Aaron Fenster
Virtual Fluoroscopy for Interventional Guidance using Magnetic Tracking
May 20, 2025Abstract:Purpose: In conventional fluoroscopy-guided interventions, the 2D projective nature of X-ray imaging limits depth perception and leads to prolonged radiation exposure. Virtual fluoroscopy, combined with spatially tracked surgical instruments, is a promising strategy to mitigate these limitations. While magnetic tracking shows unique advantages, particularly in tracking flexible instruments, it remains under-explored due to interference from ferromagnetic materials in the C-arm room. This work proposes a virtual fluoroscopy workflow by effectively integrating magnetic tracking, and demonstrates its clinical efficacy. Methods: An automatic virtual fluoroscopy workflow was developed using a radiolucent tabletop field generator prototype. Specifically, we developed a fluoro-CT registration approach with automatic 2D-3D shared landmark correspondence to establish the C-arm-patient relationship, along with a general C-arm modelling approach to calculate desired poses and generate corresponding virtual fluoroscopic images. Results: Testing on a dataset with views ranging from RAO 90 degrees to LAO 90 degrees, simulated fluoroscopic images showed visually imperceptible differences from the real ones, achieving a mean target projection distance error of 1.55 mm. An endoleak phantom insertion experiment highlighted the effectiveness of simulating multiplanar views with real-time instrument overlays, achieving a mean needle tip error of 3.42 mm. Conclusions: Results demonstrated the efficacy of virtual fluoroscopy integrated with magnetic tracking, improving depth perception during navigation. The broad capture range of virtual fluoroscopy showed promise in improving the users understanding of X-ray imaging principles, facilitating more efficient image acquisition.
Towards Seamless Integration of Magnetic Tracking into Fluoroscopy-guided Interventions
Nov 12, 2024



Abstract:The 2D projective nature of X-ray radiography presents significant limitations in fluoroscopy-guided interventions, particularly the loss of depth perception and prolonged radiation exposure. Integrating magnetic trackers into these workflows is promising; however, it remains challenging and under-explored in current research and practice. To address this, we employed a radiolucent magnetic field generator (FG) prototype as a foundational step towards seamless magnetic tracking (MT) integration. A two-layer FG mounting frame was designed for compatibility with various C-arm X-ray systems, ensuring smooth installation and optimal tracking accuracy. To overcome technical challenges, including accurate C-arm pose estimation, robust fluoro-CT registration, and 3D navigation, we proposed the incorporation of external aluminum fiducials without disrupting conventional workflows. Experimental evaluation showed no clinically significant impact of the aluminum fiducials and the C-arm on MT accuracy. Our fluoro-CT registration demonstrated high accuracy (mean projection distance approxiamtely 0.7 mm, robustness (wide capture range), and generalizability across local and public datasets. In a phantom targeting experiment, needle insertion error was between 2 mm and 3 mm, with real-time guidance using enhanced 2D and 3D navigation. Overall, our results demonstrated the efficacy and clinical applicability of the MT-assisted approach. To the best of our knowledge, this is the first study to integrate a radiolucent FG into a fluoroscopy-guided workflow.
Deep Regression 2D-3D Ultrasound Registration for Liver Motion Correction in Focal Tumor Thermal Ablation
Oct 03, 2024



Abstract:Liver tumor ablation procedures require accurate placement of the needle applicator at the tumor centroid. The lower-cost and real-time nature of ultrasound (US) has advantages over computed tomography (CT) for applicator guidance, however, in some patients, liver tumors may be occult on US and tumor mimics can make lesion identification challenging. Image registration techniques can aid in interpreting anatomical details and identifying tumors, but their clinical application has been hindered by the tradeoff between alignment accuracy and runtime performance, particularly when compensating for liver motion due to patient breathing or movement. Therefore, we propose a 2D-3D US registration approach to enable intra-procedural alignment that mitigates errors caused by liver motion. Specifically, our approach can correlate imbalanced 2D and 3D US image features and use continuous 6D rotation representations to enhance the model's training stability. The dataset was divided into 2388, 196 and 193 image pairs for training, validation and testing, respectively. Our approach achieved a mean Euclidean distance error of 2.28 mm $\pm$ 1.81 mm and a mean geodesic angular error of 2.99$^{\circ}$ $\pm$ 1.95$^{\circ}$, with a runtime of 0.22 seconds per 2D-3D US image pair. These results demonstrate that our approach can achieve accurate alignment and clinically acceptable runtime, indicating potential for clinical translation.
A multi-task learning framework for carotid plaque segmentation and classification from ultrasound images
Jul 02, 2023



Abstract:Carotid plaque segmentation and classification play important roles in the treatment of atherosclerosis and assessment for risk of stroke. Although deep learning methods have been used for carotid plaque segmentation and classification, most focused on a single task and ignored the relationship between the segmentation and classification of carotid plaques. Therefore, we propose a multi-task learning framework for ultrasound carotid plaque segmentation and classification, which utilizes a region-weight module (RWM) and a sample-weight module (SWM) to exploit the correlation between these two tasks. The RWM provides a plaque regional prior knowledge to the classification task, while the SWM is designed to learn the categorical sample weight for the segmentation task. A total of 1270 2D ultrasound images of carotid plaques were collected from Zhongnan Hospital (Wuhan, China) for our experiments. The results of the experiments showed that the proposed method can significantly improve the performance compared to existing networks trained for a single task, with an accuracy of 85.82% for classification and a Dice similarity coefficient of 84.92% for segmentation. In the ablation study, the results demonstrated that both the designed RWM and SWM were beneficial in improving the network's performance. Therefore, we believe that the proposed method could be useful for carotid plaque analysis in clinical trials and practice.
Modern Convex Optimization to Medical Image Analysis
Sep 24, 2018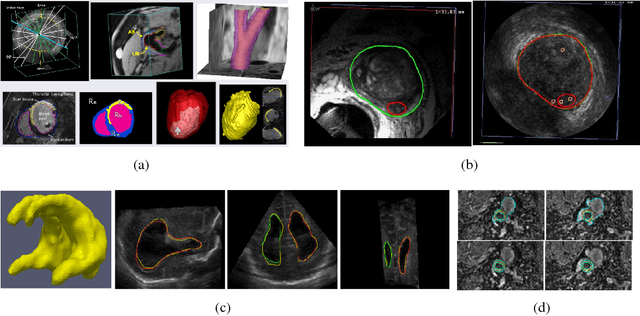

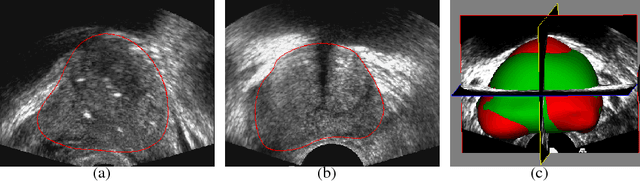
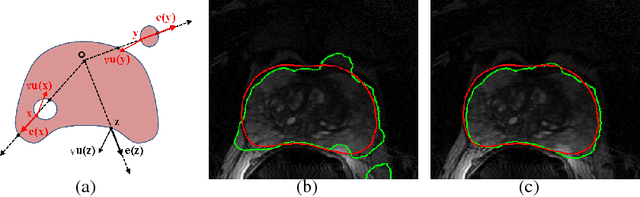
Abstract:Recently, diagnosis, therapy and monitoring of human diseases involve a variety of imaging modalities, such as magnetic resonance imaging(MRI), computed tomography(CT), Ultrasound(US) and Positron-emission tomography(PET) as well as a variety of modern optical techniques. Over the past two decade, it has been recognized that advanced image processing techniques provide valuable information to physicians for diagnosis, image guided therapy and surgery, and monitoring of the treated organ to the therapy. Many researchers and companies have invested significant efforts in the developments of advanced medical image analysis methods; especially in the two core studies of medical image segmentation and registration, segmentations of organs and lesions are used to quantify volumes and shapes used in diagnosis and monitoring treatment; registration of multimodality images of organs improves detection, diagnosis and staging of diseases as well as image-guided surgery and therapy, registration of images obtained from the same modality are used to monitor progression of therapy. These challenging clinical-motivated applications introduce novel and sophisticated mathematical problems which stimulate developments of advanced optimization and computing methods, especially convex optimization attaining optimum in a global sense, hence, bring an enormous spread of research topics for recent computational medical image analysis. Particularly, distinct from the usual image processing, most medical images have a big volume of acquired data, often in 3D or 4D (3D + t) along with great noises or incomplete image information, and form the challenging large-scale optimization problems; how to process such poor 'big data' of medical images efficiently and solve the corresponding optimization problems robustly are the key factors of modern medical image analysis.
RANCOR: Non-Linear Image Registration with Total Variation Regularization
Apr 09, 2014
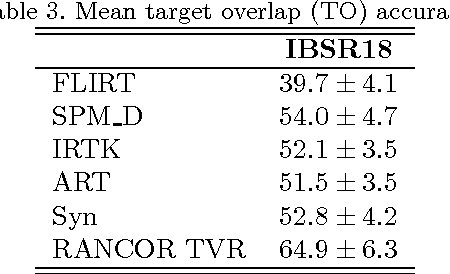
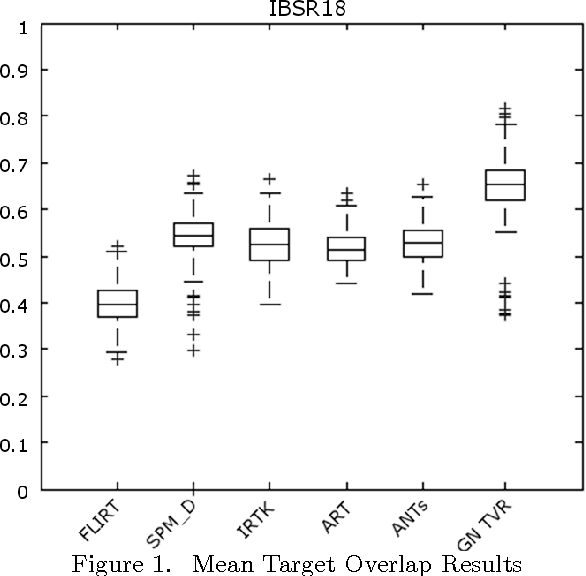

Abstract:Optimization techniques have been widely used in deformable registration, allowing for the incorporation of similarity metrics with regularization mechanisms. These regularization mechanisms are designed to mitigate the effects of trivial solutions to ill-posed registration problems and to otherwise ensure the resulting deformation fields are well-behaved. This paper introduces a novel deformable registration algorithm, RANCOR, which uses iterative convexification to address deformable registration problems under total-variation regularization. Initial comparative results against four state-of-the-art registration algorithms are presented using the Internet Brain Segmentation Repository (IBSR) database.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge