Alberto Calderone
PubSqueezer: A Text-Mining Web Tool to Transform Unstructured Documents into Structured Data
Nov 09, 2020
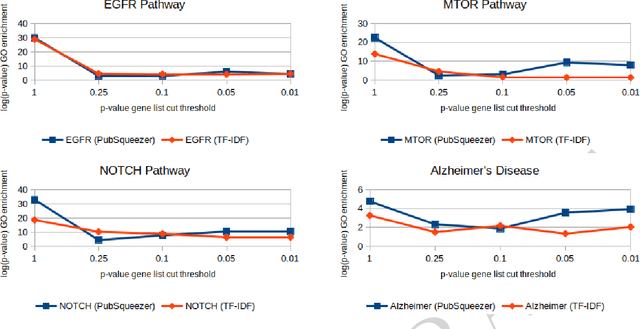
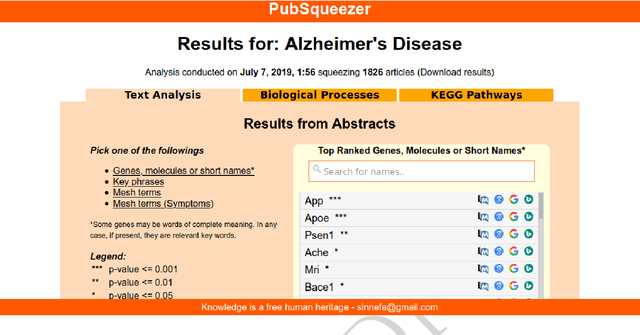
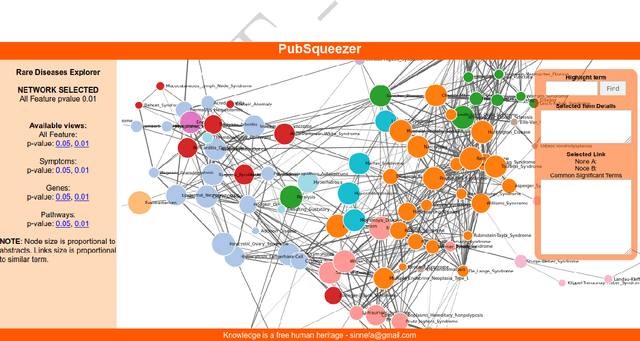
Abstract:The amount of scientific papers published every day is daunting and constantly increasing. Keeping up with literature represents a challenge. If one wants to start exploring new topics it is hard to have a big picture without reading lots of articles. Furthermore, as one reads through literature, making mental connections is crucial to ask new questions which might lead to discoveries. In this work, I present a web tool which uses a Text Mining strategy to transform large collections of unstructured biomedical articles into structured data. Generated results give a quick overview on complex topics which can possibly suggest not explicitly reported information. In particular, I show two Data Science analyses. First, I present a literature based rare diseases network build using this tool in the hope that it will help clarify some aspects of these less popular pathologies. Secondly, I show how a literature based analysis conducted with PubSqueezer results allows to describe known facts about SARS-CoV-2. In one sentence, data generated with PubSqueezer make it easy to use scientific literate in any computational analysis such as machine learning, natural language processing etc. Availability: http://www.pubsqueezer.com
A Computational Analysis of Natural Languages to Build a Sentence Structure Aware Artificial Neural Network
Jun 13, 2019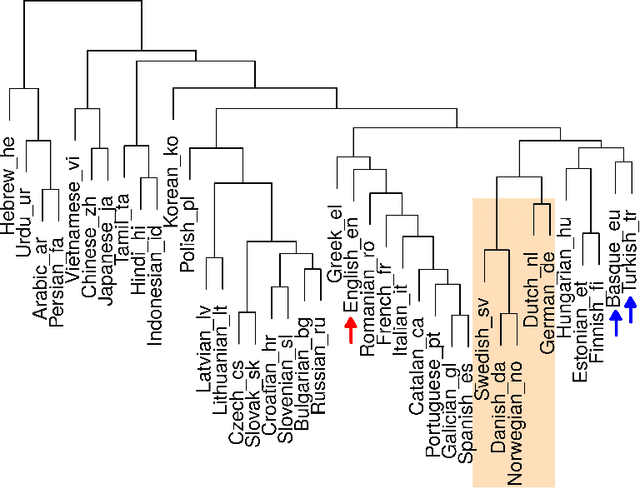
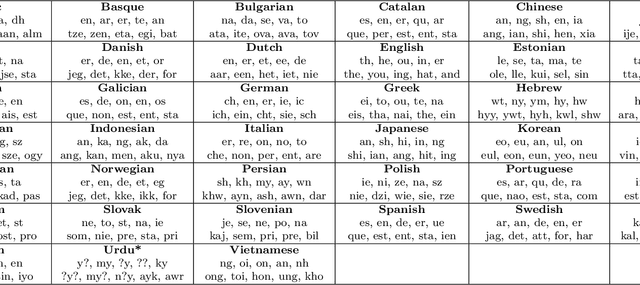
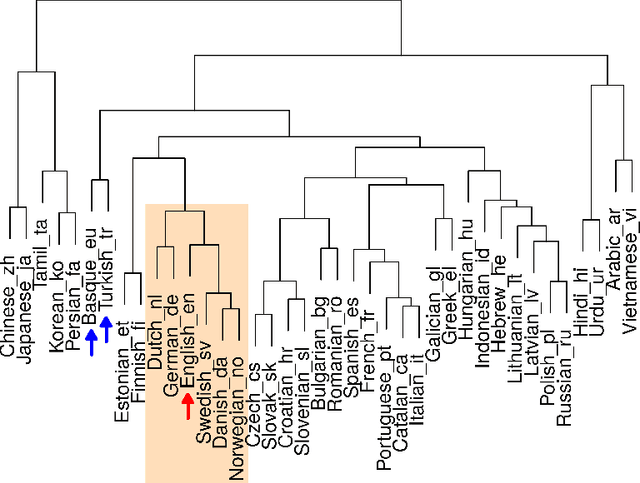

Abstract:Natural languages are complexly structured entities. They exhibit characterising regularities that can be exploited to link them one another. In this work, I compare two morphological aspects of languages: Written Patterns and Sentence Structure. I show how languages spontaneously group by similarity in both analyses and derive an average language distance. Finally, exploiting Sentence Structure I developed an Artificial Neural Network capable of distinguishing languages suggesting that not only word roots but also grammatical sentence structure is a characterising trait which alone suffice to identify them.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge