A. Banerjee
User Manual of Automatic Data Curation Tool: A bulk data curator software in Library and Information Science
Oct 31, 2022Abstract:In library and information science, document storage and user-specific document retrieval are the main aspects of digital library services. To preserve the cultural heritage, documents, and literature, we need a common platform where all types of documents are available in a specific format. Our proposed software tool, ADCT can handle a bulk amount of data and transforms different types of raw data into specified metadata information. It generalizes multiple forms of curation logic applied to the source data. As state of the art, many research activities is done in various university to manage their research data in digital library services. The author provides descriptive statistics of library and information service (LIS) activities for information storage and retrieval. The paper shows research on methodology, information gathering, and scientific communication done on library data as an activity of the LIS community. The analysis of LIS data and the change of interest in information storage and retrieval from classification and indexing to retrieval are much important. For Building these types of digitized educational services, automatic data curation is an important stepping stone. The paper shows that automation can complement expertise and knowledge. The author builds an automated data transformation and curation tool that assists users in the analysis of metadata information and effort towards conforming to generic standards using a catalog when a librarian wants to see the adjacent related metadata for an archival collection of congressional correspondence.
Personalized Survival Predictions for Cardiac Transplantation via Trees of Predictors
Apr 11, 2017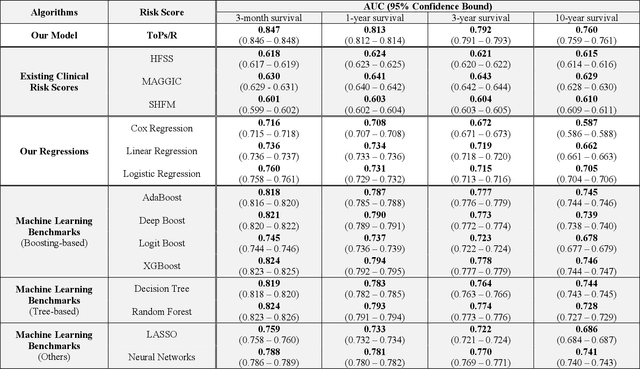
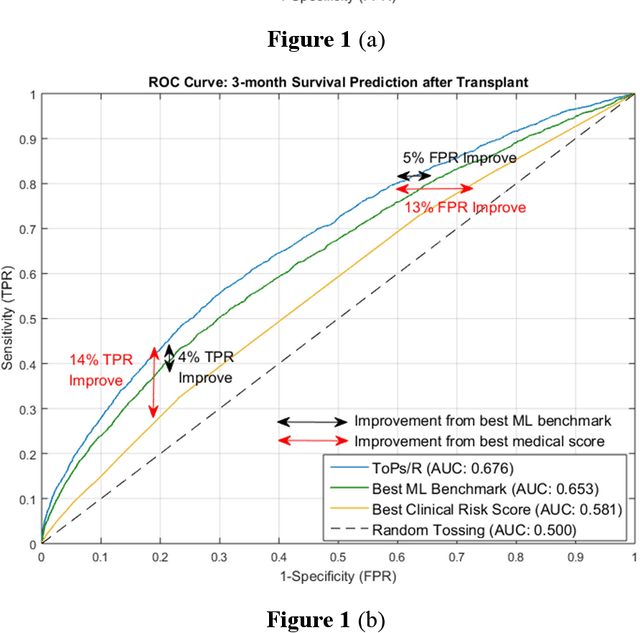
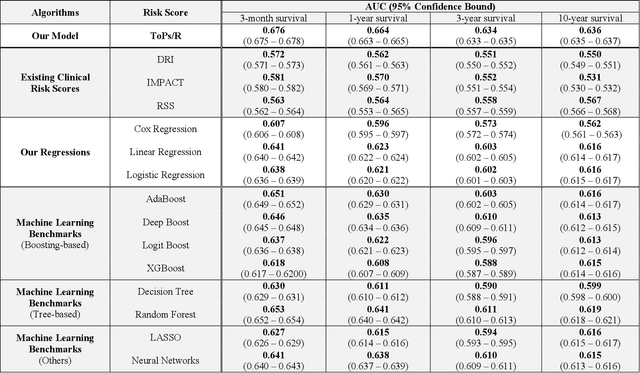
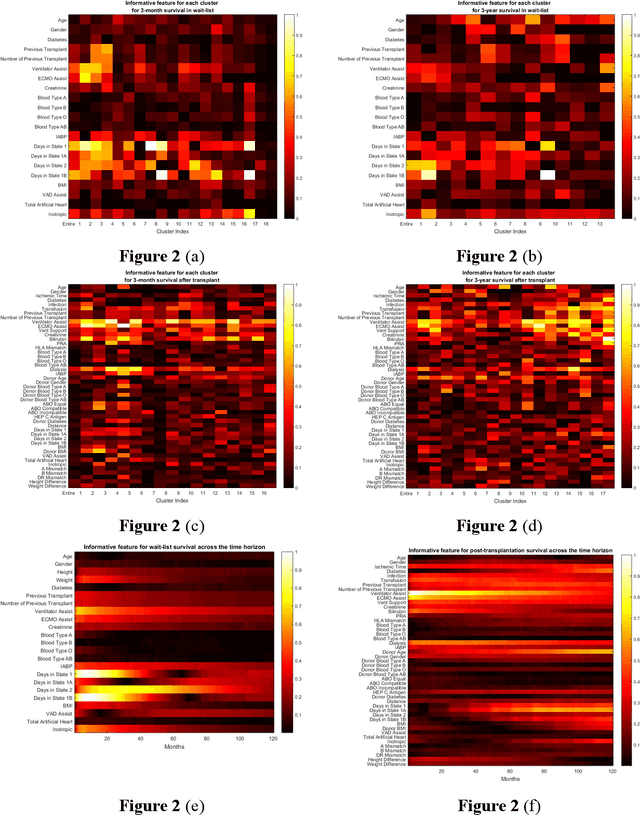
Abstract:Given the limited pool of donor organs, accurate predictions of survival on the wait list and post transplantation are crucial for cardiac transplantation decisions and policy. However, current clinical risk scores do not yield accurate predictions. We develop a new methodology (ToPs, Trees of Predictors) built on the principle that specific predictors should be used for specific clusters within the target population. ToPs discovers these specific clusters of patients and the specific predictor that perform best for each cluster. In comparison with current clinical risk scoring systems, our method provides significant improvements in the prediction of survival time on the wait list and post transplantation. For example, in terms of 3 month survival for patients who were on the US patient wait list in the period 1985 to 2015, our method achieves AUC of 0.847, the best commonly used clinical risk score (MAGGIC) achieves 0.630. In terms of 3 month survival/mortality predictions (in comparison to MAGGIC), holding specificity at 80.0 percents, our algorithm correctly predicts survival for 1,228 (26.0 percents more patients out of 4,723 who actually survived, holding sensitivity at 80.0 percents, our algorithm correctly predicts mortality for 839 (33.0 percents) more patients out of 2,542 who did not survive. Our method achieves similar improvements for other time horizons and for predictions post transplantation. Therefore, we offer a more accurate, personalized approach to survival analysis that can benefit patients, clinicians and policymakers in making clinical decisions and setting clinical policy. Because risk prediction is widely used in diagnostic and prognostic clinical decision making across diseases and clinical specialties, the implications of our methods are far reaching.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge