Ali Punjani
Building Proteins in a Day: Efficient 3D Molecular Reconstruction
Apr 14, 2015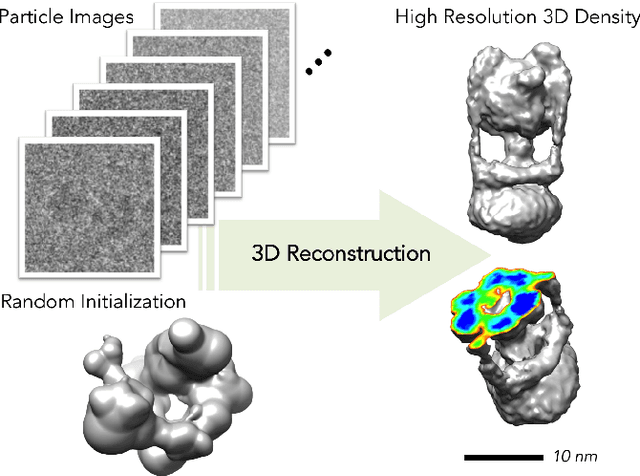
Abstract:Discovering the 3D atomic structure of molecules such as proteins and viruses is a fundamental research problem in biology and medicine. Electron Cryomicroscopy (Cryo-EM) is a promising vision-based technique for structure estimation which attempts to reconstruct 3D structures from 2D images. This paper addresses the challenging problem of 3D reconstruction from 2D Cryo-EM images. A new framework for estimation is introduced which relies on modern stochastic optimization techniques to scale to large datasets. We also introduce a novel technique which reduces the cost of evaluating the objective function during optimization by over five orders or magnitude. The net result is an approach capable of estimating 3D molecular structure from large scale datasets in about a day on a single workstation.
Microscopic Advances with Large-Scale Learning: Stochastic Optimization for Cryo-EM
Jan 27, 2015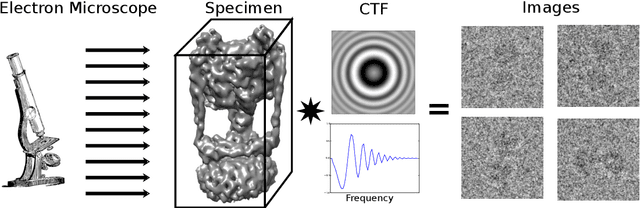
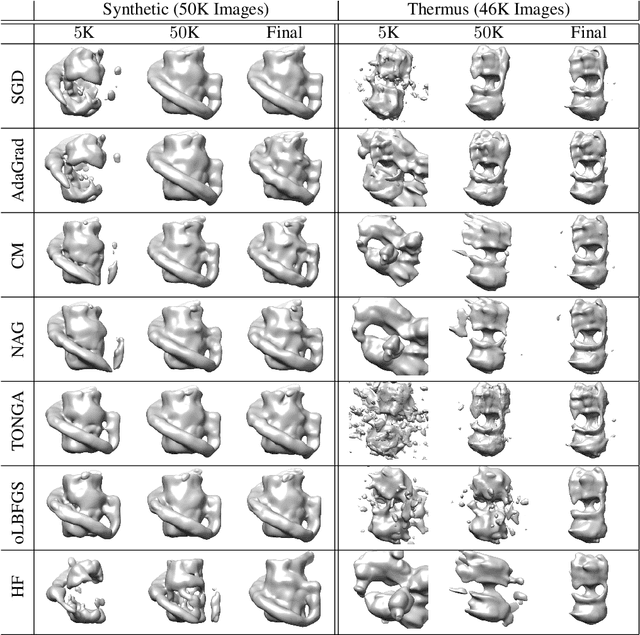

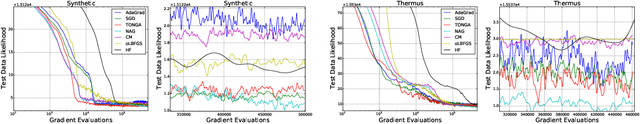

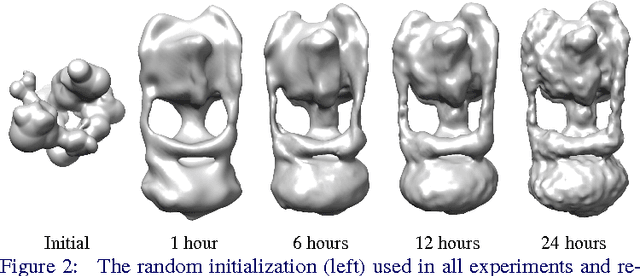
Abstract:Determining the 3D structures of biological molecules is a key problem for both biology and medicine. Electron Cryomicroscopy (Cryo-EM) is a promising technique for structure estimation which relies heavily on computational methods to reconstruct 3D structures from 2D images. This paper introduces the challenging Cryo-EM density estimation problem as a novel application for stochastic optimization techniques. Structure discovery is formulated as MAP estimation in a probabilistic latent-variable model, resulting in an optimization problem to which an array of seven stochastic optimization methods are applied. The methods are tested on both real and synthetic data, with some methods recovering reasonable structures in less than one epoch from a random initialization. Complex quasi-Newton methods are found to converge more slowly than simple gradient-based methods, but all stochastic methods are found to converge to similar optima. This method represents a major improvement over existing methods as it is significantly faster and is able to converge from a random initialization.
Fast Exact Search in Hamming Space with Multi-Index Hashing
Apr 25, 2014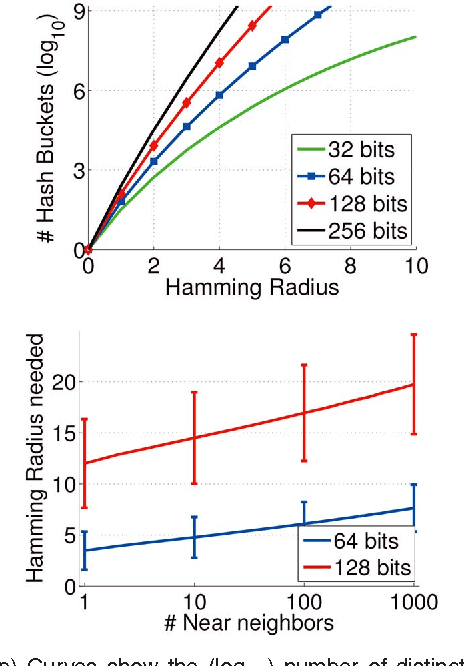
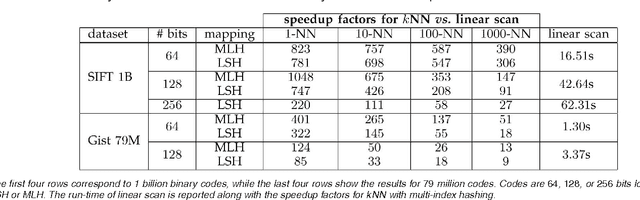
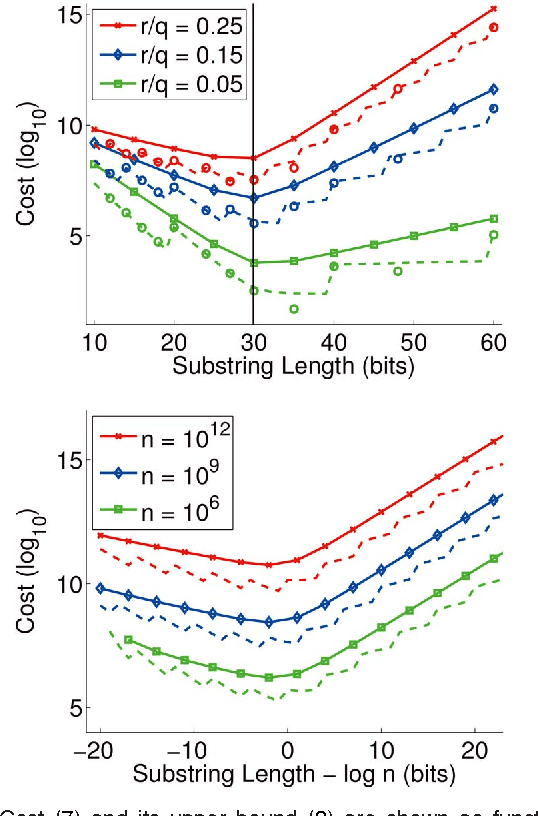
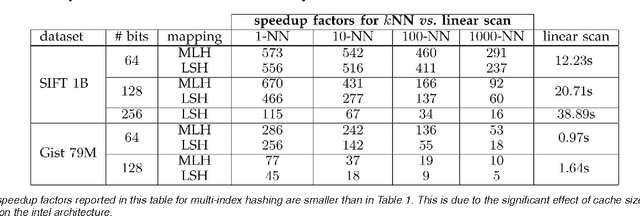
Abstract:There is growing interest in representing image data and feature descriptors using compact binary codes for fast near neighbor search. Although binary codes are motivated by their use as direct indices (addresses) into a hash table, codes longer than 32 bits are not being used as such, as it was thought to be ineffective. We introduce a rigorous way to build multiple hash tables on binary code substrings that enables exact k-nearest neighbor search in Hamming space. The approach is storage efficient and straightforward to implement. Theoretical analysis shows that the algorithm exhibits sub-linear run-time behavior for uniformly distributed codes. Empirical results show dramatic speedups over a linear scan baseline for datasets of up to one billion codes of 64, 128, or 256 bits.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge