"cancer detection": models, code, and papers
A Comparative Study on Polyp Classification using Convolutional Neural Networks
Jul 12, 2020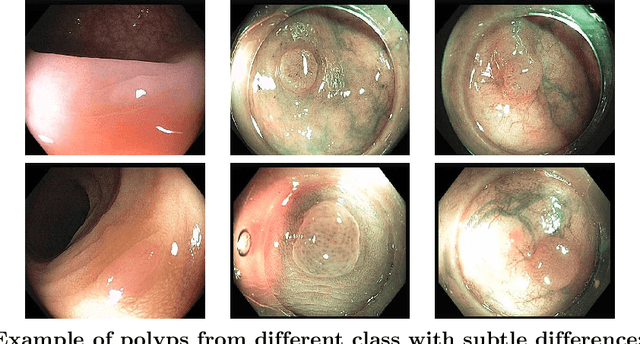
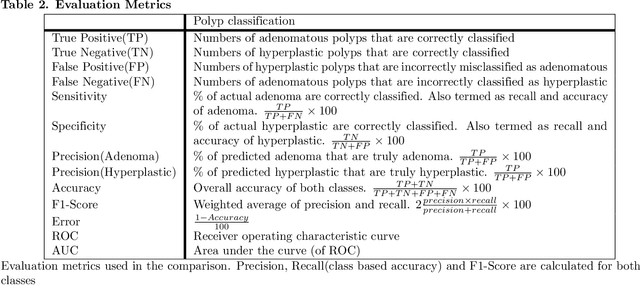
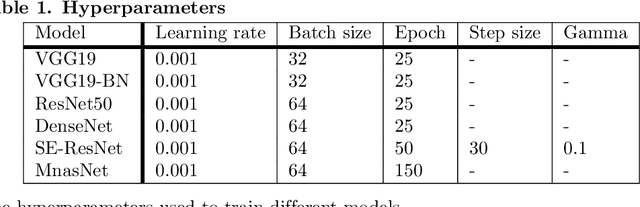
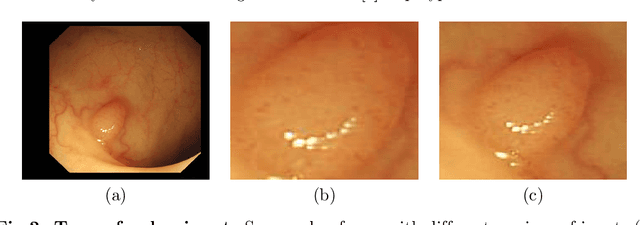

Colorectal cancer is the third most common cancer diagnosed in both men and women in the United States. Most colorectal cancers start as a growth on the inner lining of the colon or rectum, called 'polyp'. Not all polyps are cancerous, but some can develop into cancer. Early detection and recognition of the type of polyps is critical to prevent cancer and change outcomes. However, visual classification of polyps is challenging due to varying illumination conditions of endoscopy, variant texture, appearance, and overlapping morphology between polyps. More importantly, evaluation of polyp patterns by gastroenterologists is subjective leading to a poor agreement among observers. Deep convolutional neural networks have proven very successful in object classification across various object categories. In this work, we compare the performance of the state-of-the-art general object classification models for polyp classification. We trained a total of six CNN models end-to-end using a dataset of 157 video sequences composed of two types of polyps: hyperplastic and adenomatous. Our results demonstrate that the state-of-the-art CNN models can successfully classify polyps with an accuracy comparable or better than reported among gastroenterologists. The results of this study can guide future research in polyp classification.
A Deep Learning Framework for Single-Sided Sound Speed Inversion in Medical Ultrasound
Nov 02, 2018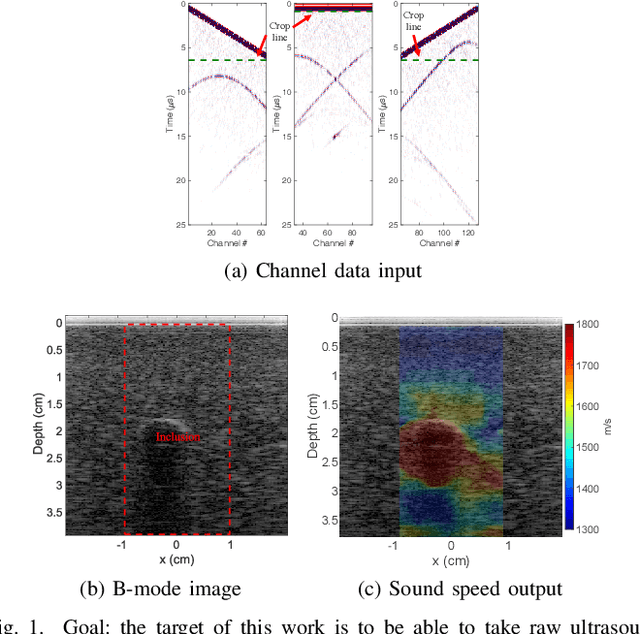
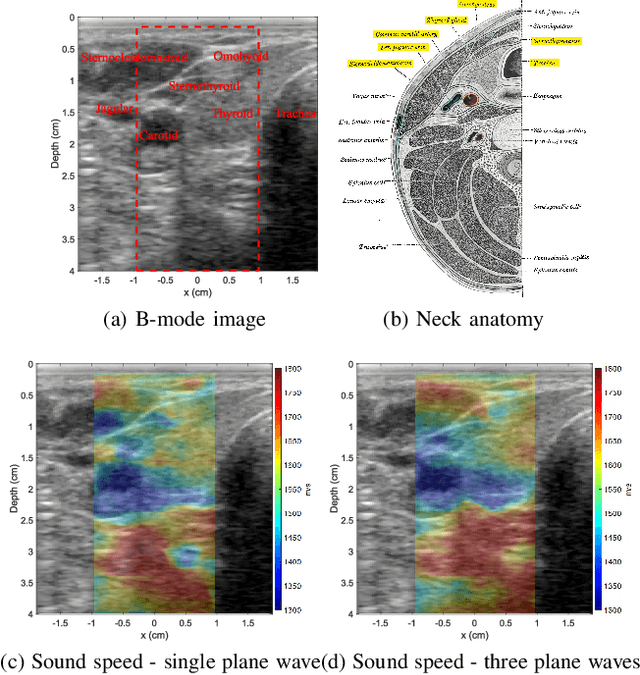
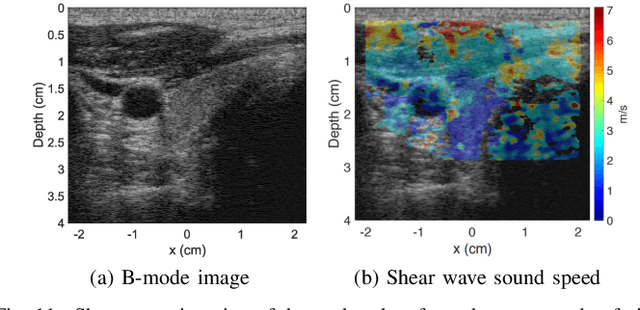
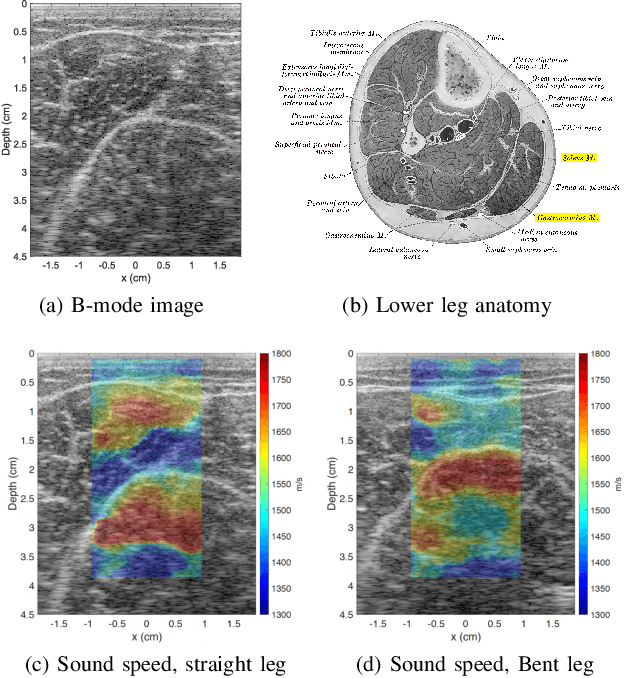
Ultrasound elastography is gaining traction as an accessible and useful diagnostic tool for such things as cancer detection and differentiation as well as liver and thyroid disease diagnostics. Unfortunately, state of the art acoustic radiation force techniques, essential to promote this goal, are limited to high end ultrasound hardware due to high power requirements; are extremely sensitive to patient and sonographer motion; and generally suffer from low frame rates. Researchers have shown that pressure wave velocity possesses similar diagnostic abilities to shear wave velocity. Using pressure waves removes the need for generating shear waves, which in turn enables elasticity based diagnostic techniques on portable and low cost devices. However, current travel time tomography and full waveform inversion techniques for recovering pressure wave velocities require a full circumferential field of view. Focus based techniques, on the other hand, provide only localized measurements, are sensitive to the intermediate medium and require capturing multiple frames. In this paper, we present a single sided sound speed inversion solution using a fully convolutional deep neural network. We show that it is possible to invert for longitudinal sound speed in soft tissue at real time frame rates. For the computation, analysis is performed on channel data information from three diagonal plane waves. This is the first step towards a full waveform solver using a Deep Learning framework for the elastic and viscoelastic inverse problem.
MS-GWNN:multi-scale graph wavelet neural network for breast cancer diagnosis
Dec 29, 2020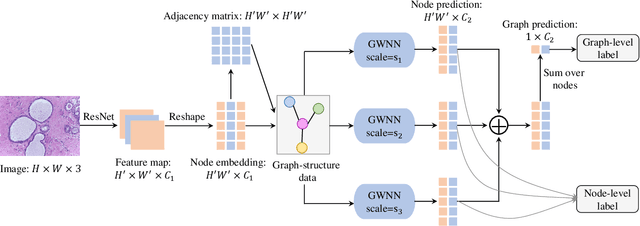
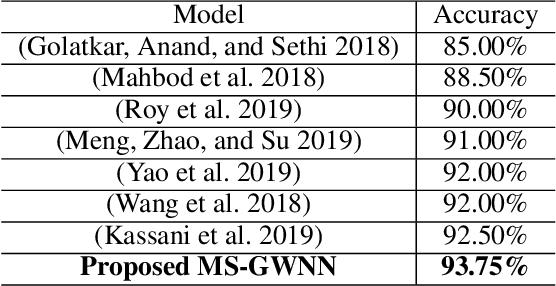
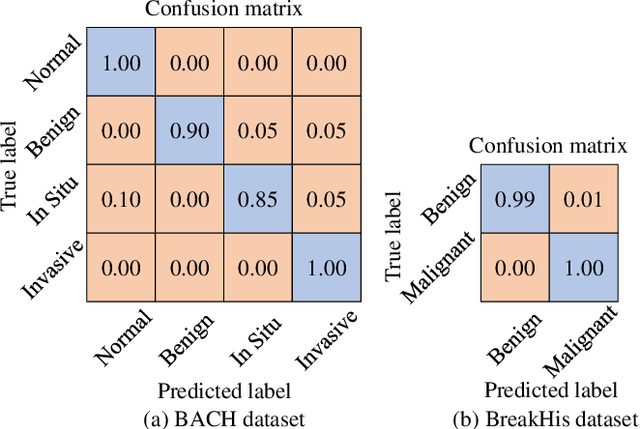
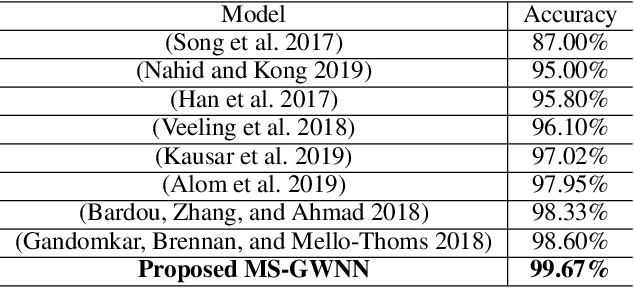
Breast cancer is one of the most common cancers in women worldwide, and early detection can significantly reduce the mortality rate of breast cancer. It is crucial to take multi-scale information of tissue structure into account in the detection of breast cancer. And thus, it is the key to design an accurate computer-aided detection (CAD) system to capture multi-scale contextual features in a cancerous tissue. In this work, we present a novel graph convolutional neural network for histopathological image classification of breast cancer. The new method, named multi-scale graph wavelet neural network (MS-GWNN), leverages the localization property of spectral graph wavelet to perform multi-scale analysis. By aggregating features at different scales, MS-GWNN can encode the multi-scale contextual interactions in the whole pathological slide. Experimental results on two public datasets demonstrate the superiority of the proposed method. Moreover, through ablation studies, we find that multi-scale analysis has a significant impact on the accuracy of cancer diagnosis.
Wavelet based approach for tissue fractal parameter measurement: Pre cancer detection
Mar 21, 2015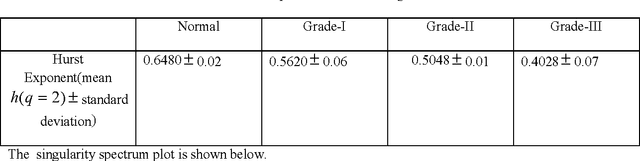
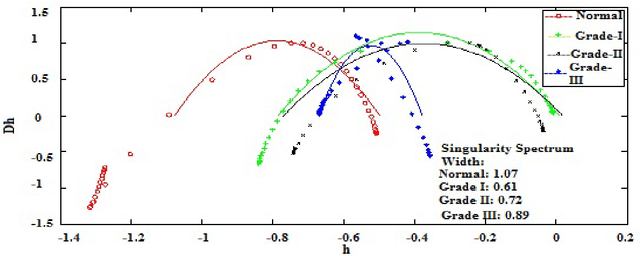
In this paper, we have carried out the detail studies of pre-cancer by wavelet coherency and multifractal based detrended fluctuation analysis (MFDFA) on differential interference contrast (DIC) images of stromal region among different grades of pre-cancer tissues. Discrete wavelet transform (DWT) through Daubechies basis has been performed for identifying fluctuations over polynomial trends for clear characterization and differentiation of tissues. Wavelet coherence plots are performed for identifying the level of correlation in time scale plane between normal and various grades of DIC samples. Applying MFDFA on refractive index variations of cervical tissues, we have observed that the values of Hurst exponent (correlation) decreases from healthy (normal) to pre-cancer tissues. The width of singularity spectrum has a sudden degradation at grade-I in comparison of healthy (normal) tissue but later on it increases as cancer progresses from grade-II to grade-III.
Deep Multi-Resolution Dictionary Learning for Histopathology Image Analysis
Apr 01, 2021
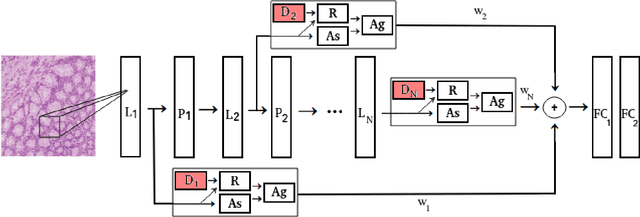
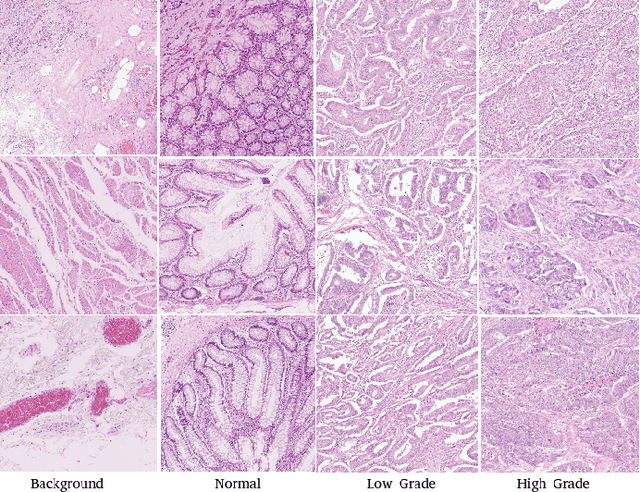
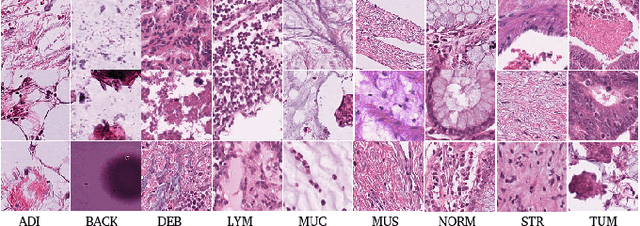
The problem of recognizing various types of tissues present in multi-gigapixel histology images is an important fundamental pre-requisite for downstream analysis of the tumor microenvironment in a bottom-up analysis paradigm for computational pathology. In this paper, we propose a deep dictionary learning approach to solve the problem of tissue phenotyping in histology images. We propose deep Multi-Resolution Dictionary Learning (deepMRDL) in order to benefit from deep texture descriptors at multiple different spatial resolutions. We show the efficacy of the proposed approach through extensive experiments on four benchmark histology image datasets from different organs (colorectal cancer, breast cancer and breast lymphnodes) and tasks (namely, cancer grading, tissue phenotyping, tumor detection and tissue type classification). We also show that the proposed framework can employ most off-the-shelf CNNs models to generate effective deep texture descriptors.
Augmented Networks for Faster Brain Metastases Detection in T1-Weighted Contrast-Enhanced 3D MRI
May 27, 2021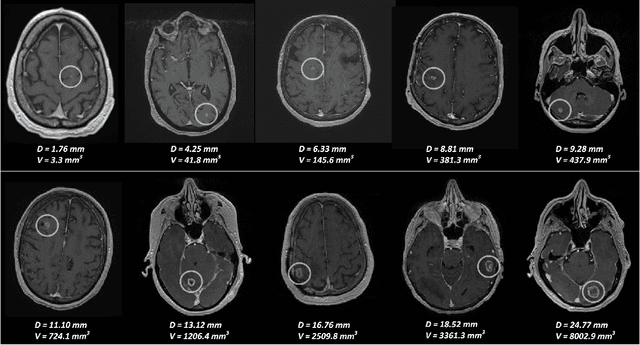
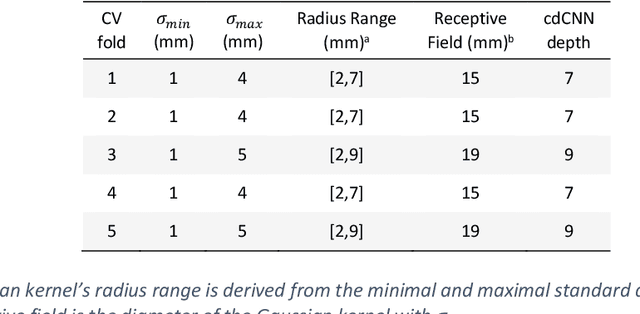
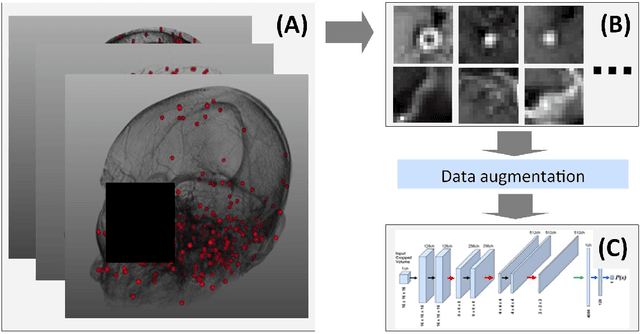
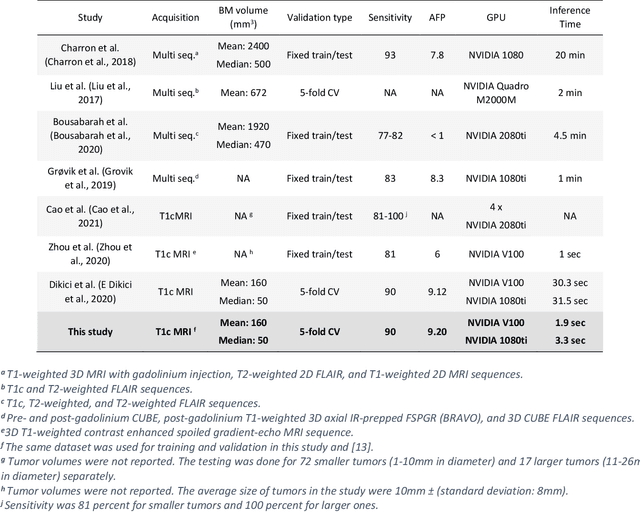
Early detection of brain metastases (BM) is one of the determining factors for the successful treatment of patients with cancer; however, the accurate detection of small BM lesions (< 15mm) remains a challenging task. We previously described a framework for the detection of small BM in single-sequence gadolinium-enhanced T1-weighted 3D MRI datasets. It combined classical image processing (IP) with a dedicated convolutional neural network, taking approximately 30 seconds to process each dataset due to computation-intensive IP stages. To overcome the speed limitation, this study aims to reformulate the framework via an augmented pair of CNNs (eliminating the IP) to reduce the processing times while preserving the BM detection performance. Our previous implementation of the BM detection algorithm utilized Laplacian of Gaussians (LoG) for the candidate selection portion of the solution. In this study, we introduce a novel BM candidate detection CNN (cdCNN) to replace this classical IP stage. The network is formulated to have (1) a similar receptive field as the LoG method, and (2) a bias for the detection of BM lesion loci. The proposed CNN is later augmented with a classification CNN to perform the BM detection task. The cdCNN achieved 97.4% BM detection sensitivity when producing 60K candidates per 3D MRI dataset, while the LoG achieved 96.5% detection sensitivity with 73K candidates. The augmented BM detection framework generated on average 9.20 false-positive BM detections per patient for 90% sensitivity, which is comparable with our previous results. However, it processes each 3D data in 1.9 seconds, presenting a 93.5% reduction in the computation time.
Densely Dilated Spatial Pooling Convolutional Network using benign loss functions for imbalanced volumetric prostate segmentation
Feb 01, 2018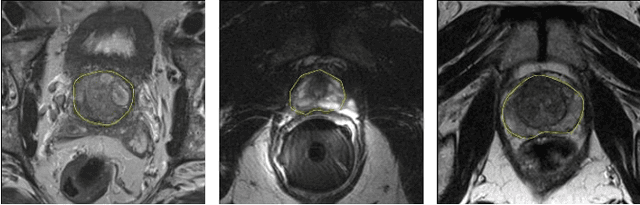
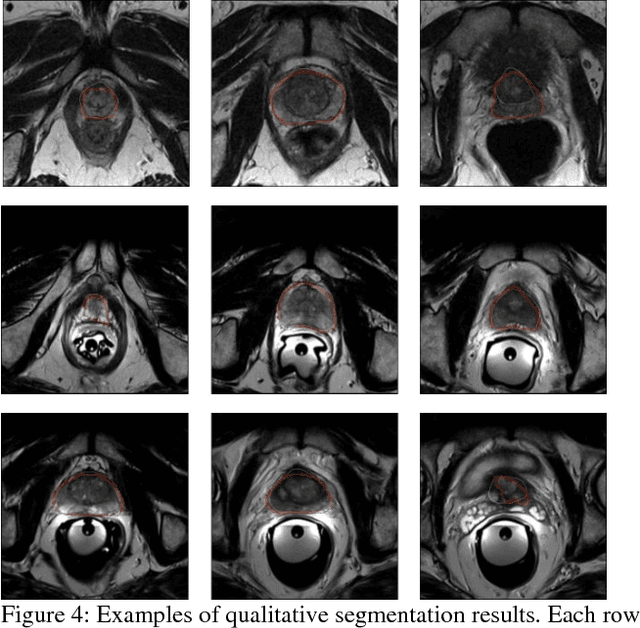
The high incidence rate of prostate disease poses a requirement in early detection for diagnosis. As one of the main imaging methods used for prostate cancer detection, Magnetic Resonance Imaging (MRI) has wide range of appearance and imbalance problems, making automated prostate segmentation fundamental but challenging. Here we propose a novel Densely Dilated Spatial Pooling Convolutional Network (DDSP ConNet) in encoder-decoder structure. It employs dense structure to combine dilated convolution and global pooling, thus supplies coarse segmentation results from encoder and decoder subnet and preserves more contextual information. To obtain richer hierarchical feature maps, residual long connection is furtherly adopted to fuse contexture features. Meanwhile, we adopt DSC loss and Jaccard loss functions to train our DDSP ConNet. We surprisingly found and proved that, in contrast to re-weighted cross entropy, DSC loss and Jaccard loss have a lot of benign properties in theory, including symmetry, continuity and differentiability about the parameters of network. Extensive experiments on the MICCAI PROMISE12 challenge dataset have been done to corroborate the effectiveness of our DDSP ConNet with DSC loss and Jaccard loss. Totally, our method achieves a score of 85.78 in the test dataset, outperforming most of other competitors.
Positive-unlabeled Learning for Cell Detection in Histopathology Images with Incomplete Annotations
Jun 30, 2021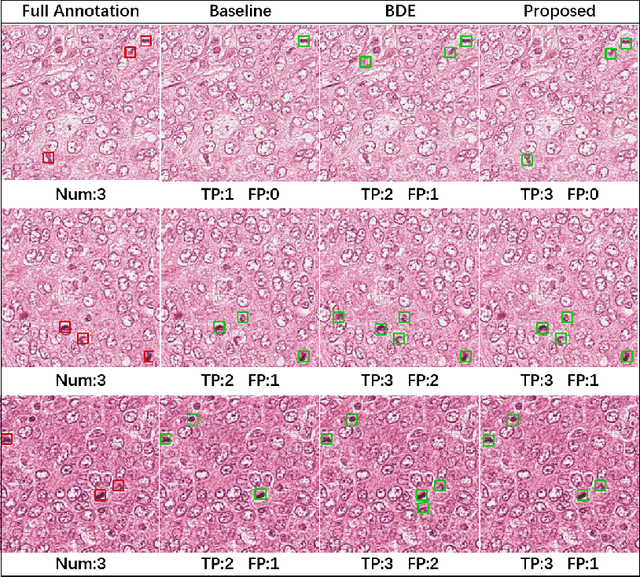
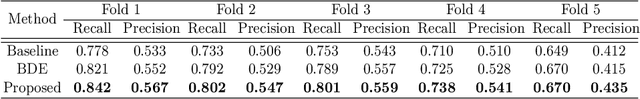
Cell detection in histopathology images is of great value in clinical practice. \textit{Convolutional neural networks} (CNNs) have been applied to cell detection to improve the detection accuracy, where cell annotations are required for network training. However, due to the variety and large number of cells, complete annotations that include every cell of interest in the training images can be challenging. Usually, incomplete annotations can be achieved, where positive labeling results are carefully examined to ensure their reliability but there can be other positive instances, i.e., cells of interest, that are not included in the annotations. This annotation strategy leads to a lack of knowledge about true negative samples. Most existing methods simply treat instances that are not labeled as positive as truly negative during network training, which can adversely affect the network performance. In this work, to address the problem of incomplete annotations, we formulate the training of detection networks as a positive-unlabeled learning problem. Specifically, the classification loss in network training is revised to take into account incomplete annotations, where the terms corresponding to negative samples are approximated with the true positive samples and the other samples of which the labels are unknown. To evaluate the proposed method, experiments were performed on a publicly available dataset for mitosis detection in breast cancer cells, and the experimental results show that our method improves the performance of cell detection given incomplete annotations for training.
Skin Lesion Synthesis with Generative Adversarial Networks
Feb 08, 2019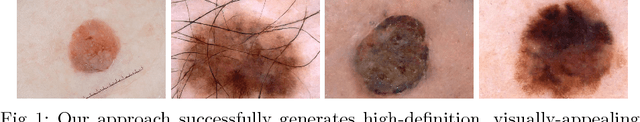
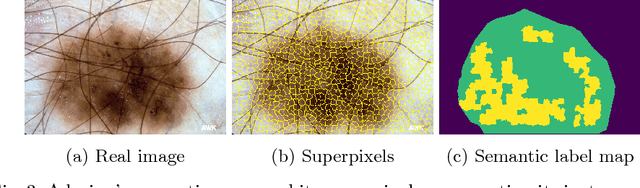
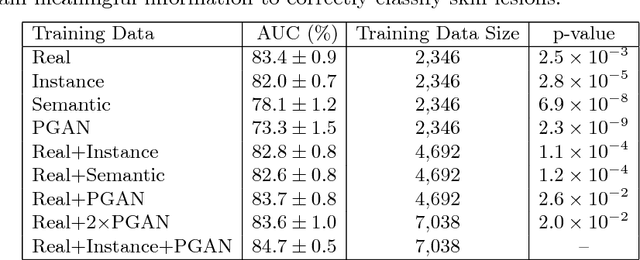
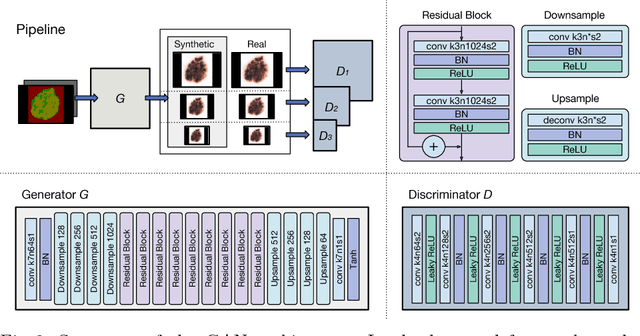

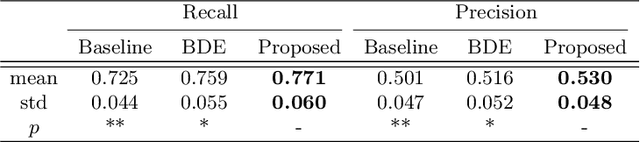
Skin cancer is by far the most common type of cancer. Early detection is the key to increase the chances for successful treatment significantly. Currently, Deep Neural Networks are the state-of-the-art results on automated skin cancer classification. To push the results further, we need to address the lack of annotated data, which is expensive and require much effort from specialists. To bypass this problem, we propose using Generative Adversarial Networks for generating realistic synthetic skin lesion images. To the best of our knowledge, our results are the first to show visually-appealing synthetic images that comprise clinically-meaningful information.
Diagnosis of Breast Cancer using Hybrid Transfer Learning
Mar 23, 2020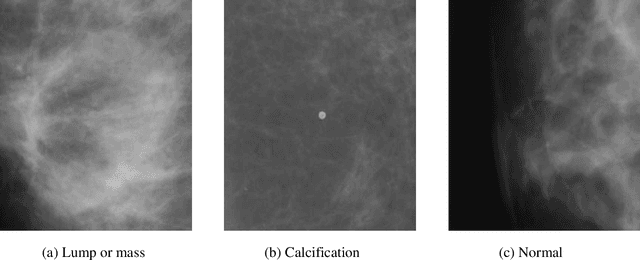
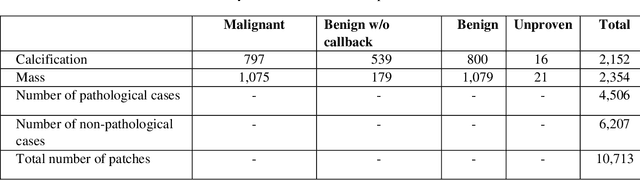
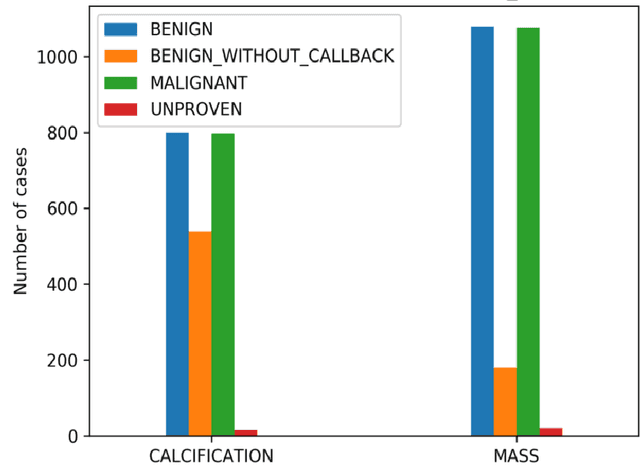

Breast cancer is a common cancer for women. Early detection of breast cancer can considerably increase the survival rate of women. This paper mainly focuses on transfer learning process to detect breast cancer. Modified VGG (MVGG), residual network, mobile network is proposed and implemented in this paper. DDSM dataset is used in this paper. Experimental results show that our proposed hybrid transfers learning model (Fusion of MVGG16 and ImageNet) provides an accuracy of 88.3% where the number of epoch is 15. On the other hand, only modified VGG 16 architecture (MVGG 16) provides an accuracy 80.8% and MobileNet provides an accuracy of 77.2%. So, it is clearly stated that the proposed hybrid pre-trained network outperforms well compared to single architecture. This architecture can be considered as an effective tool for the radiologists in order to reduce the false negative and false positive rate. Therefore, the efficiency of mammography analysis will be improved.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge