Bradley Wright
Semi-Supervised Natural Language Approach for Fine-Grained Classification of Medical Reports
Nov 14, 2019
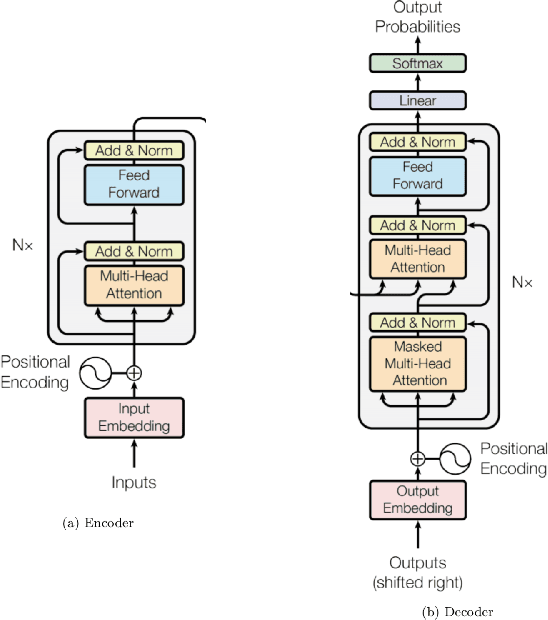
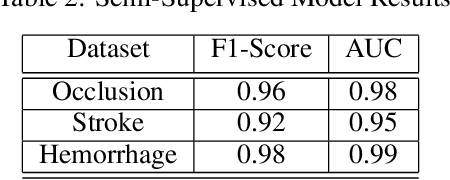
Abstract:Although machine learning has become a powerful tool to augment doctors in clinical analysis, the immense amount of labeled data that is necessary to train supervised learning approaches burdens each development task as time and resource intensive. The vast majority of dense clinical information is stored in written reports, detailing pertinent patient information. The challenge with utilizing natural language data for standard model development is due to the complex nature of the modality. In this research, a model pipeline was developed to utilize an unsupervised approach to train an encoder-language model, a recurrent network, to generate document encodings; which then can be used as features passed into a decoder-classifier model that requires magnitudes less labeled data than previous approaches to differentiate between fine-grained disease classes accurately. The language model was trained on unlabeled radiology reports from the Massachusetts General Hospital Radiology Department (n=218,159) and terminated with a loss of 1.62. The classification models were trained on three labeled datasets of head CT studies of reported patients, presenting large vessel occlusion (n=1403), acute ischemic strokes (n=331), and intracranial hemorrhage (n=4350), to identify a variety of different findings directly from the radiology report data; resulting in AUCs of 0.98, 0.95, and 0.99, respectively, for the large vessel occlusion, acute ischemic stroke, and intracranial hemorrhage datasets. The output encodings are able to be used in conjunction with imaging data, to create models that can process a multitude of different modalities. The ability to automatically extract relevant features from textual data allows for faster model development and integration of textual modality, overall, allowing clinical reports to become a more viable input for more encompassing and accurate deep learning models.
Fully-Automated Analysis of Body Composition from CT in Cancer Patients Using Convolutional Neural Networks
Aug 11, 2018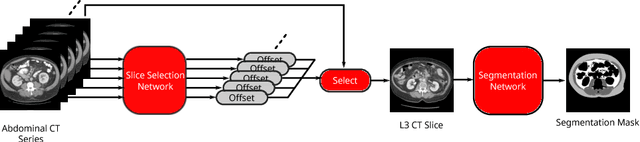
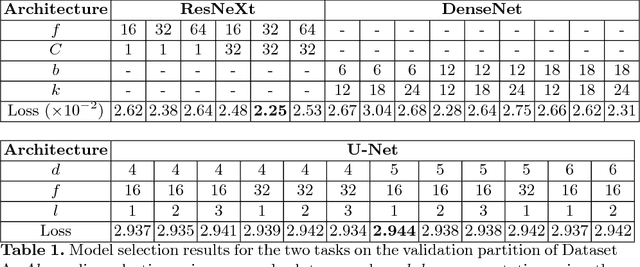
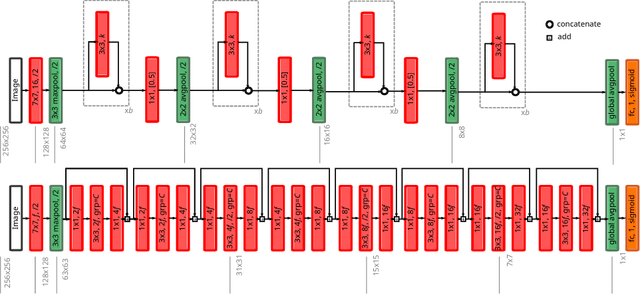
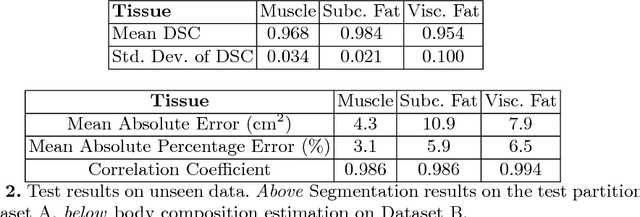
Abstract:The amounts of muscle and fat in a person's body, known as body composition, are correlated with cancer risks, cancer survival, and cardiovascular risk. The current gold standard for measuring body composition requires time-consuming manual segmentation of CT images by an expert reader. In this work, we describe a two-step process to fully automate the analysis of CT body composition using a DenseNet to select the CT slice and U-Net to perform segmentation. We train and test our methods on independent cohorts. Our results show Dice scores (0.95-0.98) and correlation coefficients (R=0.99) that are favorable compared to human readers. These results suggest that fully automated body composition analysis is feasible, which could enable both clinical use and large-scale population studies.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge