Chihye Han
MHSAN: Multi-Head Self-Attention Network for Visual Semantic Embedding
Jan 11, 2020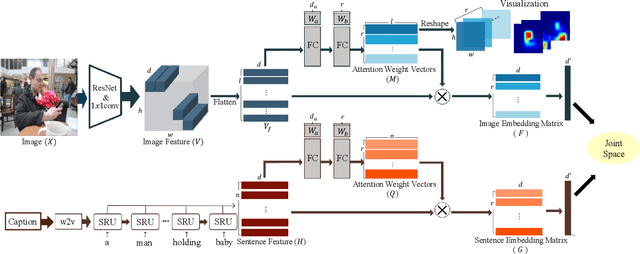
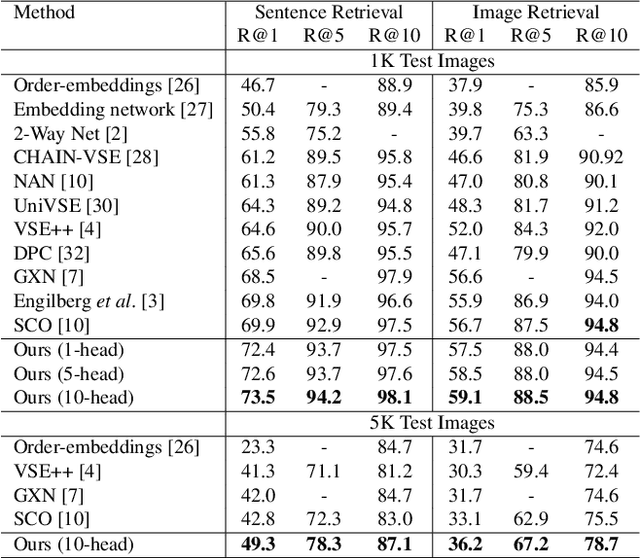
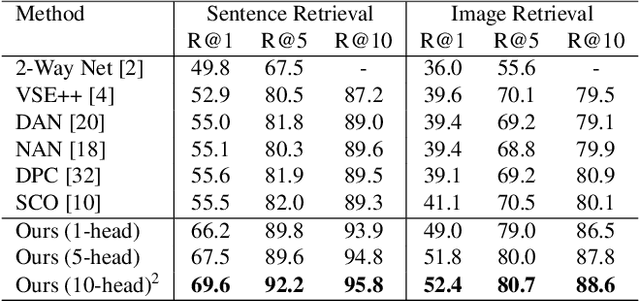

Abstract:Visual-semantic embedding enables various tasks such as image-text retrieval, image captioning, and visual question answering. The key to successful visual-semantic embedding is to express visual and textual data properly by accounting for their intricate relationship. While previous studies have achieved much advance by encoding the visual and textual data into a joint space where similar concepts are closely located, they often represent data by a single vector ignoring the presence of multiple important components in an image or text. Thus, in addition to the joint embedding space, we propose a novel multi-head self-attention network to capture various components of visual and textual data by attending to important parts in data. Our approach achieves the new state-of-the-art results in image-text retrieval tasks on MS-COCO and Flicker30K datasets. Through the visualization of the attention maps that capture distinct semantic components at multiple positions in the image and the text, we demonstrate that our method achieves an effective and interpretable visual-semantic joint space.
Generation of 3D Brain MRI Using Auto-Encoding Generative Adversarial Networks
Aug 07, 2019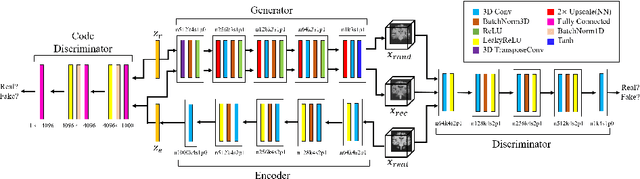
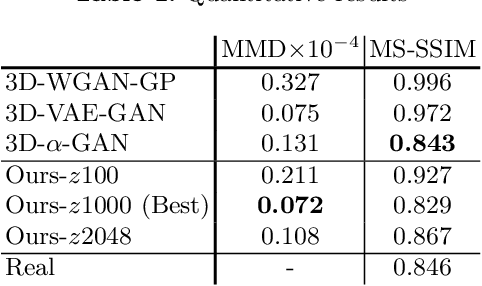
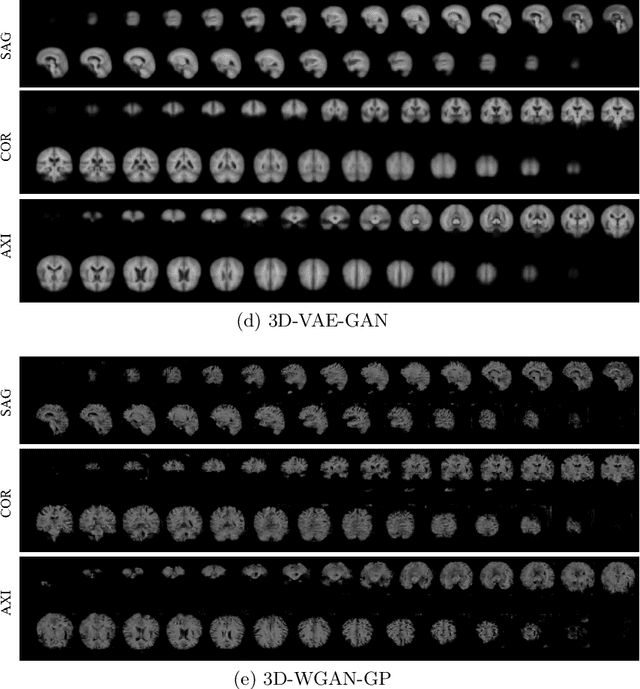
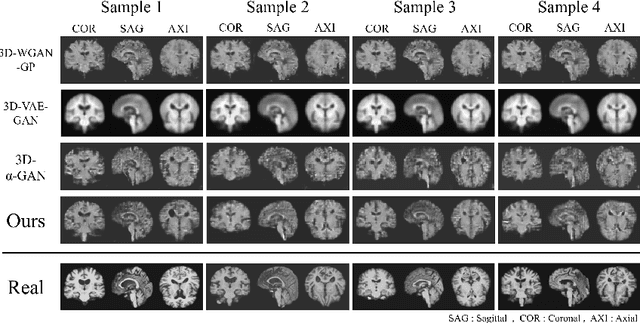
Abstract:As deep learning is showing unprecedented success in medical image analysis tasks, the lack of sufficient medical data is emerging as a critical problem. While recent attempts to solve the limited data problem using Generative Adversarial Networks (GAN) have been successful in generating realistic images with diversity, most of them are based on image-to-image translation and thus require extensive datasets from different domains. Here, we propose a novel model that can successfully generate 3D brain MRI data from random vectors by learning the data distribution. Our 3D GAN model solves both image blurriness and mode collapse problems by leveraging alpha-GAN that combines the advantages of Variational Auto-Encoder (VAE) and GAN with an additional code discriminator network. We also use the Wasserstein GAN with Gradient Penalty (WGAN-GP) loss to lower the training instability. To demonstrate the effectiveness of our model, we generate new images of normal brain MRI and show that our model outperforms baseline models in both quantitative and qualitative measurements. We also train the model to synthesize brain disorder MRI data to demonstrate the wide applicability of our model. Our results suggest that the proposed model can successfully generate various types and modalities of 3D whole brain volumes from a small set of training data.
Representation of White- and Black-Box Adversarial Examples in Deep Neural Networks and Humans: A Functional Magnetic Resonance Imaging Study
May 07, 2019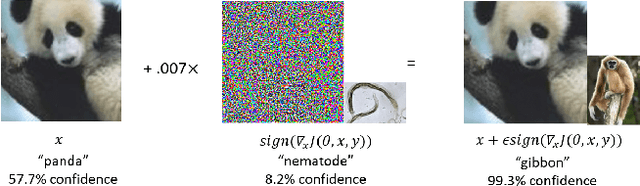
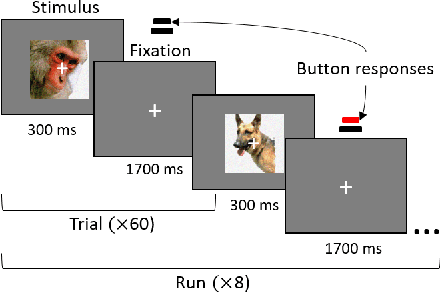
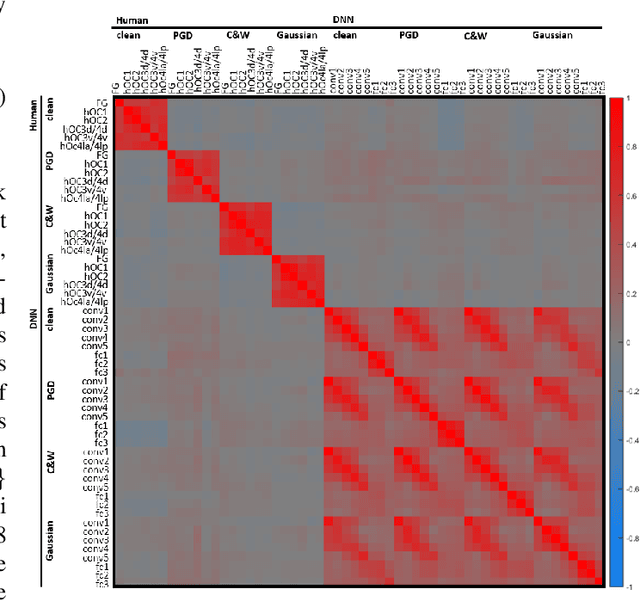
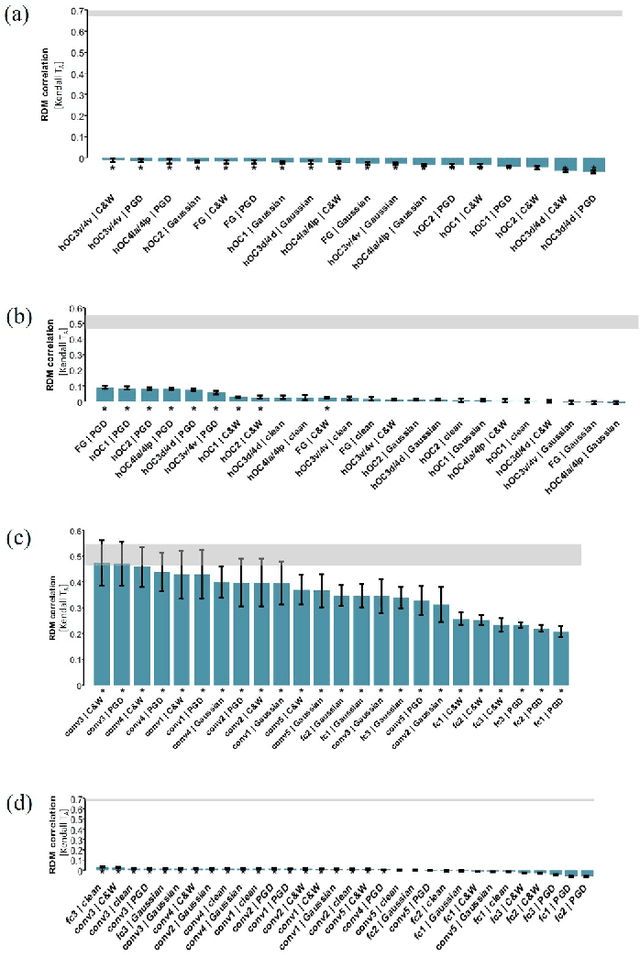
Abstract:The recent success of brain-inspired deep neural networks (DNNs) in solving complex, high-level visual tasks has led to rising expectations for their potential to match the human visual system. However, DNNs exhibit idiosyncrasies that suggest their visual representation and processing might be substantially different from human vision. One limitation of DNNs is that they are vulnerable to adversarial examples, input images on which subtle, carefully designed noises are added to fool a machine classifier. The robustness of the human visual system against adversarial examples is potentially of great importance as it could uncover a key mechanistic feature that machine vision is yet to incorporate. In this study, we compare the visual representations of white- and black-box adversarial examples in DNNs and humans by leveraging functional magnetic resonance imaging (fMRI). We find a small but significant difference in representation patterns for different (i.e. white- versus black- box) types of adversarial examples for both humans and DNNs. However, human performance on categorical judgment is not degraded by noise regardless of the type unlike DNN. These results suggest that adversarial examples may be differentially represented in the human visual system, but unable to affect the perceptual experience.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge