Brendon K. Colbert
Efficient Convex Algorithms for Universal Kernel Learning
Apr 15, 2023Abstract:The accuracy and complexity of machine learning algorithms based on kernel optimization are determined by the set of kernels over which they are able to optimize. An ideal set of kernels should: admit a linear parameterization (for tractability); be dense in the set of all kernels (for robustness); be universal (for accuracy). Recently, a framework was proposed for using positive matrices to parameterize a class of positive semi-separable kernels. Although this class can be shown to meet all three criteria, previous algorithms for optimization of such kernels were limited to classification and furthermore relied on computationally complex Semidefinite Programming (SDP) algorithms. In this paper, we pose the problem of learning semiseparable kernels as a minimax optimization problem and propose a SVD-QCQP primal-dual algorithm which dramatically reduces the computational complexity as compared with previous SDP-based approaches. Furthermore, we provide an efficient implementation of this algorithm for both classification and regression -- an implementation which enables us to solve problems with 100 features and up to 30,000 datums. Finally, when applied to benchmark data, the algorithm demonstrates the potential for significant improvement in accuracy over typical (but non-convex) approaches such as Neural Nets and Random Forest with similar or better computation time.
Employing Feature Selection Algorithms to Determine the Immune State of Mice with Rheumatoid Arthritis
Jul 12, 2022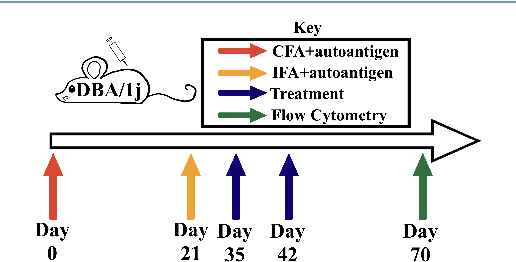
Abstract:The immune response is a dynamic process by which the body determines whether an antigen is self or nonself. The state of this dynamic process is defined by the relative balance and population of inflammatory and regulatory actors which comprise this decision making process. The goal of immunotherapy as applied to, e.g. Rheumatoid Arthritis (RA), then, is to bias the immune state in favor of the regulatory actors - thereby shutting down autoimmune pathways in the response. While there are several known approaches to immunotherapy, the effectiveness of the therapy will depend on how this intervention alters the evolution of this state. Unfortunately, this process is determined not only by the dynamics of the process, but the state of the system at the time of intervention - a state which is difficult if not impossible to determine prior to application of the therapy.
A New Algorithm for Tessellated Kernel Learning
Jun 13, 2020
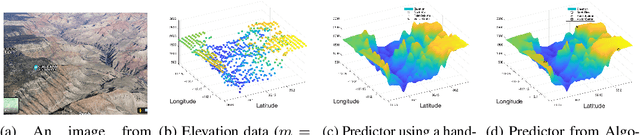
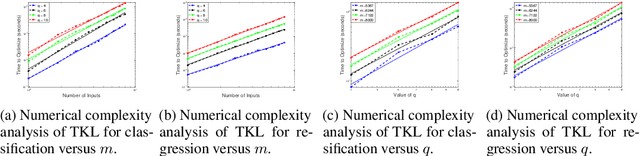
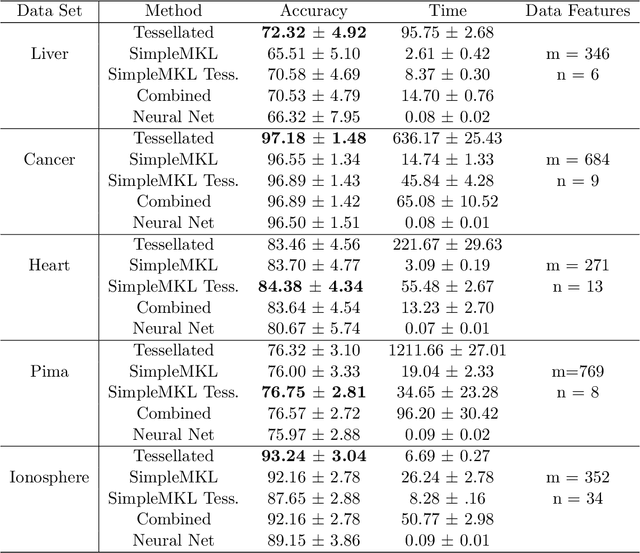
Abstract:The accuracy and complexity of machine learning algorithms based on kernel optimization are limited by the set of kernels over which they are able to optimize. An ideal set of kernels should: admit a linear parameterization (for tractability); be dense in the set of all kernels (for robustness); be universal (for accuracy). The recently proposed Tesselated Kernels (TKs) is currently the only known class which meets all three criteria. However, previous algorithms for optimizing TKs were limited to classification and relied on Semidefinite Programming (SDP) - limiting them to relatively small datasets. By contrast, the 2-step algorithm proposed here scales to 10,000 data points and extends to the regression problem. Furthermore, when applied to benchmark data, the algorithm demonstrates significant improvement in performance over Neural Nets and SimpleMKL with similar computation time.
A Convex Parametrization of a New Class of Universal Kernel Functions for use in Kernel Learning
Nov 15, 2017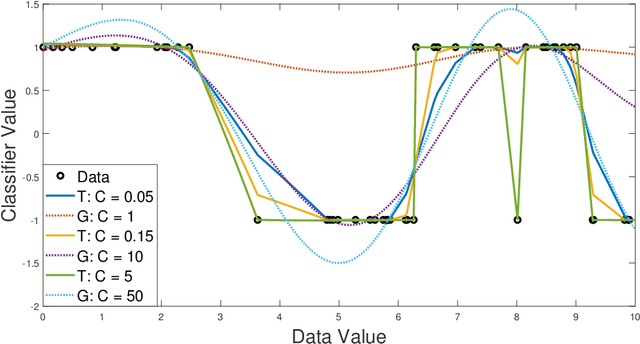
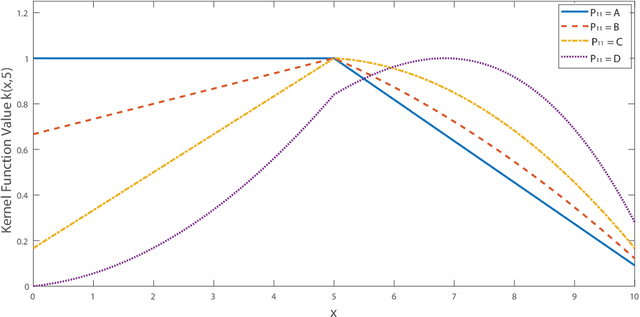
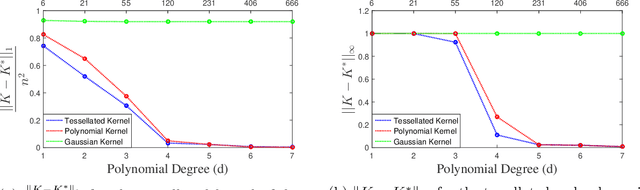
Abstract:We propose a new class of universal kernel functions which admit a linear parametrization using positive semidefinite matrices. These kernels are generalizations of the Sobolev kernel and are defined by piecewise-polynomial functions. The class of kernels is termed "tessellated" as the resulting discriminant is defined piecewise with hyper-rectangular domains whose corners are determined by the training data. The kernels have scalable complexity, but each instance is universal in the sense that its hypothesis space is dense in $L_2$. Using numerical testing, we show that for the soft margin SVM, this class can eliminate the need for Gaussian kernels. Furthermore, we demonstrate that when the ratio of the number of training data to features is high, this method will significantly outperform other kernel learning algorithms. Finally, to reduce the complexity associated with SDP-based kernel learning methods, we use a randomized basis for the positive matrices to integrate with existing multiple kernel learning algorithms such as SimpleMKL.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge