Arvind Seshan
A Neural Network Based Automated IFT-20 Sensory Neuron Classifier for Caenorhabditis elegans
Oct 24, 2022Abstract:Determining neuronal identity in imaging data is an essential task in neuroscience, facilitating the comparison of neural activity across organisms. Cross-organism comparison, in turn, enables a wide variety of research including whole-brain analysis of functional networks and linking the activity of specific neurons to behavior or environmental stimuli. The recent development of three-dimensional, pan-neuronal imaging with single-cell resolution within Caenorhabditis elegans has brought neuron identification, tracking, and activity monitoring all within reach. The nematode C. elegans is often used as a model organism to study neuronal activity due to factors such as its transparency and well-understood nervous system. The principal barrier to high-accuracy neuron identification is that in adult C. elegans, the position of neuronal cell bodies is not stereotyped. Existing approaches to address this issue use genetically encoded markers as an additional identifying feature. For example, the NeuroPAL strain uses multicolored fluorescent reporters. However, this approach has limited use due to the negative effects of excessive genetic modification. In this study, I propose an alternative neuronal identification technique using only single-color fluorescent images. I designed a novel neural network based classifier that automatically labels sensory neurons using an iterative, landmark-based neuron identification process inspired by the manual annotation procedures that humans employ. This design labels sensory neurons in C. elegans with 91.61% accuracy.
Using Machine Learning to Augment Dynamic Time Warping Based Signal Classification
Jun 14, 2022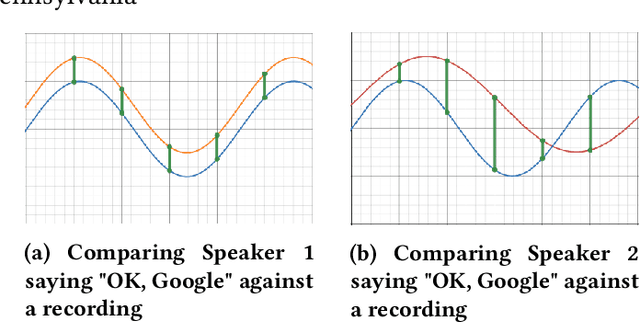
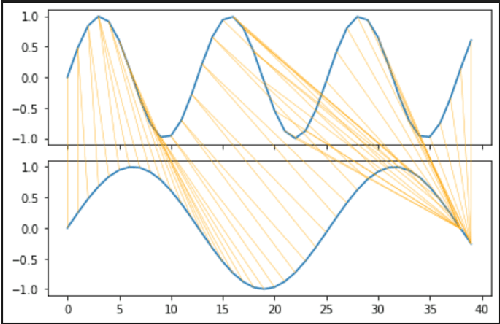
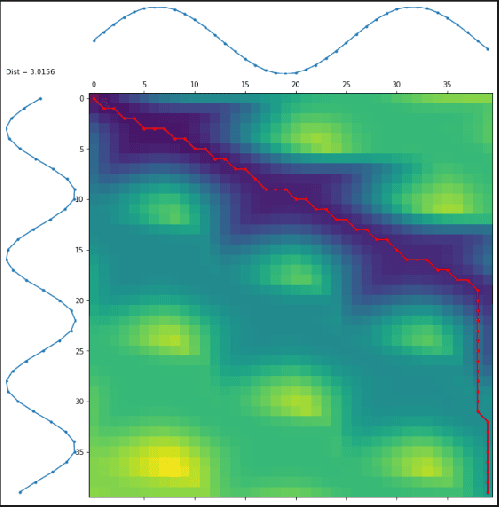
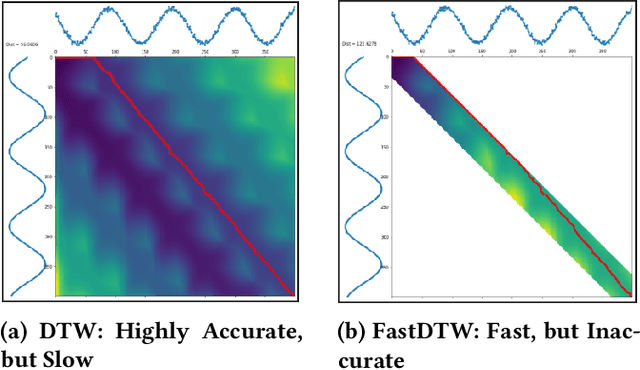
Abstract:Modern applications such as voice recognition rely on the ability to compare signals to pre-recorded ones to classify them. However, this comparison typically needs to ignore differences due to signal noise, temporal offset, signal magnitude, and other external factors. The Dynamic Time Warping (DTW) algorithm quantifies this similarity by finding corresponding regions between the signals and non-linearly warping one signal by stretching and shrinking it. Unfortunately, searching through all "warps" of a signal to find the best corresponding regions is computationally expensive. The FastDTW algorithm improves performance, but sacrifices accuracy by only considering small signal warps. My goal is to improve the speed of DTW while maintaining high accuracy. My key insight is that in any particular application domain, signals exhibit specific types of variation. For example, the accelerometer signal measured for two different people would differ based on their stride length and weight. My system, called Machine Learning DTW (MLDTW), uses machine learning to learn the types of warps that are common in a particular domain. It then uses the learned model to improve DTW performance by limiting the search of potential warps appropriately. My results show that compared to FastDTW, MLDTW is at least as fast and reduces errors by 60% on average across four different data sets. These improvements will significantly impact a wide variety of applications (e.g. health monitoring) and enable more scalable processing of multivariate, higher frequency, and longer signal recordings.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge