Adam Howes
Measuring Mid-2025 LLM-Assistance on Novice Performance in Biology
Feb 18, 2026Abstract:Large language models (LLMs) perform strongly on biological benchmarks, raising concerns that they may help novice actors acquire dual-use laboratory skills. Yet, whether this translates to improved human performance in the physical laboratory remains unclear. To address this, we conducted a pre-registered, investigator-blinded, randomized controlled trial (June-August 2025; n = 153) evaluating whether LLMs improve novice performance in tasks that collectively model a viral reverse genetics workflow. We observed no significant difference in the primary endpoint of workflow completion (5.2% LLM vs. 6.6% Internet; P = 0.759), nor in the success rate of individual tasks. However, the LLM arm had numerically higher success rates in four of the five tasks, most notably for the cell culture task (68.8% LLM vs. 55.3% Internet; P = 0.059). Post-hoc Bayesian modeling of pooled data estimates an approximate 1.4-fold increase (95% CrI 0.74-2.62) in success for a "typical" reverse genetics task under LLM assistance. Ordinal regression modelling suggests that participants in the LLM arm were more likely to progress through intermediate steps across all tasks (posterior probability of a positive effect: 81%-96%). Overall, mid-2025 LLMs did not substantially increase novice completion of complex laboratory procedures but were associated with a modest performance benefit. These results reveal a gap between in silico benchmarks and real-world utility, underscoring the need for physical-world validation of AI biosecurity assessments as model capabilities and user proficiency evolve.
Encoding spatiotemporal priors with VAEs for small-area estimation
Oct 20, 2021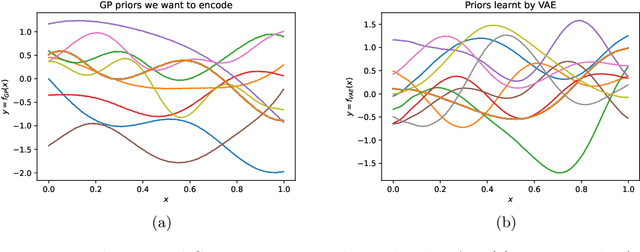
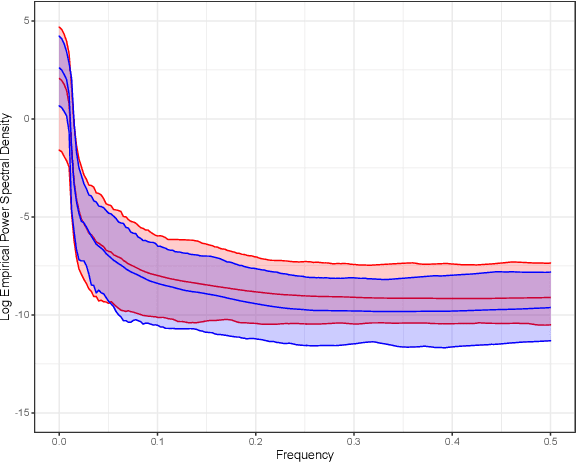
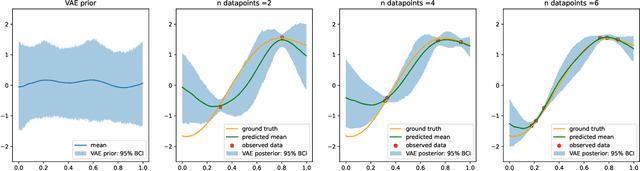
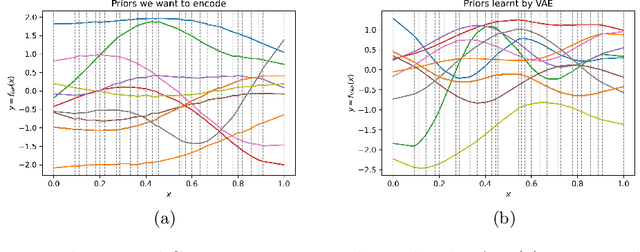
Abstract:Gaussian processes (GPs), implemented through multivariate Gaussian distributions for a finite collection of data, are the most popular approach in small-area spatiotemporal statistical modelling. In this context they are used to encode correlation structures over space and time and can generalise well in interpolation tasks. Despite their flexibility, off-the-shelf GPs present serious computational challenges which limit their scalability and practical usefulness in applied settings. Here, we propose a novel, deep generative modelling approach to tackle this challenge: for a particular spatiotemporal setting, we approximate a class of GP priors through prior sampling and subsequent fitting of a variational autoencoder (VAE). Given a trained VAE, the resultant decoder allows spatiotemporal inference to become incredibly efficient due to the low dimensional, independently distributed latent Gaussian space representation of the VAE. Once trained, inference using the VAE decoder replaces the GP within a Bayesian sampling framework. This approach provides tractable and easy-to-implement means of approximately encoding spatiotemporal priors and facilitates efficient statistical inference. We demonstrate the utility of our VAE two stage approach on Bayesian, small-area estimation tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge