Bernhard Renger
Whole-body CT attenuation and volume charts from routine clinical scans via evidence-grounded LLM report filtering
May 07, 2026Abstract:Interpreting quantitative CT biomarkers, such as organ volume and tissue attenuation, requires large-scale healthy reference distributions. However, creating these is challenging because clinical datasets are often heavily enriched with pathology. Here, we develop an evidence-grounded, cross-verified large language model (LLM) ensemble to filter pathological findings from radiology reports, enabling the construction of pathology-reduced cohorts from over 350,000 CT examinations. Five LLMs, first, flag structure-level abnormality candidates grounded in verbatim report evidence and, second, resolve disagreements via cross-verification. Using distribution-aware generalized additive models for location, scale, and shape, we establish comprehensive whole-body reference charts for 106 anatomical structures (volumes and attenuation) across adulthood, accounting for age, sex, contrast enhancement, and acquisition parameters. Longitudinal analyses reveal structure- and contrast-dependent changes distinct from cross-sectional trends. These resources facilitate covariate-adjusted centile scoring from routine CT, supporting standardized quantitative phenotyping, multi-site imaging studies, and scalable opportunistic screening research.
Per-Pixel Lung Thickness and Lung Capacity Estimation on Chest X-Rays using Convolutional Neural Networks
Oct 27, 2021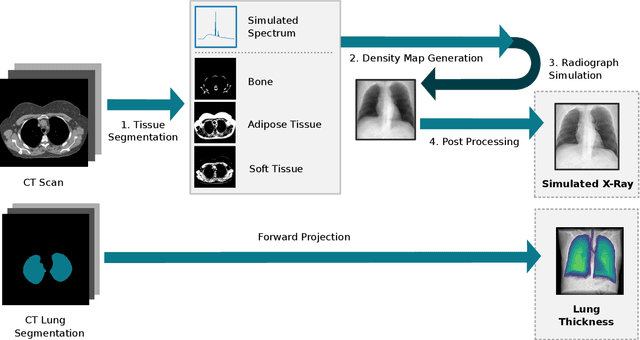
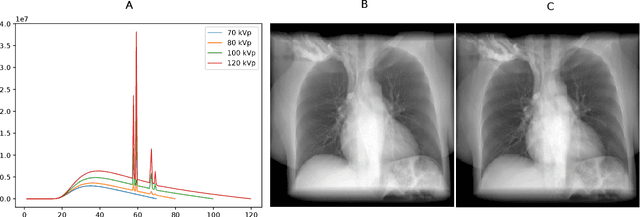
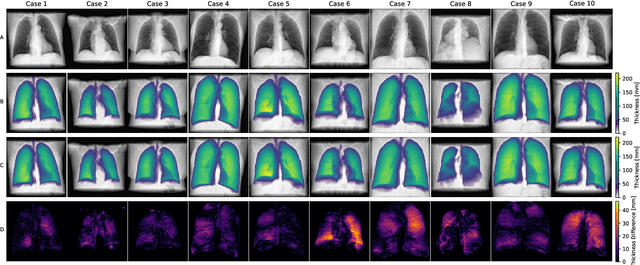
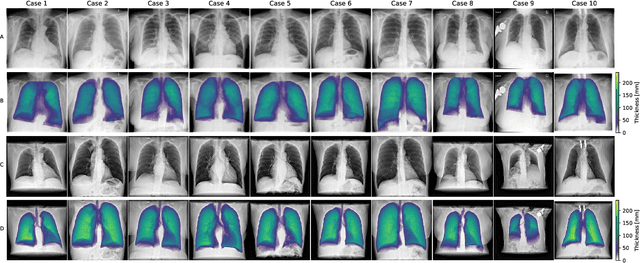
Abstract:Estimating the lung depth on x-ray images could provide both an accurate opportunistic lung volume estimation during clinical routine and improve image contrast in modern structural chest imaging techniques like x-ray dark-field imaging. We present a method based on a convolutional neural network that allows a per-pixel lung thickness estimation and subsequent total lung capacity estimation. The network was trained and validated using 5250 simulated radiographs generated from 525 real CT scans. Furthermore, we are able to infer the model trained with simulation data on real radiographs. For 35 patients, quantitative and qualitative evaluation was performed on standard clinical radiographs. The ground-truth for each patient's total lung volume was defined based on the patients' corresponding CT scan. The mean-absolute error between the estimated lung volume on the 35 real radiographs and groundtruth volume was 0.73 liter. Additionally, we predicted the lung thicknesses on a synthetic dataset of 131 radiographs, where the mean-absolute error was 0.27 liter. The results show, that it is possible to transfer the knowledge obtained in a simulation model to real x-ray images.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge