"cancer detection": models, code, and papers
Semi-supervised multi-task learning for lung cancer diagnosis
May 04, 2018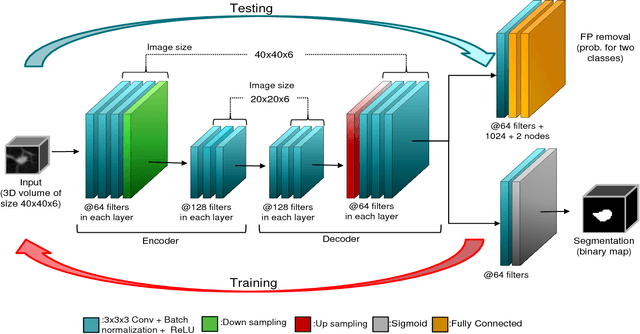

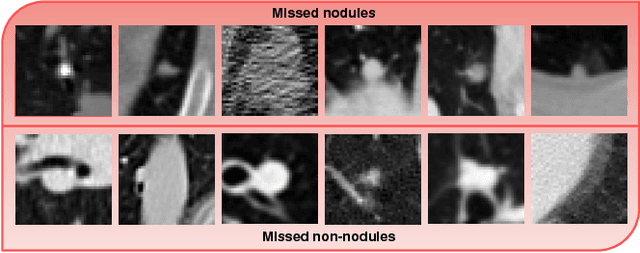
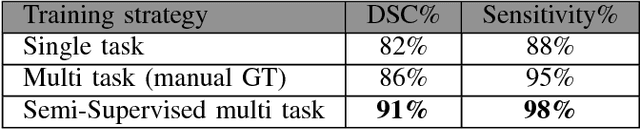
Early detection of lung nodules is of great importance in lung cancer screening. Existing research recognizes the critical role played by CAD systems in early detection and diagnosis of lung nodules. However, many CAD systems, which are used as cancer detection tools, produce a lot of false positives (FP) and require a further FP reduction step. Furthermore, guidelines for early diagnosis and treatment of lung cancer are consist of different shape and volume measurements of abnormalities. Segmentation is at the heart of our understanding of nodules morphology making it a major area of interest within the field of computer aided diagnosis systems. This study set out to test the hypothesis that joint learning of false positive (FP) nodule reduction and nodule segmentation can improve the computer aided diagnosis (CAD) systems' performance on both tasks. To support this hypothesis we propose a 3D deep multi-task CNN to tackle these two problems jointly. We tested our system on LUNA16 dataset and achieved an average dice similarity coefficient (DSC) of 91% as segmentation accuracy and a score of nearly 92% for FP reduction. As a proof of our hypothesis, we showed improvements of segmentation and FP reduction tasks over two baselines. Our results support that joint training of these two tasks through a multi-task learning approach improves system performance on both. We also showed that a semi-supervised approach can be used to overcome the limitation of lack of labeled data for the 3D segmentation task.
Discriminative Localized Sparse Representations for Breast Cancer Screening
Nov 20, 2020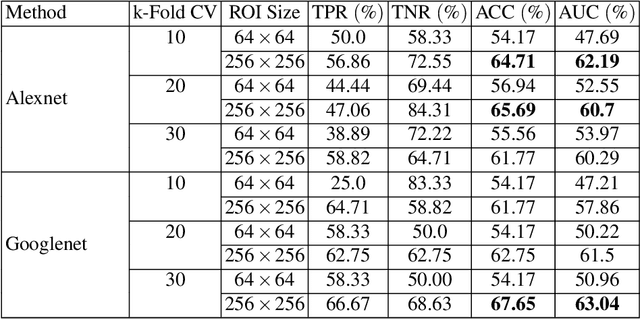
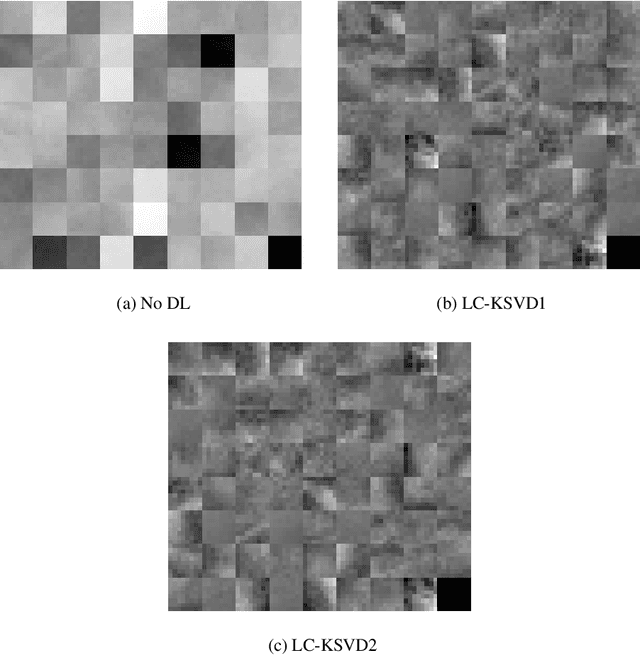
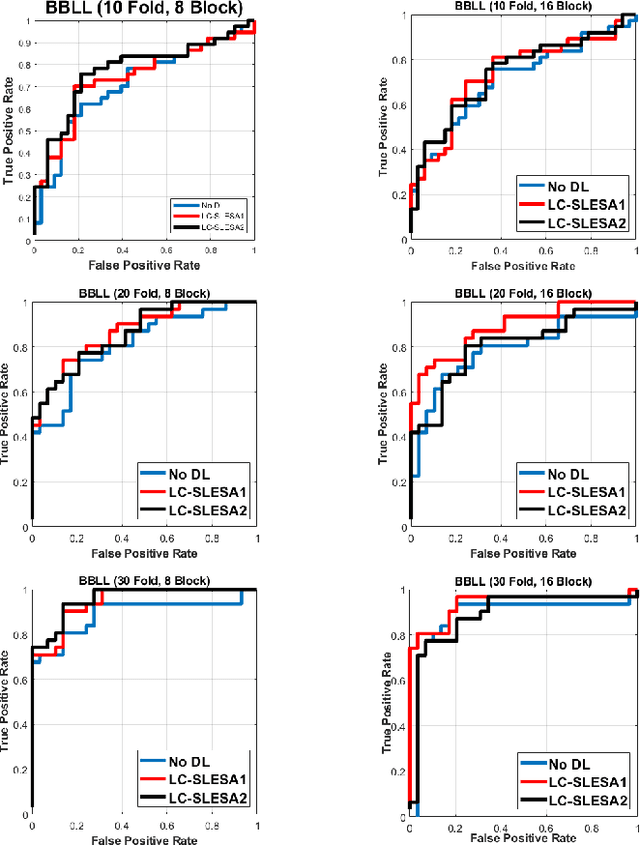
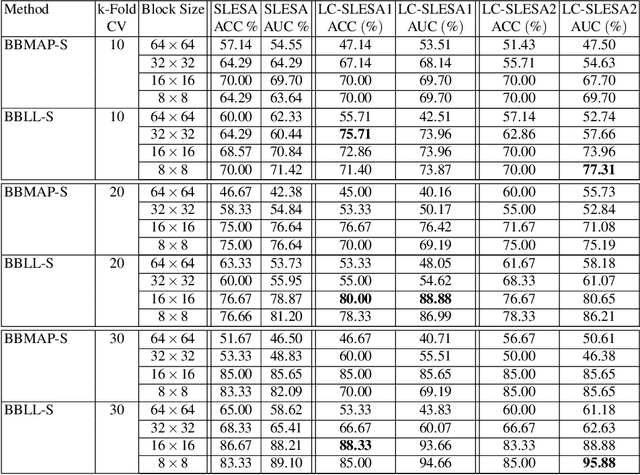

Breast cancer is the most common cancer among women both in developed and developing countries. Early detection and diagnosis of breast cancer may reduce its mortality and improve the quality of life. Computer-aided detection (CADx) and computer-aided diagnosis (CAD) techniques have shown promise for reducing the burden of human expert reading and improve the accuracy and reproducibility of results. Sparse analysis techniques have produced relevant results for representing and recognizing imaging patterns. In this work we propose a method for Label Consistent Spatially Localized Ensemble Sparse Analysis (LC-SLESA). In this work we apply dictionary learning to our block based sparse analysis method to classify breast lesions as benign or malignant. The performance of our method in conjunction with LC-KSVD dictionary learning is evaluated using 10-, 20-, and 30-fold cross validation on the MIAS dataset. Our results indicate that the proposed sparse analyses may be a useful component for breast cancer screening applications.
Colorectal Polyp Detection in Real-world Scenario: Design and Experiment Study
Jan 11, 2021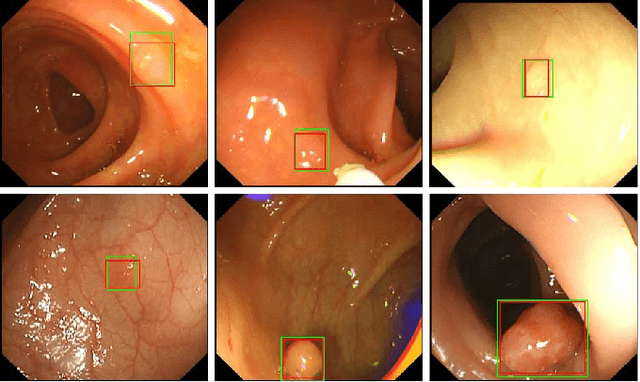
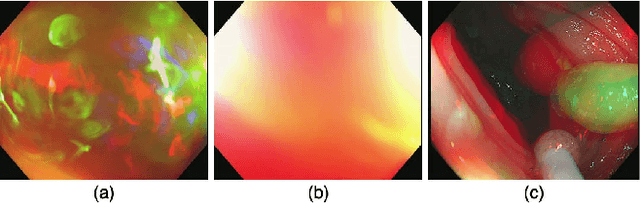
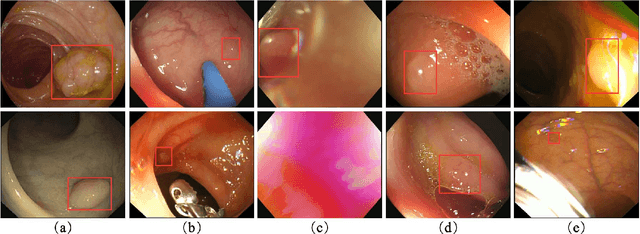
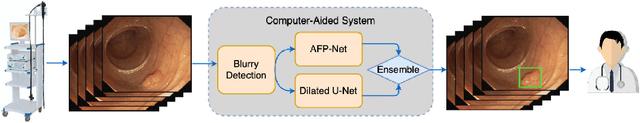

Colorectal polyps are abnormal tissues growing on the intima of the colon or rectum with a high risk of developing into colorectal cancer, the third leading cause of cancer death worldwide. Early detection and removal of colon polyps via colonoscopy have proved to be an effective approach to prevent colorectal cancer. Recently, various CNN-based computer-aided systems have been developed to help physicians detect polyps. However, these systems do not perform well in real-world colonoscopy operations due to the significant difference between images in a real colonoscopy and those in the public datasets. Unlike the well-chosen clear images with obvious polyps in the public datasets, images from a colonoscopy are often blurry and contain various artifacts such as fluid, debris, bubbles, reflection, specularity, contrast, saturation, and medical instruments, with a wide variety of polyps of different sizes, shapes, and textures. All these factors pose a significant challenge to effective polyp detection in a colonoscopy. To this end, we collect a private dataset that contains 7,313 images from 224 complete colonoscopy procedures. This dataset represents realistic operation scenarios and thus can be used to better train the models and evaluate a system's performance in practice. We propose an integrated system architecture to address the unique challenges for polyp detection. Extensive experiments results show that our system can effectively detect polyps in a colonoscopy with excellent performance in real time.
DeepPap: Deep Convolutional Networks for Cervical Cell Classification
Jan 25, 2018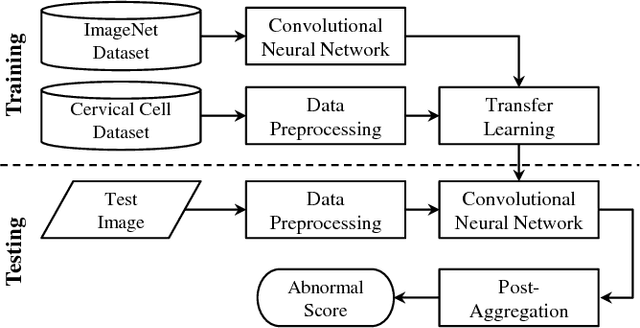
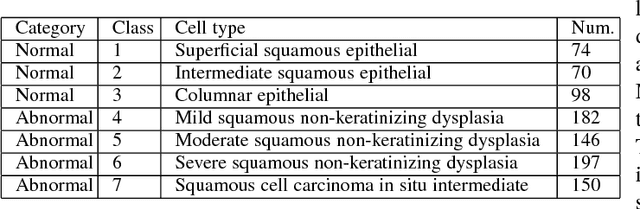

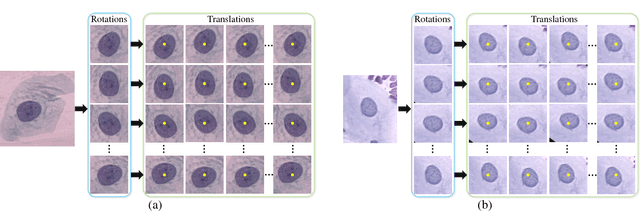

Automation-assisted cervical screening via Pap smear or liquid-based cytology (LBC) is a highly effective cell imaging based cancer detection tool, where cells are partitioned into "abnormal" and "normal" categories. However, the success of most traditional classification methods relies on the presence of accurate cell segmentations. Despite sixty years of research in this field, accurate segmentation remains a challenge in the presence of cell clusters and pathologies. Moreover, previous classification methods are only built upon the extraction of hand-crafted features, such as morphology and texture. This paper addresses these limitations by proposing a method to directly classify cervical cells - without prior segmentation - based on deep features, using convolutional neural networks (ConvNets). First, the ConvNet is pre-trained on a natural image dataset. It is subsequently fine-tuned on a cervical cell dataset consisting of adaptively re-sampled image patches coarsely centered on the nuclei. In the testing phase, aggregation is used to average the prediction scores of a similar set of image patches. The proposed method is evaluated on both Pap smear and LBC datasets. Results show that our method outperforms previous algorithms in classification accuracy (98.3%), area under the curve (AUC) (0.99) values, and especially specificity (98.3%), when applied to the Herlev benchmark Pap smear dataset and evaluated using five-fold cross-validation. Similar superior performances are also achieved on the HEMLBC (H&E stained manual LBC) dataset. Our method is promising for the development of automation-assisted reading systems in primary cervical screening.
Two-Dimensional ARMA Modeling for Breast Cancer Detection and Classification
Jun 19, 2009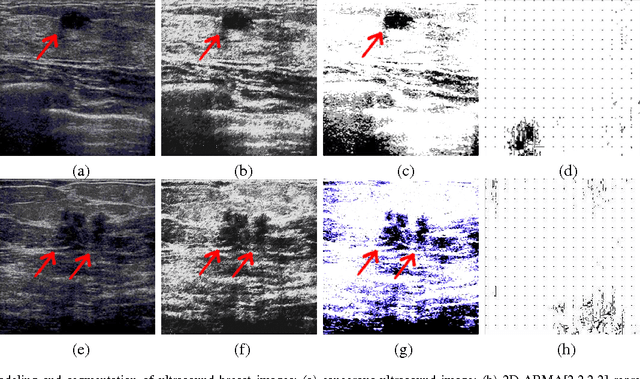
We propose a new model-based computer-aided diagnosis (CAD) system for tumor detection and classification (cancerous v.s. benign) in breast images. Specifically, we show that (x-ray, ultrasound and MRI) images can be accurately modeled by two-dimensional autoregressive-moving average (ARMA) random fields. We derive a two-stage Yule-Walker Least-Squares estimates of the model parameters, which are subsequently used as the basis for statistical inference and biophysical interpretation of the breast image. We use a k-means classifier to segment the breast image into three regions: healthy tissue, benign tumor, and cancerous tumor. Our simulation results on ultrasound breast images illustrate the power of the proposed approach.
Ensembles of Radial Basis Function Networks for Spectroscopic Detection of Cervical Pre-Cancer
May 20, 1999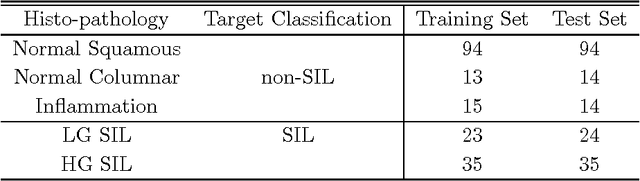
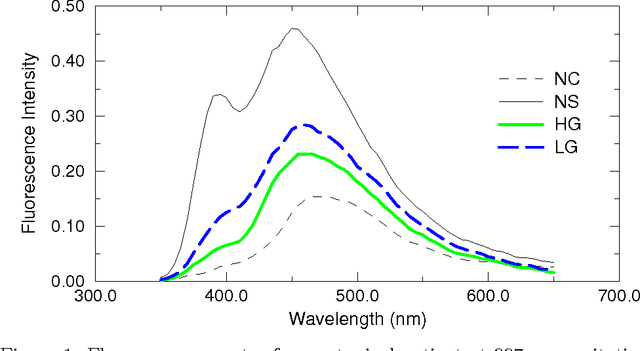
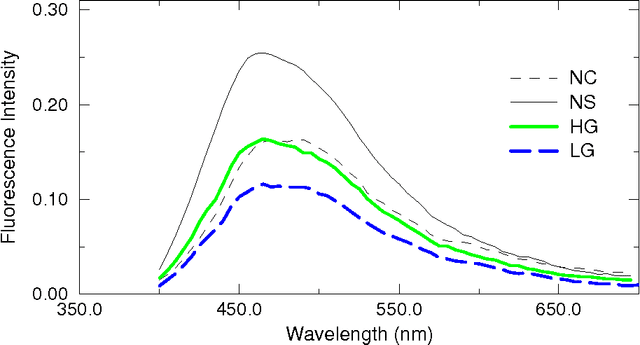
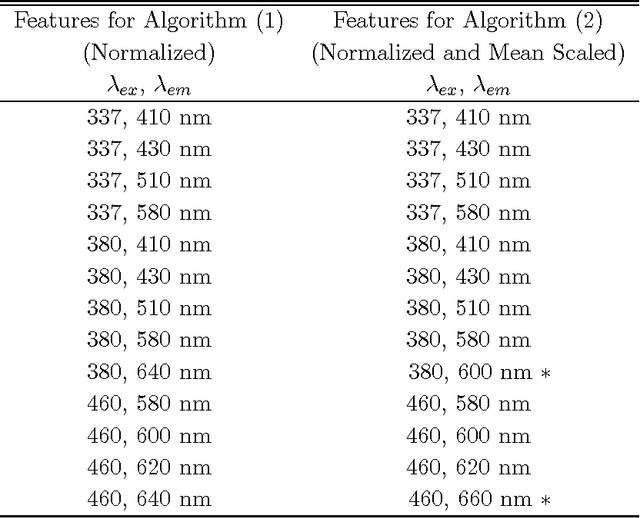
The mortality related to cervical cancer can be substantially reduced through early detection and treatment. However, current detection techniques, such as Pap smear and colposcopy, fail to achieve a concurrently high sensitivity and specificity. In vivo fluorescence spectroscopy is a technique which quickly, non-invasively and quantitatively probes the biochemical and morphological changes that occur in pre-cancerous tissue. A multivariate statistical algorithm was used to extract clinically useful information from tissue spectra acquired from 361 cervical sites from 95 patients at 337, 380 and 460 nm excitation wavelengths. The multivariate statistical analysis was also employed to reduce the number of fluorescence excitation-emission wavelength pairs required to discriminate healthy tissue samples from pre-cancerous tissue samples. The use of connectionist methods such as multi layered perceptrons, radial basis function networks, and ensembles of such networks was investigated. RBF ensemble algorithms based on fluorescence spectra potentially provide automated, and near real-time implementation of pre-cancer detection in the hands of non-experts. The results are more reliable, direct and accurate than those achieved by either human experts or multivariate statistical algorithms.
* 23 pages
Added value of morphological features to breast lesion diagnosis in ultrasound
Jun 06, 2017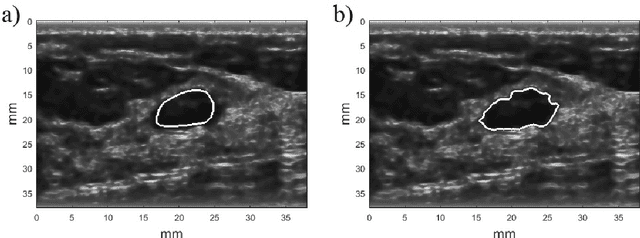
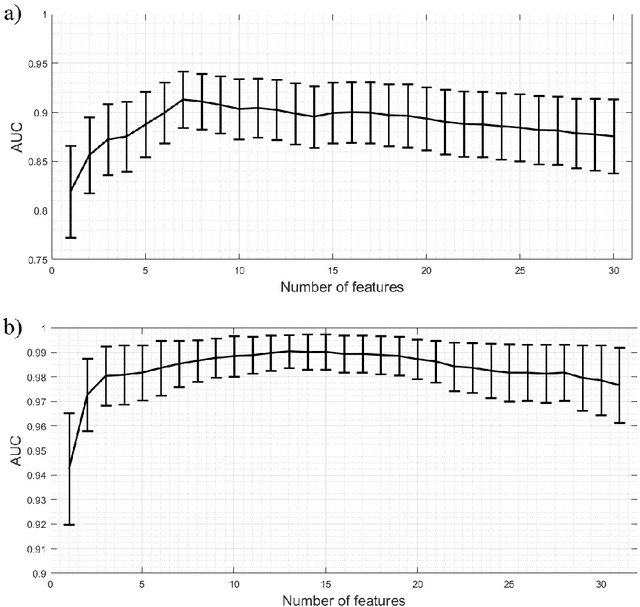
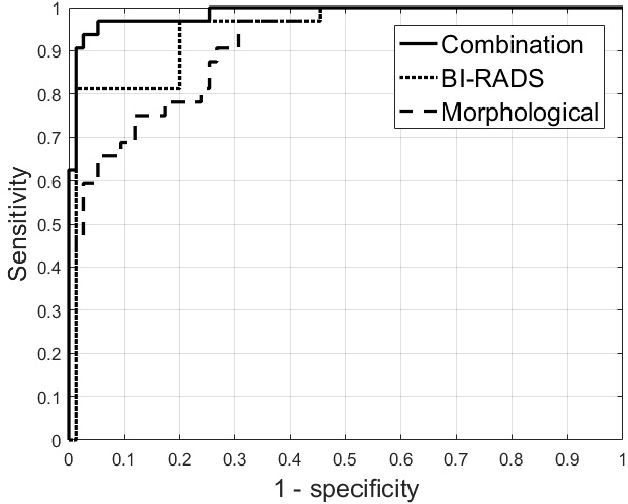
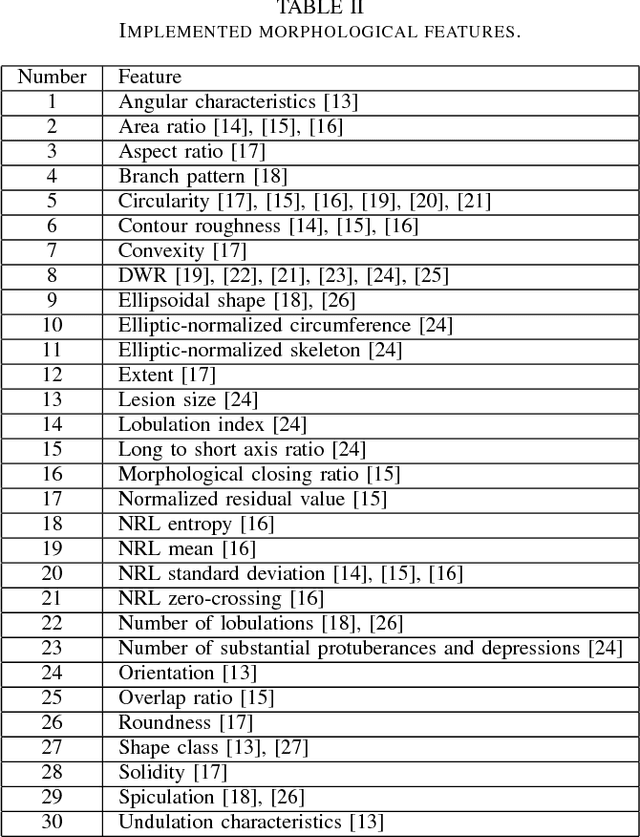
Ultrasound imaging plays an important role in breast lesion differentiation. However, diagnostic accuracy depends on ultrasonographer experience. Various computer aided diagnosis systems has been developed to improve breast cancer detection and reduce the number of unnecessary biopsies. In this study, our aim was to improve breast lesion classification based on the BI-RADS (Breast Imaging - Reporting and Data System). This was accomplished by combining the BI-RADS with morphological features which assess lesion boundary. A dataset of 214 lesion images was used for analysis. 30 morphological features were extracted and feature selection scheme was applied to find features which improve the BI-RADS classification performance. Additionally, the best performing morphological feature subset was indicated. We obtained a better classification by combining the BI-RADS with six morphological features. These features were the extent, overlap ratio, NRL entropy, circularity, elliptic-normalized circumference and the normalized residual value. The area under the receiver operating curve calculated with the use of the combined classifier was 0.986. The best performing morphological feature subset contained six features: the DWR, NRL entropy, normalized residual value, overlap ratio, extent and the morphological closing ratio. For this set, the area under the curve was 0.901. The combination of the radiologist's experience related to the BI-RADS and the morphological features leads to a more effective breast lesion classification.
Scalable Reinforcement-Learning-Based Neural Architecture Search for Cancer Deep Learning Research
Sep 01, 2019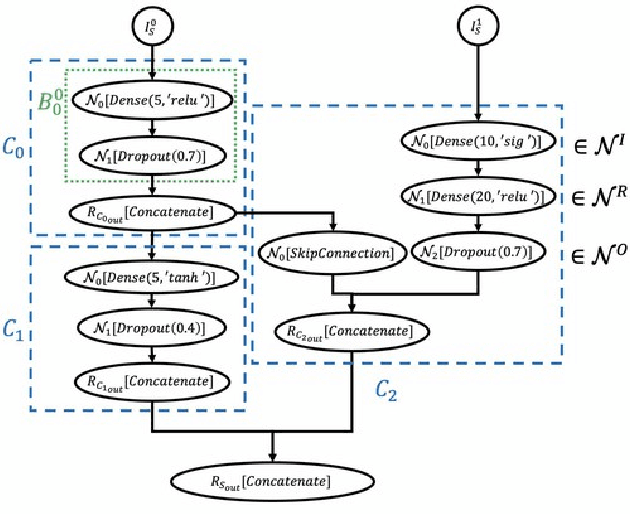
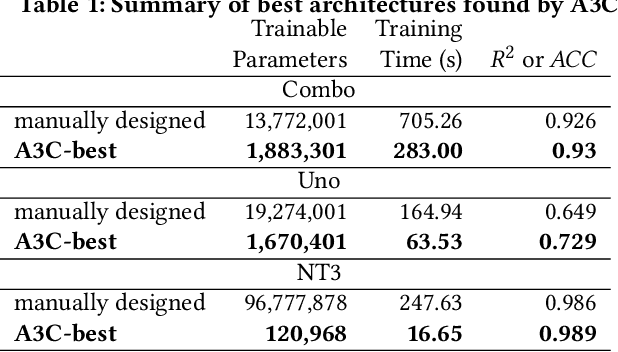
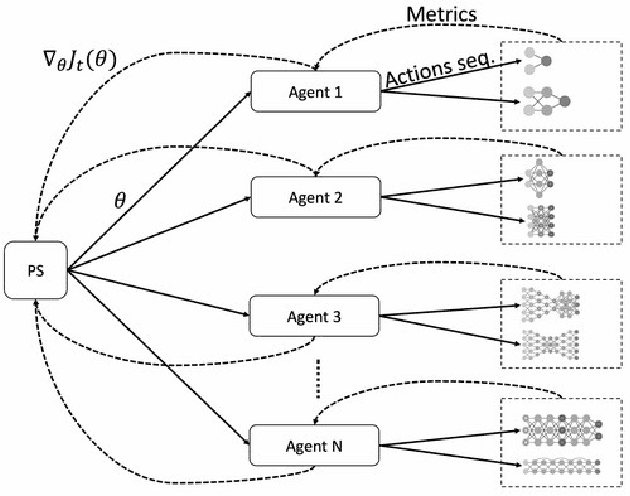
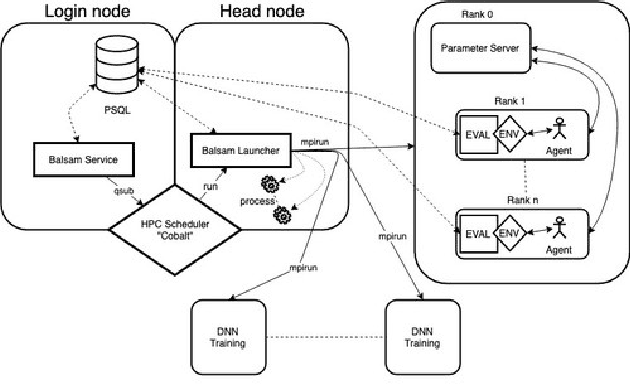
Cancer is a complex disease, the understanding and treatment of which are being aided through increases in the volume of collected data and in the scale of deployed computing power. Consequently, there is a growing need for the development of data-driven and, in particular, deep learning methods for various tasks such as cancer diagnosis, detection, prognosis, and prediction. Despite recent successes, however, designing high-performing deep learning models for nonimage and nontext cancer data is a time-consuming, trial-and-error, manual task that requires both cancer domain and deep learning expertise. To that end, we develop a reinforcement-learning-based neural architecture search to automate deep-learning-based predictive model development for a class of representative cancer data. We develop custom building blocks that allow domain experts to incorporate the cancer-data-specific characteristics. We show that our approach discovers deep neural network architectures that have significantly fewer trainable parameters, shorter training time, and accuracy similar to or higher than those of manually designed architectures. We study and demonstrate the scalability of our approach on up to 1,024 Intel Knights Landing nodes of the Theta supercomputer at the Argonne Leadership Computing Facility.
Sample Efficient Semantic Segmentation using Rotation Equivariant Convolutional Networks
Jul 02, 2018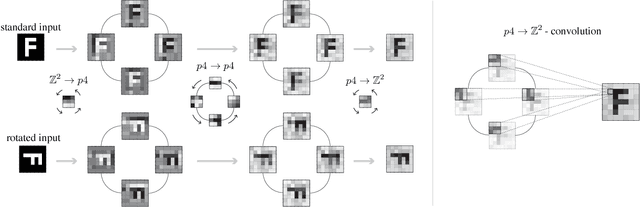
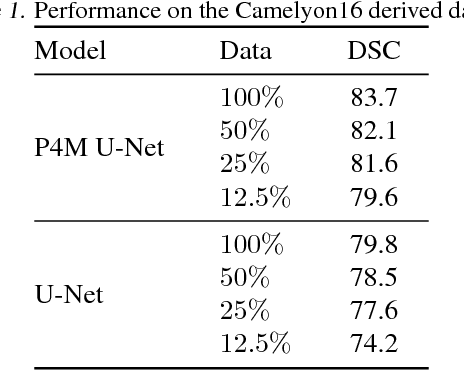
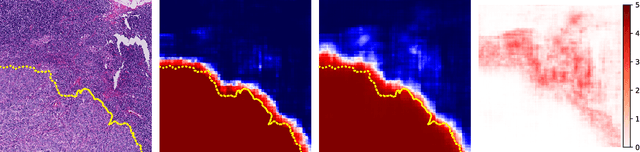
We propose a semantic segmentation model that exploits rotation and reflection symmetries. We demonstrate significant gains in sample efficiency due to increased weight sharing, as well as improvements in robustness to symmetry transformations. The group equivariant CNN framework is extended for segmentation by introducing a new equivariant (G->Z2)-convolution that transforms feature maps on a group to planar feature maps. Also, equivariant transposed convolution is formulated for up-sampling in an encoder-decoder network. To demonstrate improvements in sample efficiency we evaluate on multiple data regimes of a rotation-equivariant segmentation task: cancer metastases detection in histopathology images. We further show the effectiveness of exploiting more symmetries by varying the size of the group.
Interpretative Computer-aided Lung Cancer Diagnosis: from Radiology Analysis to Malignancy Evaluation
Feb 22, 2021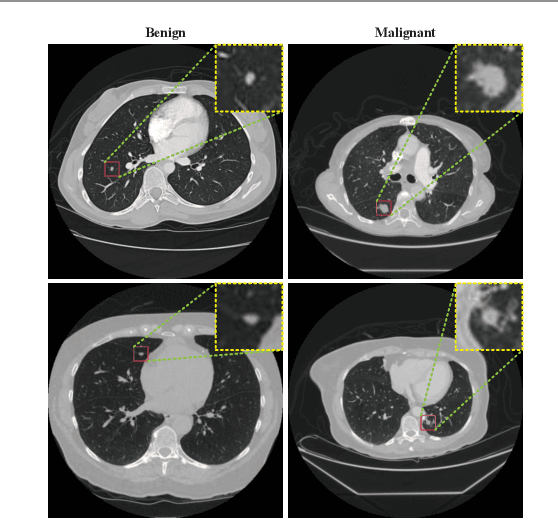
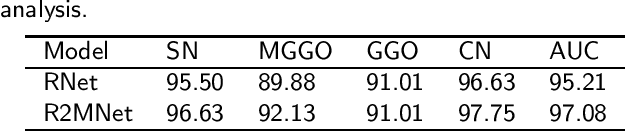
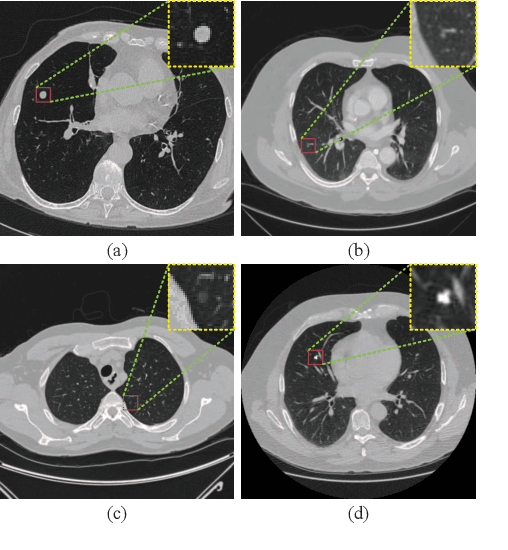
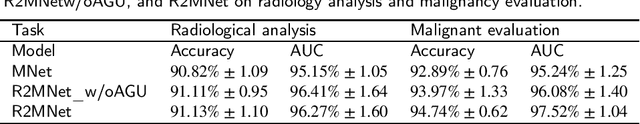
Background and Objective:Computer-aided diagnosis (CAD) systems promote diagnosis effectiveness and alleviate pressure of radiologists. A CAD system for lung cancer diagnosis includes nodule candidate detection and nodule malignancy evaluation. Recently, deep learning-based pulmonary nodule detection has reached satisfactory performance ready for clinical application. However, deep learning-based nodule malignancy evaluation depends on heuristic inference from low-dose computed tomography volume to malignant probability, which lacks clinical cognition. Methods:In this paper, we propose a joint radiology analysis and malignancy evaluation network (R2MNet) to evaluate the pulmonary nodule malignancy via radiology characteristics analysis. Radiological features are extracted as channel descriptor to highlight specific regions of the input volume that are critical for nodule malignancy evaluation. In addition, for model explanations, we propose channel-dependent activation mapping to visualize the features and shed light on the decision process of deep neural network. Results:Experimental results on the LIDC-IDRI dataset demonstrate that the proposed method achieved area under curve of 96.27% on nodule radiology analysis and AUC of 97.52% on nodule malignancy evaluation. In addition, explanations of CDAM features proved that the shape and density of nodule regions were two critical factors that influence a nodule to be inferred as malignant, which conforms with the diagnosis cognition of experienced radiologists. Conclusion:Incorporating radiology analysis with nodule malignant evaluation, the network inference process conforms to the diagnostic procedure of radiologists and increases the confidence of evaluation results. Besides, model interpretation with CDAM features shed light on the regions which DNNs focus on when they estimate nodule malignancy probabilities.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge