Anindo Saha
Leveraging Adaptive Color Augmentation in Convolutional Neural Networks for Deep Skin Lesion Segmentation
Oct 31, 2020
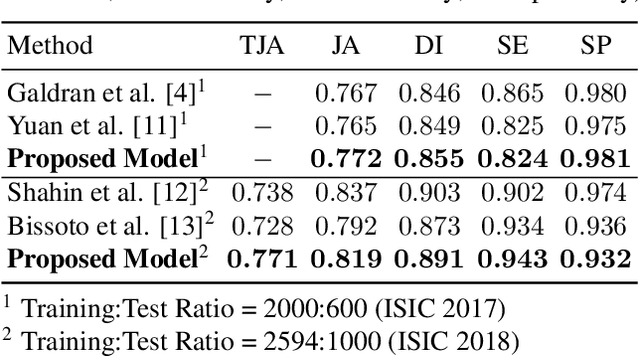
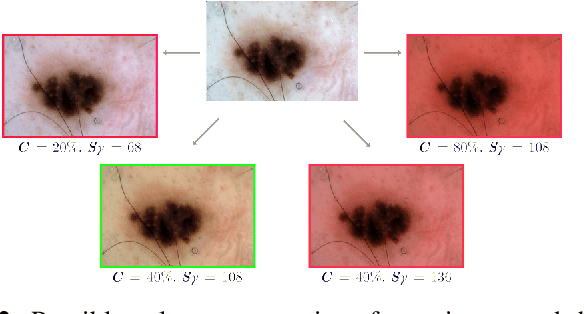
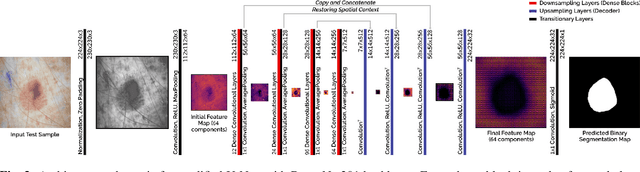
Abstract:Fully automatic detection of skin lesions in dermatoscopic images can facilitate early diagnosis and repression of malignant melanoma and non-melanoma skin cancer. Although convolutional neural networks are a powerful solution, they are limited by the illumination spectrum of annotated dermatoscopic screening images, where color is an important discriminative feature. In this paper, we propose an adaptive color augmentation technique to amplify data expression and model performance, while regulating color difference and saturation to minimize the risks of using synthetic data. Through deep visualization, we qualitatively identify and verify the semantic structural features learned by the network for discriminating skin lesions against normal skin tissue. The overall system achieves a Dice Ratio of 0.891 with 0.943 sensitivity and 0.932 specificity on the ISIC 2018 Testing Set for segmentation.
Leveraging SLIC Superpixel Segmentation and Cascaded Ensemble SVM for Fully Automated Mass Detection In Mammograms
Oct 20, 2020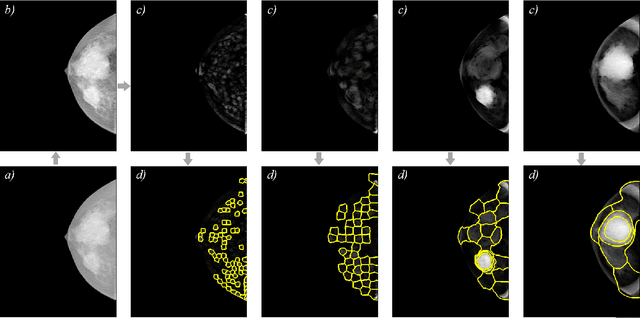
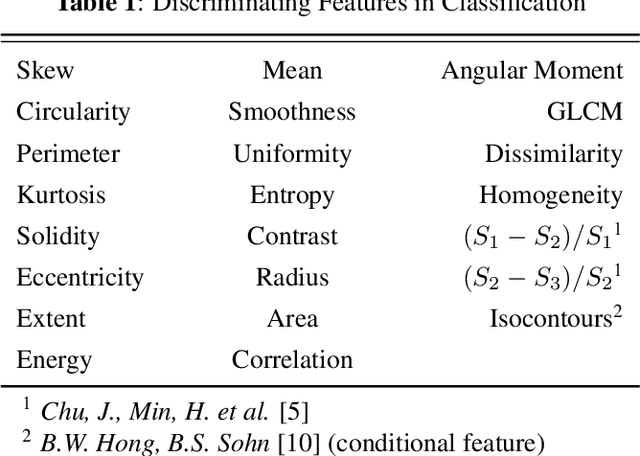

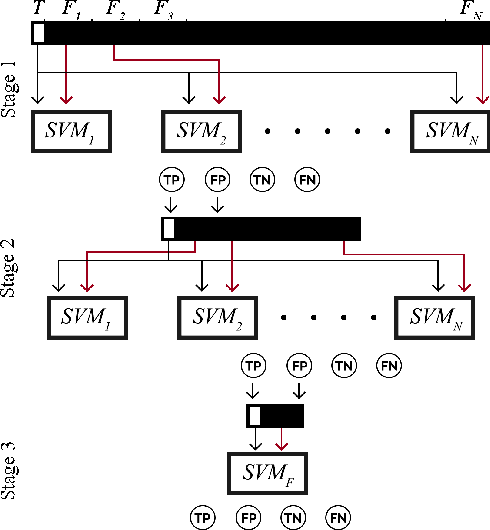
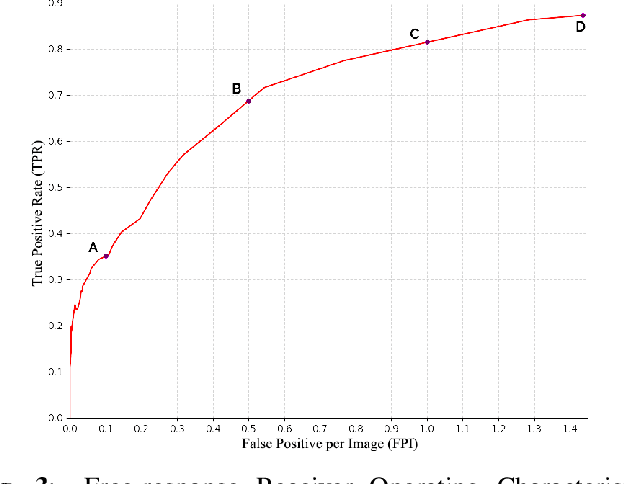
Abstract:Identification and segmentation of breast masses in mammograms face complex challenges, owing to the highly variable nature of malignant densities with regards to their shape, contours, texture and orientation. Additionally, classifiers typically suffer from high class imbalance in region candidates, where normal tissue regions vastly outnumber malignant masses. This paper proposes a rigorous segmentation method, supported by morphological enhancement using grayscale linear filters. A novel cascaded ensemble of support vector machines (SVM) is used to effectively tackle the class imbalance and provide significant predictions. For True Positive Rate (TPR) of 0.35, 0.69 and 0.82, the system generates only 0.1, 0.5 and 1.0 False Positives/Image (FPI), respectively.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge