B. Scellier
Frequency propagation: Multi-mechanism learning in nonlinear physical networks
Aug 10, 2022Abstract:We introduce frequency propagation, a learning algorithm for nonlinear physical networks. In a resistive electrical circuit with variable resistors, an activation current is applied at a set of input nodes at one frequency, and an error current is applied at a set of output nodes at another frequency. The voltage response of the circuit to these boundary currents is the superposition of an `activation signal' and an `error signal' whose coefficients can be read in different frequencies of the frequency domain. Each conductance is updated proportionally to the product of the two coefficients. The learning rule is local and proved to perform gradient descent on a loss function. We argue that frequency propagation is an instance of a multi-mechanism learning strategy for physical networks, be it resistive, elastic, or flow networks. Multi-mechanism learning strategies incorporate at least two physical quantities, potentially governed by independent physical mechanisms, to act as activation and error signals in the training process. Locally available information about these two signals is then used to update the trainable parameters to perform gradient descent. We demonstrate how earlier work implementing learning via chemical signaling in flow networks also falls under the rubric of multi-mechanism learning.
Learning by non-interfering feedback chemical signaling in physical networks
Mar 22, 2022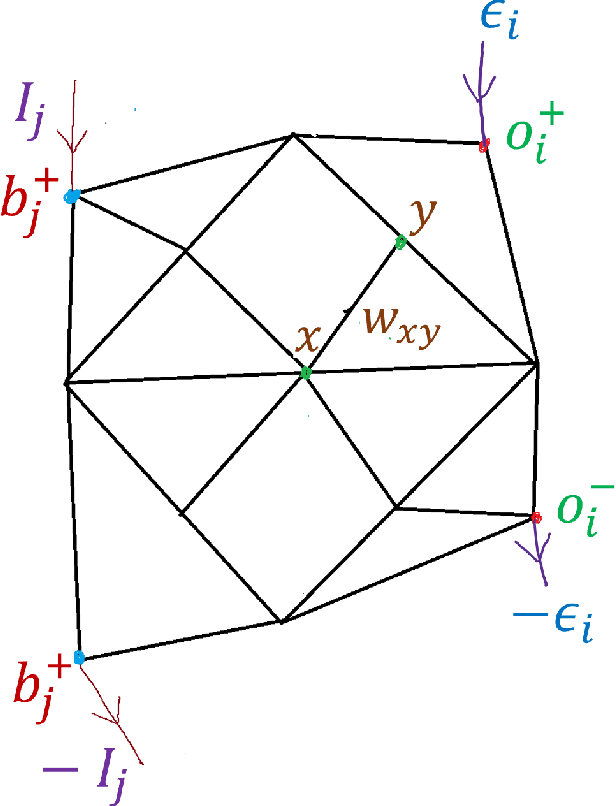
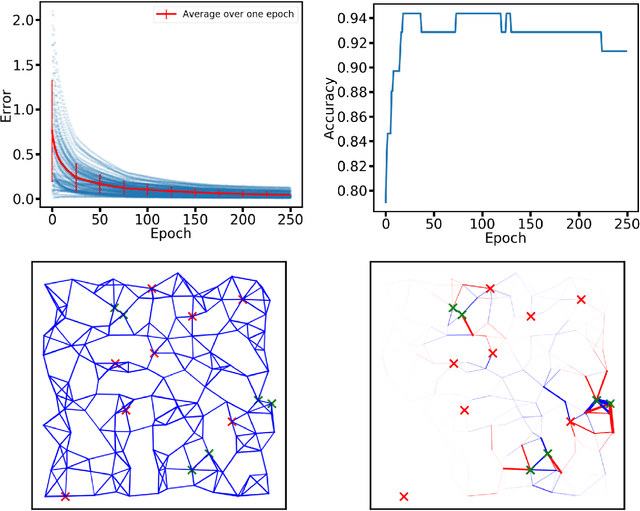
Abstract:Both non-neural and neural biological systems can learn. So rather than focusing on purely brain-like learning, efforts are underway to study learning in physical systems. Such efforts include equilibrium propagation (EP) and coupled learning (CL), which require storage of two different states-the free state and the perturbed state-during the learning process to retain information about gradients. Inspired by slime mold, we propose a new learning algorithm rooted in chemical signaling that does not require storage of two different states. Rather, the output error information is encoded in a chemical signal that diffuses into the network in a similar way as the activation/feedforward signal. The steady state feedback chemical concentration, along with the activation signal, stores the required gradient information locally. We apply our algorithm using a physical, linear flow network and test it using the Iris data set with 93% accuracy. We also prove that our algorithm performs gradient descent. Finally, in addition to comparing our algorithm directly with EP and CL, we address the biological plausibility of the algorithm.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge