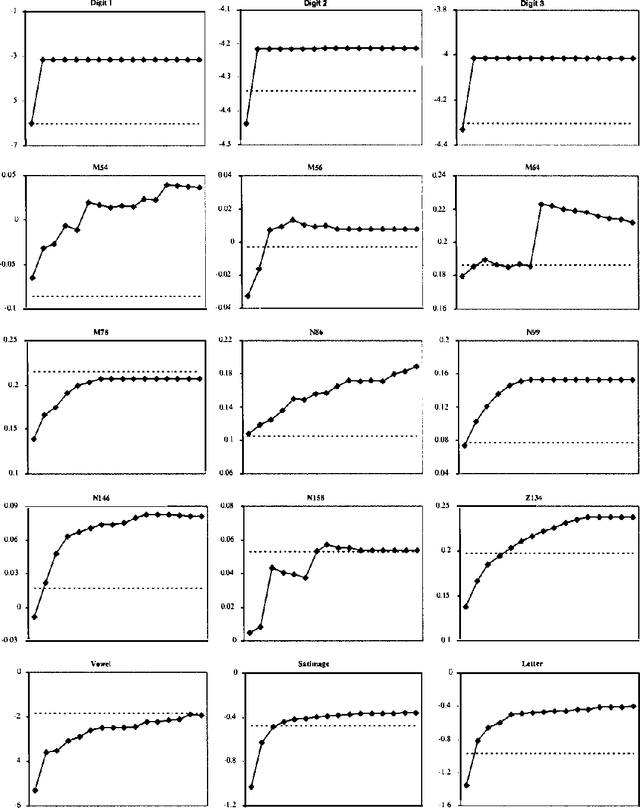

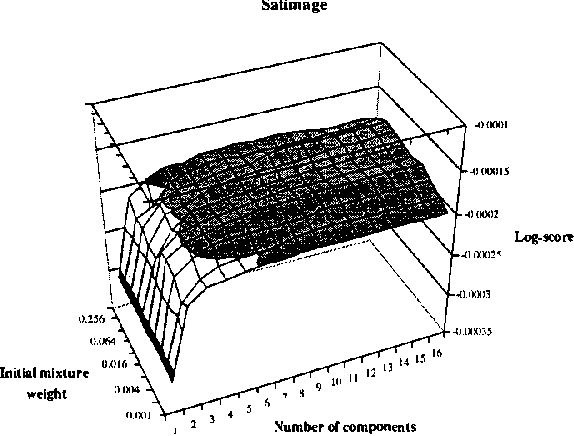
Abstract:In this paper, we introduce and evaluate a data-driven staged mixture modeling technique for building density, regression, and classification models. Our basic approach is to sequentially add components to a finite mixture model using the structural expectation maximization (SEM) algorithm. We show that our technique is qualitatively similar to boosting. This correspondence is a natural byproduct of the fact that we use the SEM algorithm to sequentially fit the mixture model. Finally, in our experimental evaluation, we demonstrate the effectiveness of our approach on a variety of prediction and density estimation tasks using real-world data.
Factorization of Discrete Probability Distributions
Dec 12, 2012Abstract:We formulate necessary and sufficient conditions for an arbitrary discrete probability distribution to factor according to an undirected graphical model, or a log-linear model, or other more general exponential models. This result generalizes the well known Hammersley-Clifford Theorem.
Finding Optimal Bayesian Networks
Dec 12, 2012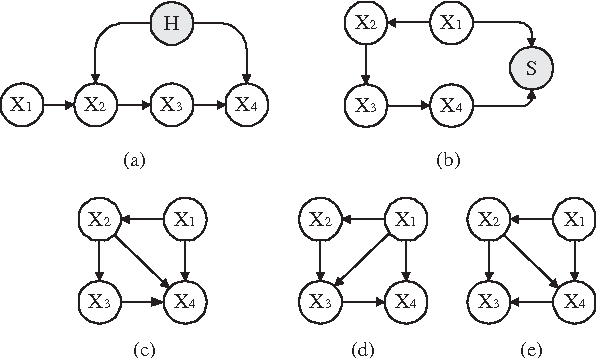
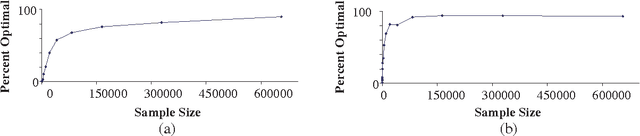
Abstract:In this paper, we derive optimality results for greedy Bayesian-network search algorithms that perform single-edge modifications at each step and use asymptotically consistent scoring criteria. Our results extend those of Meek (1997) and Chickering (2002), who demonstrate that in the limit of large datasets, if the generative distribution is perfect with respect to a DAG defined over the observable variables, such search algorithms will identify this optimal (i.e. generative) DAG model. We relax their assumption about the generative distribution, and assume only that this distribution satisfies the {em composition property} over the observable variables, which is a more realistic assumption for real domains. Under this assumption, we guarantee that the search algorithms identify an {em inclusion-optimal} model; that is, a model that (1) contains the generative distribution and (2) has no sub-model that contains this distribution. In addition, we show that the composition property is guaranteed to hold whenever the dependence relationships in the generative distribution can be characterized by paths between singleton elements in some generative graphical model (e.g. a DAG, a chain graph, or a Markov network) even when the generative model includes unobserved variables, and even when the observed data is subject to selection bias.
Practically Perfect
Oct 19, 2012Abstract:The property of perfectness plays an important role in the theory of Bayesian networks. First, the existence of perfect distributions for arbitrary sets of variables and directed acyclic graphs implies that various methods for reading independence from the structure of the graph (e.g., Pearl, 1988; Lauritzen, Dawid, Larsen & Leimer, 1990) are complete. Second, the asymptotic reliability of various search methods is guaranteed under the assumption that the generating distribution is perfect (e.g., Spirtes, Glymour & Scheines, 2000; Chickering & Meek, 2002). We provide a lower-bound on the probability of sampling a non-perfect distribution when using a fixed number of bits to represent the parameters of the Bayesian network. This bound approaches zero exponentially fast as one increases the number of bits used to represent the parameters. This result implies that perfect distributions with fixed-length representations exist. We also provide a lower-bound on the number of bits needed to guarantee that a distribution sampled from a uniform Dirichlet distribution is perfect with probability greater than 1/2. This result is useful for constructing randomized reductions for hardness proofs.
Large-Sample Learning of Bayesian Networks is NP-Hard
Oct 19, 2012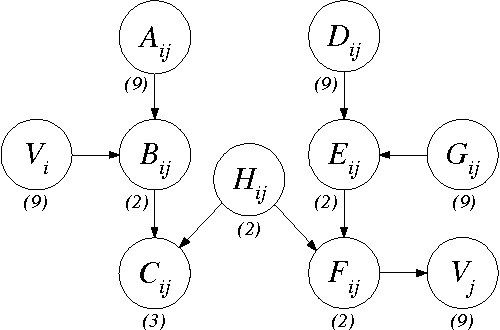
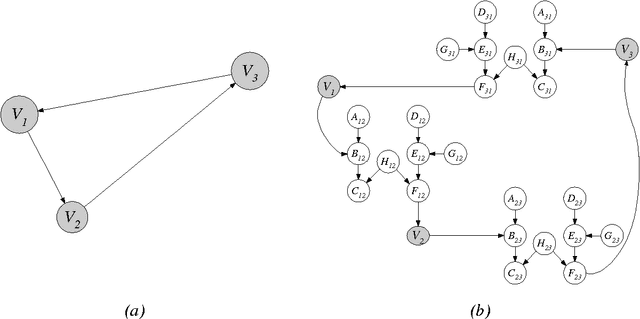
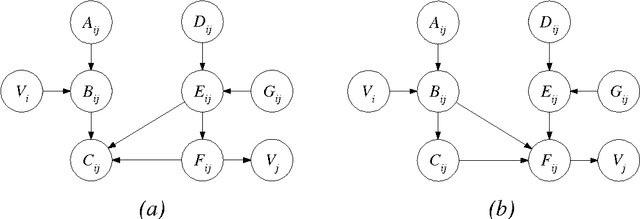
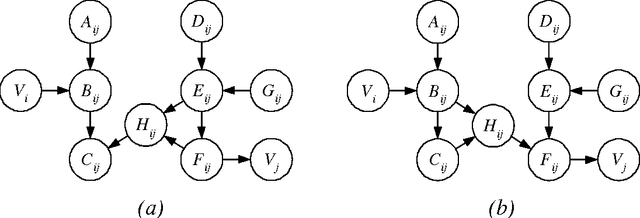
Abstract:In this paper, we provide new complexity results for algorithms that learn discrete-variable Bayesian networks from data. Our results apply whenever the learning algorithm uses a scoring criterion that favors the simplest model able to represent the generative distribution exactly. Our results therefore hold whenever the learning algorithm uses a consistent scoring criterion and is applied to a sufficiently large dataset. We show that identifying high-scoring structures is hard, even when we are given an independence oracle, an inference oracle, and/or an information oracle. Our negative results also apply to the learning of discrete-variable Bayesian networks in which each node has at most k parents, for all k > 3.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge