Ali Mammadov
Tribvn Healthcare
Self-Supervision Enhances Instance-based Multiple Instance Learning Methods in Digital Pathology: A Benchmark Study
May 02, 2025Abstract:Multiple Instance Learning (MIL) has emerged as the best solution for Whole Slide Image (WSI) classification. It consists of dividing each slide into patches, which are treated as a bag of instances labeled with a global label. MIL includes two main approaches: instance-based and embedding-based. In the former, each patch is classified independently, and then the patch scores are aggregated to predict the bag label. In the latter, bag classification is performed after aggregating patch embeddings. Even if instance-based methods are naturally more interpretable, embedding-based MILs have usually been preferred in the past due to their robustness to poor feature extractors. However, recently, the quality of feature embeddings has drastically increased using self-supervised learning (SSL). Nevertheless, many authors continue to endorse the superiority of embedding-based MIL. To investigate this further, we conduct 710 experiments across 4 datasets, comparing 10 MIL strategies, 6 self-supervised methods with 4 backbones, 4 foundation models, and various pathology-adapted techniques. Furthermore, we introduce 4 instance-based MIL methods never used before in the pathology domain. Through these extensive experiments, we show that with a good SSL feature extractor, simple instance-based MILs, with very few parameters, obtain similar or better performance than complex, state-of-the-art (SOTA) embedding-based MIL methods, setting new SOTA results on the BRACS and Camelyon16 datasets. Since simple instance-based MIL methods are naturally more interpretable and explainable to clinicians, our results suggest that more effort should be put into well-adapted SSL methods for WSI rather than into complex embedding-based MIL methods.
Robust Mitosis Detection Using a Cascade Mask-RCNN Approach With Domain-Specific Residual Cycle-GAN Data Augmentation
Sep 28, 2021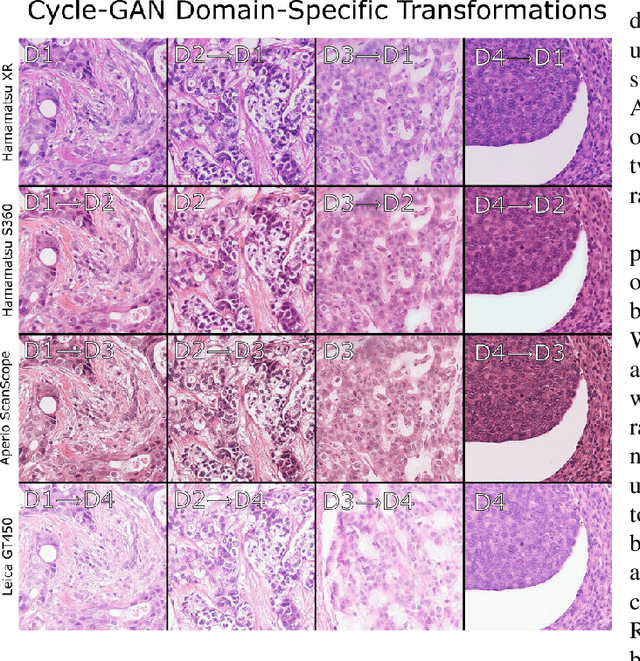
Abstract:For the MIDOG mitosis detection challenge, we created a cascade algorithm consisting of a Mask-RCNN detector, followed by a classification ensemble consisting of ResNet50 and DenseNet201 to refine detected mitotic candidates. The MIDOG training data consists of 200 frames originating from four scanners, three of which are annotated for mitotic instances with centroid annotations. Our main algorithmic choices are as follows: first, to enhance the generalizability of our detector and classification networks, we use a state-of-the-art residual Cycle-GAN to transform each scanner domain to every other scanner domain. During training, we then randomly load, for each image, one of the four domains. In this way, our networks can learn from the fourth non-annotated scanner domain even if we don't have annotations for it. Second, for training the detector network, rather than using centroid-based fixed-size bounding boxes, we create mitosis-specific bounding boxes. We do this by manually annotating a small selection of mitoses, training a Mask-RCNN on this small dataset, and applying it to the rest of the data to obtain full annotations. We trained the follow-up classification ensemble using only the challenge-provided positive and hard-negative examples. On the preliminary test set, the algorithm scores an F1 score of 0.7578, putting us as the second-place team on the leaderboard.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge