Ali K. Z. Tehrani
Constrained and Regularized Quantitative Ultrasound Parameter Estimation using ADMM
Jan 07, 2025



Abstract:Regularized estimation of quantitative ultrasound (QUS) parameters, such as attenuation and backscatter coefficients, has gained research interest. Recently, the alternating direction method of multipliers (ADMM) has been applied successfully to estimate these parameters, by utilizing L2 and L1 norms for attenuation and backscatter coefficient regularization, respectively. While this method improves upon previous approaches, it does not fully leverage the prior knowledge of minimum physically feasible parameter values, sometimes yielding values outside the realistic range. This work addresses this limitation by incorporating minimum QUS parameter values as constraints to enhance ADMM estimation. The proposed method is validated using experimental phantom data.
Uncertainty Decomposition and Error Margin Detection of Homodyned-K Distribution in Quantitative Ultrasound
Sep 17, 2024Abstract:Homodyned K-distribution (HK-distribution) parameter estimation in quantitative ultrasound (QUS) has been recently addressed using Bayesian Neural Networks (BNNs). BNNs have been shown to significantly reduce computational time in speckle statistics-based QUS without compromising accuracy and precision. Additionally, they provide estimates of feature uncertainty, which can guide the clinician's trust in the reported feature value. The total predictive uncertainty in Bayesian modeling can be decomposed into epistemic (uncertainty over the model parameters) and aleatoric (uncertainty inherent in the data) components. By decomposing the predictive uncertainty, we can gain insights into the factors contributing to the total uncertainty. In this study, we propose a method to compute epistemic and aleatoric uncertainties for HK-distribution parameters ($\alpha$ and $k$) estimated by a BNN, in both simulation and experimental data. In addition, we investigate the relationship between the prediction error and both uncertainties, shedding light on the interplay between these uncertainties and HK parameters errors.
Homodyned K-Distribution Parameter Estimation in Quantitative Ultrasound: Autoencoder and Bayesian Neural Network Approaches
Jan 19, 2024



Abstract:Quantitative ultrasound (QUS) analyzes the ultrasound backscattered data to find the properties of scatterers that correlate with the tissue microstructure. Statistics of the envelope of the backscattered radiofrequency (RF) data can be utilized to estimate several QUS parameters. Different distributions have been proposed to model envelope data. The homodyned K-distribution (HK distribution) is one of the most comprehensive distributions that can model ultrasound backscattered envelope data under diverse scattering conditions (varying scatterer number density and coherent scattering). The scatterer clustering parameter (alpha) and the ratio of the coherent to diffuse scattering power (k) are the parameters of this distribution that have been used extensively for tissue characterization in diagnostic ultrasound. The estimation of these two parameters (which we refer to as HK parameters) is done using optimization algorithms in which statistical features such as the envelope point-wise signalto-noise ratio (SNR), skewness, kurtosis, and the log-based moments have been utilized as input to such algorithms. The optimization methods minimize the difference between features and their theoretical value from the HK model. We propose that the true value of these statistical features is a hyperplane that covers a small portion of the feature space. In this paper, we follow two approaches to reduce the effect of sample features' error. We propose a model projection neural network based on denoising autoencoders to project the noisy features into this space based on this assumption. We also investigate if the noise distribution can be learned by the deep estimators. We compare the proposed methods with conventional methods using simulations, an experimental phantom, and data from an in vivo animal model of hepatic steatosis. A demo code are available online at http://code.sonography.ai
Exploiting Mechanics-Based Priors for Lateral Displacement Estimation in Ultrasound Elastography
May 31, 2023



Abstract:Tracking the displacement between the pre- and post-deformed radio-frequency (RF) frames is a pivotal step of ultrasound elastography, which depicts tissue mechanical properties to identify pathologies. Due to ultrasound's poor ability to capture information pertaining to the lateral direction, the existing displacement estimation techniques fail to generate an accurate lateral displacement or strain map. The attempts made in the literature to mitigate this well-known issue suffer from one of the following limitations: 1) Sampling size is substantially increased, rendering the method computationally and memory expensive. 2) The lateral displacement estimation entirely depends on the axial one, ignoring data fidelity and creating large errors. This paper proposes exploiting the effective Poisson's ratio (EPR)-based mechanical correspondence between the axial and lateral strains along with the RF data fidelity and displacement continuity to improve the lateral displacement and strain estimation accuracies. We call our techniques MechSOUL (Mechanically-constrained Second-Order Ultrasound eLastography) and L1-MechSOUL (L1-norm-based MechSOUL), which optimize L2- and L1-norm-based penalty functions, respectively. Extensive validation experiments with simulated, phantom, and in vivo datasets demonstrate that MechSOUL and L1-MechSOUL's lateral strain and EPR estimation abilities are substantially superior to those of the recently-published elastography techniques. We have published the MATLAB codes of MechSOUL and L1-MechSOUL at http://code.sonography.ai.
A deep learning approach for patchless estimation of ultrasound quantitative parametric image with uncertainty measurement
Feb 24, 2023Abstract:Quantitative ultrasound (QUS) aims to find properties of scatterers which are related to the tissue microstructure. Among different QUS parameters, scatterer number density has been found to be a reliable biomarker for detecting different abnormalities. The homodyned K-distribution (HK-distribution) is a model for the probability density function of the ultrasound echo amplitude that can model different scattering scenarios but requires a large number of samples to be estimated reliably. Parametric images of HK-distribution parameters can be formed by dividing the envelope data into small overlapping patches and estimating parameters within the patches independently. This approach imposes two limiting constraints, the HK-distribution parameters are assumed to be constant within each patch, and each patch requires enough independent samples. In order to mitigate those problems, we employ a deep learning approach to estimate parametric images of scatterer number density (related to HK-distribution shape parameter) without patching. Furthermore, an uncertainty map of the network's prediction is quantified to provide insight about the confidence of the network about the estimated HK parameter values.
Lateral Strain Imaging using Self-supervised and Physically Inspired Constraints in Unsupervised Regularized Elastography
Dec 16, 2022



Abstract:Convolutional Neural Networks (CNN) have shown promising results for displacement estimation in UltraSound Elastography (USE). Many modifications have been proposed to improve the displacement estimation of CNNs for USE in the axial direction. However, the lateral strain, which is essential in several downstream tasks such as the inverse problem of elasticity imaging, remains a challenge. The lateral strain estimation is complicated since the motion and the sampling frequency in this direction are substantially lower than the axial one, and a lack of carrier signal in this direction. In computer vision applications, the axial and the lateral motions are independent. In contrast, the tissue motion pattern in USE is governed by laws of physics which link the axial and lateral displacements. In this paper, inspired by Hooke's law, we first propose Physically Inspired ConsTraint for Unsupervised Regularized Elastography (PICTURE), where we impose a constraint on the Effective Poisson's ratio (EPR) to improve the lateral strain estimation. In the next step, we propose self-supervised PICTURE (sPICTURE) to further enhance the strain image estimation. Extensive experiments on simulation, experimental phantom and in vivo data demonstrate that the proposed methods estimate accurate axial and lateral strain maps.
Homodyned K-distribution: parameter estimation and uncertainty quantification using Bayesian neural networks
Oct 31, 2022Abstract:Quantitative ultrasound (QUS) allows estimating the intrinsic tissue properties. Speckle statistics are the QUS parameters that describe the first order statistics of ultrasound (US) envelope data. The parameters of Homodyned K-distribution (HK-distribution) are the speckle statistics that can model the envelope data in diverse scattering conditions. However, they require a large amount of data to be estimated reliably. Consequently, finding out the intrinsic uncertainty of the estimated parameters can help us to have a better understanding of the estimated parameters. In this paper, we propose a Bayesian Neural Network (BNN) to estimate the parameters of HK-distribution and quantify the uncertainty of the estimator.
Infusing known operators in convolutional neural networks for lateral strain imaging in ultrasound elastography
Oct 31, 2022Abstract:Convolutional Neural Networks (CNN) have been employed for displacement estimation in ultrasound elastography (USE). High-quality axial strains (derivative of the axial displacement in the axial direction) can be estimated by the proposed networks. In contrast to axial strain, lateral strain, which is highly required in Poisson's ratio imaging and elasticity reconstruction, has a poor quality. The main causes include low sampling frequency, limited motion, and lack of phase information in the lateral direction. Recently, physically inspired constraint in unsupervised regularized elastography (PICTURE) has been proposed. This method took into account the range of the feasible lateral strain defined by the rules of physics of motion and employed a regularization strategy to improve the lateral strains. Despite the substantial improvement, the regularization was only applied during the training; hence it did not guarantee during the test that the lateral strain is within the feasible range. Furthermore, only the feasible range was employed, other constraints such as incompressibility were not investigated. In this paper, we address these two issues and propose kPICTURE in which two iterative algorithms were infused into the network architecture in the form of known operators to ensure the lateral strain is within the feasible range and impose incompressibility during the test phase.
Deep Estimation of Speckle Statistics Parametric Images
Jun 08, 2022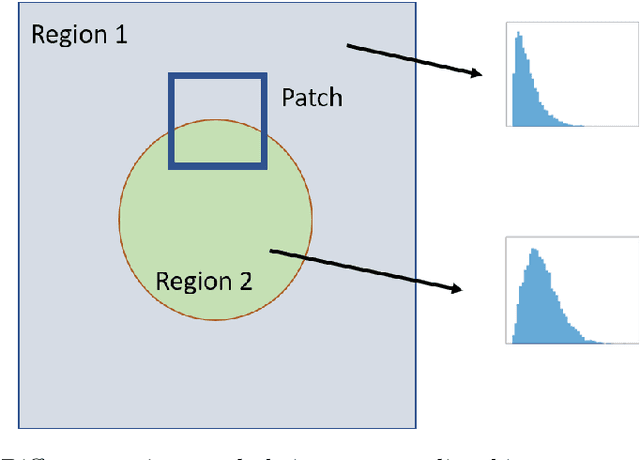

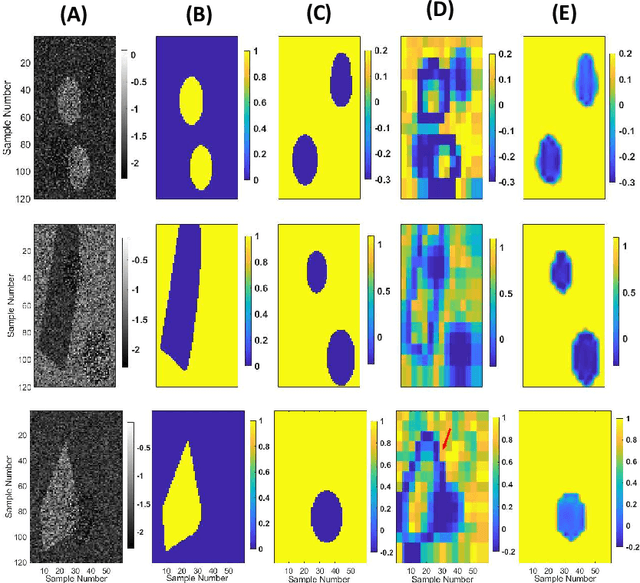

Abstract:Quantitative Ultrasound (QUS) provides important information about the tissue properties. QUS parametric image can be formed by dividing the envelope data into small overlapping patches and computing different speckle statistics such as parameters of the Nakagami and Homodyned K-distributions (HK-distribution). The calculated QUS parametric images can be erroneous since only a few independent samples are available inside the patches. Another challenge is that the envelope samples inside the patch are assumed to come from the same distribution, an assumption that is often violated given that the tissue is usually not homogenous. In this paper, we propose a method based on Convolutional Neural Networks (CNN) to estimate QUS parametric images without patching. We construct a large dataset sampled from the HK-distribution, having regions with random shapes and QUS parameter values. We then use a well-known network to estimate QUS parameters in a multi-task learning fashion. Our results confirm that the proposed method is able to reduce errors and improve border definition in QUS parametric images.
Physically Inspired Constraint for Unsupervised Regularized Ultrasound Elastography
Jun 05, 2022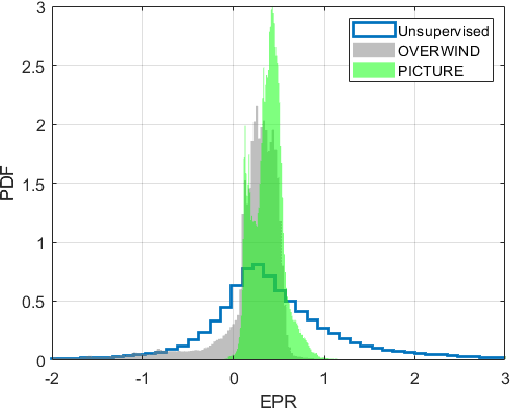
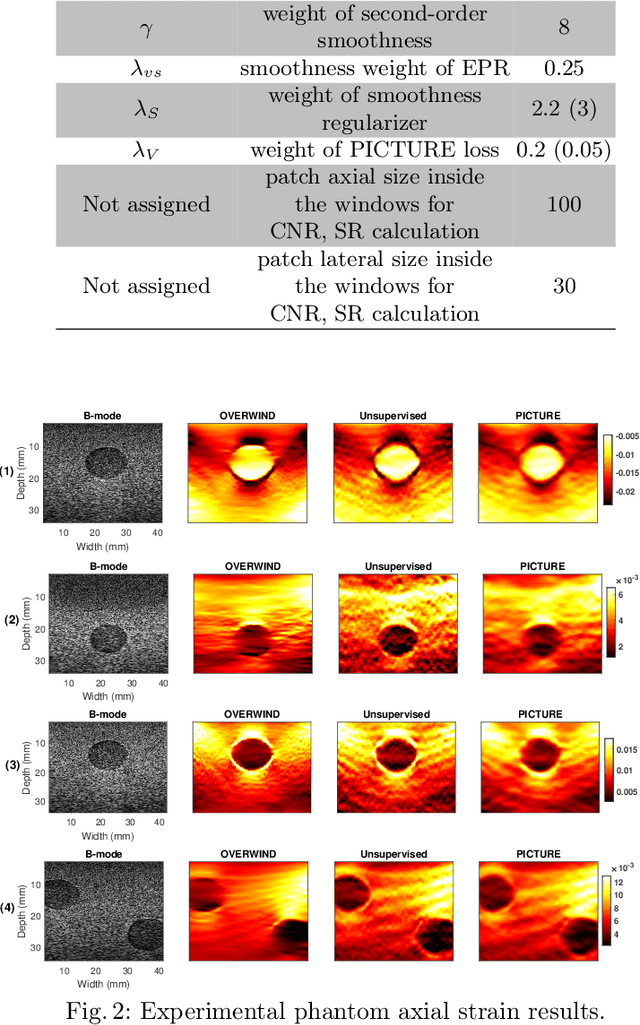
Abstract:Displacement estimation is a critical step of virtually all Ultrasound Elastography (USE) techniques. Two main features make this task unique compared to the general optical flow problem: the high-frequency nature of ultrasound radio-frequency (RF) data and the governing laws of physics on the displacement field. Recently, the architecture of the optical flow networks has been modified to be able to use RF data. Also, semi-supervised and unsupervised techniques have been employed for USE by considering prior knowledge of displacement continuity in the form of the first- and second-derivative regularizers. Despite these attempts, no work has considered the tissue compression pattern, and displacements in axial and lateral directions have been assumed to be independent. However, tissue motion pattern is governed by laws of physics in USE, rendering the axial and the lateral displacements highly correlated. In this paper, we propose Physically Inspired ConsTraint for Unsupervised Regularized Elastography (PICTURE), where we impose constraints on the Poisson's ratio to improve lateral displacement estimates. Experiments on phantom and in vivo data show that PICTURE substantially improves the quality of the lateral displacement estimation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge