Atabey Kaygun
Classification of Stochastic Processes with Topological Data Analysis
Jun 08, 2022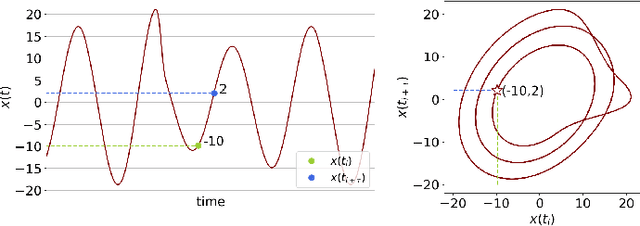
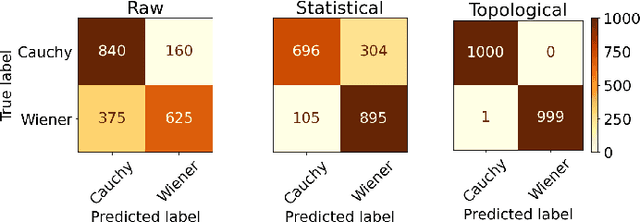
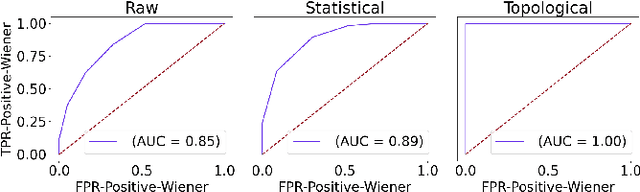
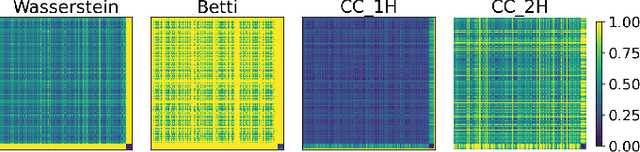
Abstract:In this study, we examine if engineered topological features can distinguish time series sampled from different stochastic processes with different noise characteristics, in both balanced and unbalanced sampling schemes. We compare our classification results against the results of the same classification tasks built on statistical and raw features. We conclude that in classification tasks of time series, different machine learning models built on engineered topological features perform consistently better than those built on standard statistical and raw features.
Time Series Classification via Topological Data Analysis
Feb 03, 2021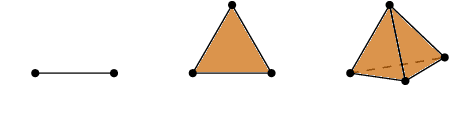
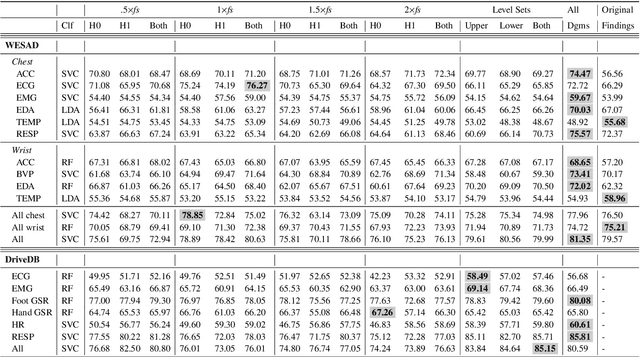
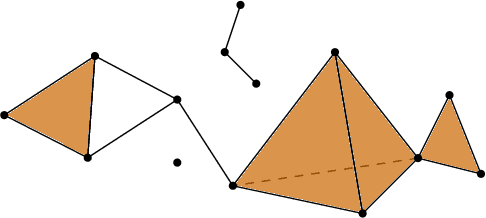
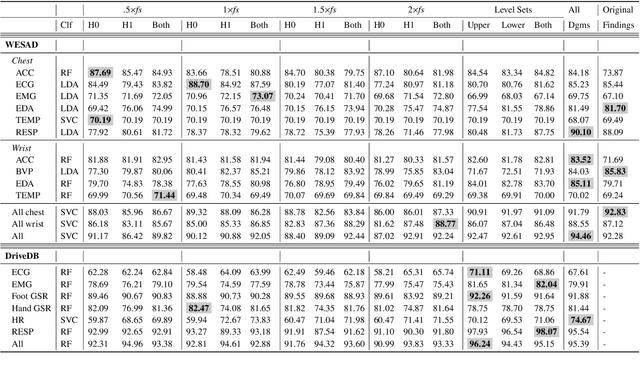
Abstract:In this paper, we develop topological data analysis methods for classification tasks on univariate time series. As an application we perform binary and ternary classification tasks on two public datasets that consist of physiological signals collected under stress and non-stress conditions. We accomplish our goal by using persistent homology to engineer stable topological features after we use a time delay embedding of the signals and perform a subwindowing instead of using windows of fixed length. The combination of methods we use can be applied to any univariate time series and in this application allows us to reduce noise and use long window sizes without incurring an extra computational cost. We then use machine learning models on the features we algorithmically engineered to obtain higher accuracies with fewer features.
Hierarchical Clustering and Zeroth Persistent Homology
Dec 04, 2020
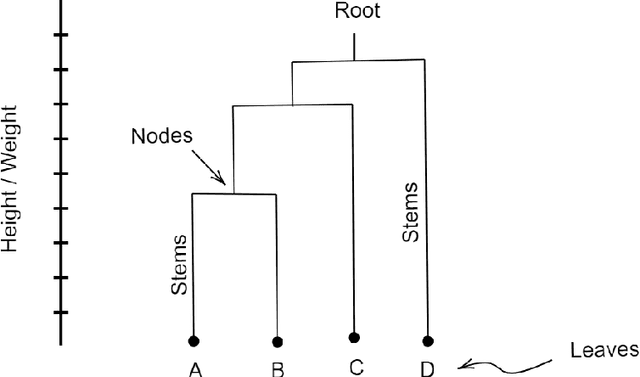
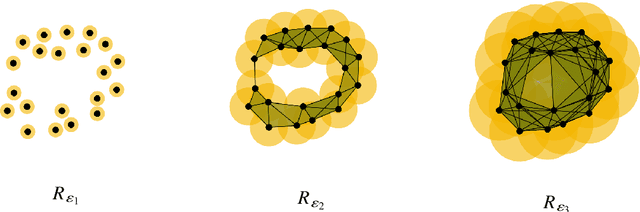
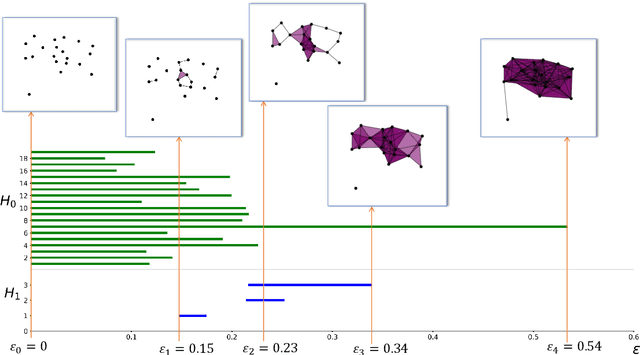
Abstract:In this article, we show that hierarchical clustering and the zeroth persistent homology do deliver the same topological information about a given data set. We show this fact using cophenetic matrices constructed out of the filtered Vietoris-Rips complex of the data set at hand. As in any cophenetic matrix, one can also display the inter-relations of zeroth homology classes via a rooted tree, also known as a dendogram. Since homological cophenetic matrices can be calculated for higher homologies, one can also sketch similar dendograms for higher persistent homology classes.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge