"cancer detection": models, code, and papers
On Breast Cancer Detection: An Application of Machine Learning Algorithms on the Wisconsin Diagnostic Dataset
Mar 06, 2018

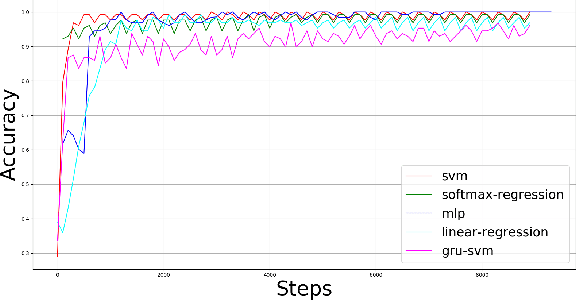
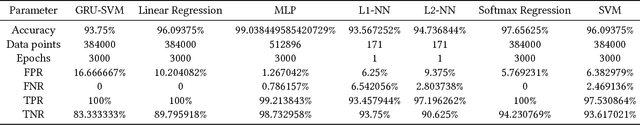
This paper presents a comparison of six machine learning (ML) algorithms: GRU-SVM (Agarap, 2017), Linear Regression, Multilayer Perceptron (MLP), Nearest Neighbor (NN) search, Softmax Regression, and Support Vector Machine (SVM) on the Wisconsin Diagnostic Breast Cancer (WDBC) dataset (Wolberg, Street, & Mangasarian, 1992) by measuring their classification test accuracy and their sensitivity and specificity values. The said dataset consists of features which were computed from digitized images of FNA tests on a breast mass (Wolberg, Street, & Mangasarian, 1992). For the implementation of the ML algorithms, the dataset was partitioned in the following fashion: 70% for training phase, and 30% for the testing phase. The hyper-parameters used for all the classifiers were manually assigned. Results show that all the presented ML algorithms performed well (all exceeded 90% test accuracy) on the classification task. The MLP algorithm stands out among the implemented algorithms with a test accuracy of ~99.04%.
DenseNet approach to segmentation and classification of dermatoscopic skin lesions images
Oct 09, 2021
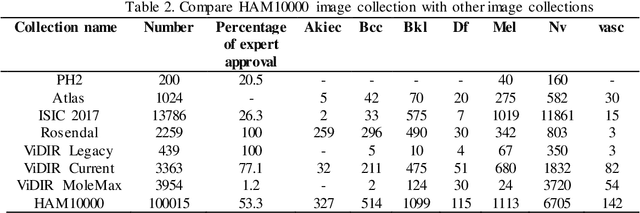
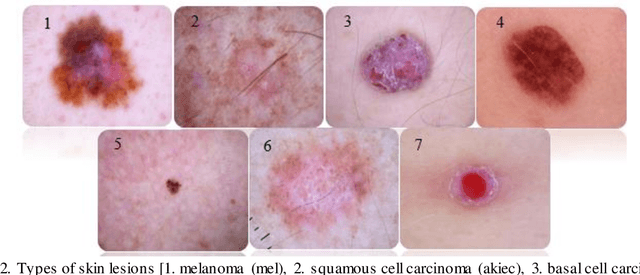

At present, cancer is one of the most important health issues in the world. Because early detection and appropriate treatment in cancer are very effective in the recovery and survival of patients, image processing as a diagnostic tool can help doctors to diagnose in the first recognition of cancer. One of the most important steps in diagnosing a skin lesion is to automatically detect the border of the skin image because the accuracy of the next steps depends on it. If these subtleties are identified, they can have a great impact on the diagnosis of the disease. Therefore, there is a good opportunity to develop more accurate algorithms to analyze such images. This paper proposes an improved method for segmentation and classification for skin lesions using two architectures, the U-Net for image segmentation and the DenseNet121 for image classification which have excellent accuracy. We tested the segmentation architecture of our model on the ISIC-2018 dataset and the classification on the HAM10000 dataset. Our results show that the combination of U-Net and DenseNet121 architectures provides acceptable results in dermatoscopic image analysis compared to previous research. Another classification examined in this study is cancerous and non-cancerous samples. In this classification, cancerous and non-cancerous samples were detected in DenseNet121 network with 79.49% and 93.11% accuracy respectively.
Discovery Radiomics via Evolutionary Deep Radiomic Sequencer Discovery for Pathologically-Proven Lung Cancer Detection
Oct 20, 2017While lung cancer is the second most diagnosed form of cancer in men and women, a sufficiently early diagnosis can be pivotal in patient survival rates. Imaging-based, or radiomics-driven, detection methods have been developed to aid diagnosticians, but largely rely on hand-crafted features which may not fully encapsulate the differences between cancerous and healthy tissue. Recently, the concept of discovery radiomics was introduced, where custom abstract features are discovered from readily available imaging data. We propose a novel evolutionary deep radiomic sequencer discovery approach based on evolutionary deep intelligence. Motivated by patient privacy concerns and the idea of operational artificial intelligence, the evolutionary deep radiomic sequencer discovery approach organically evolves increasingly more efficient deep radiomic sequencers that produce significantly more compact yet similarly descriptive radiomic sequences over multiple generations. As a result, this framework improves operational efficiency and enables diagnosis to be run locally at the radiologist's computer while maintaining detection accuracy. We evaluated the evolved deep radiomic sequencer (EDRS) discovered via the proposed evolutionary deep radiomic sequencer discovery framework against state-of-the-art radiomics-driven and discovery radiomics methods using clinical lung CT data with pathologically-proven diagnostic data from the LIDC-IDRI dataset. The evolved deep radiomic sequencer shows improved sensitivity (93.42%), specificity (82.39%), and diagnostic accuracy (88.78%) relative to previous radiomics approaches.
Evaluation of Joint Multi-Instance Multi-Label Learning For Breast Cancer Diagnosis
Oct 10, 2015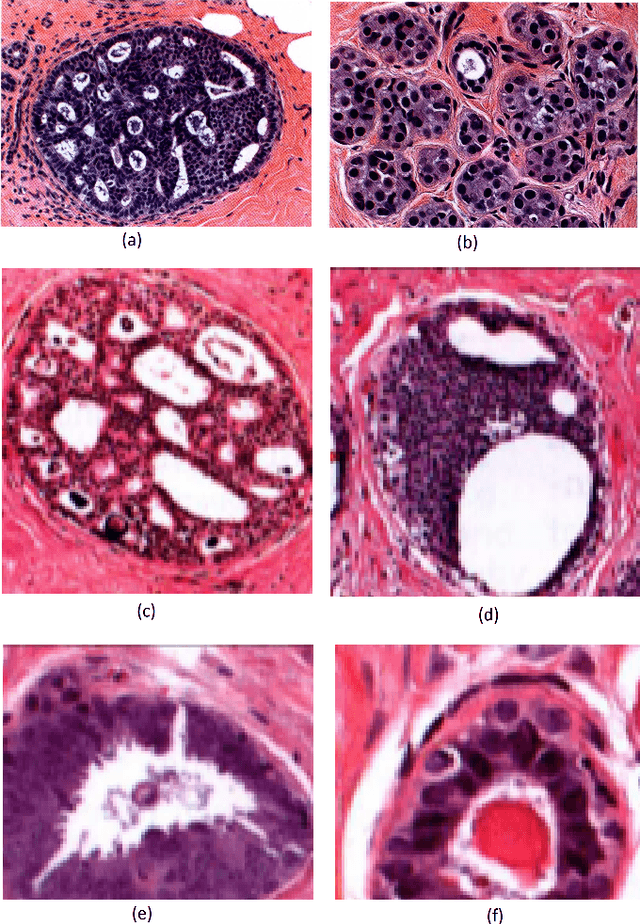
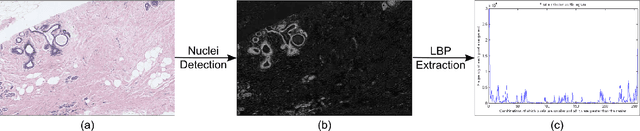
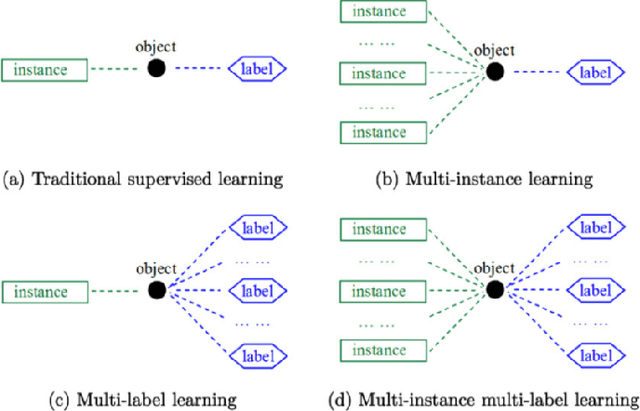

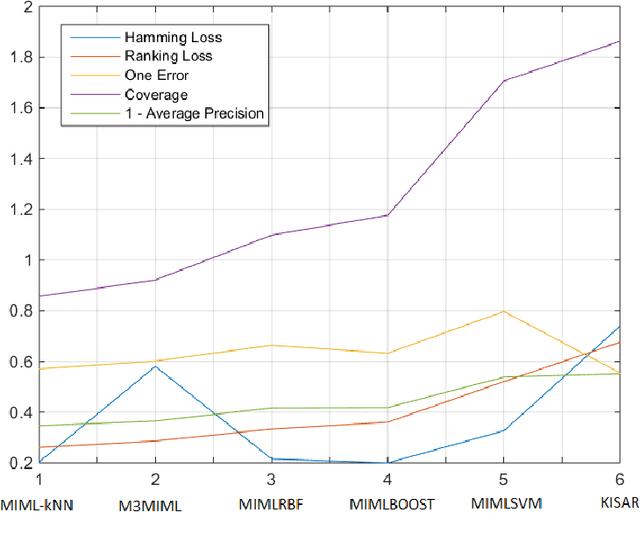
Multi-instance multi-label (MIML) learning is a challenging problem in many aspects. Such learning approaches might be useful for many medical diagnosis applications including breast cancer detection and classification. In this study subset of digiPATH dataset (whole slide digital breast cancer histopathology images) are used for training and evaluation of six state-of-the-art MIML methods. At the end, performance comparison of these approaches are given by means of effective evaluation metrics. It is shown that MIML-kNN achieve the best performance that is %65.3 average precision, where most of other methods attain acceptable results as well.
Ensemble of CNN classifiers using Sugeno Fuzzy Integral Technique for Cervical Cytology Image Classification
Aug 21, 2021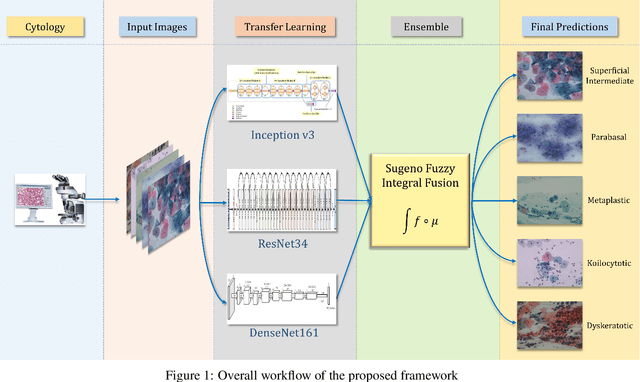
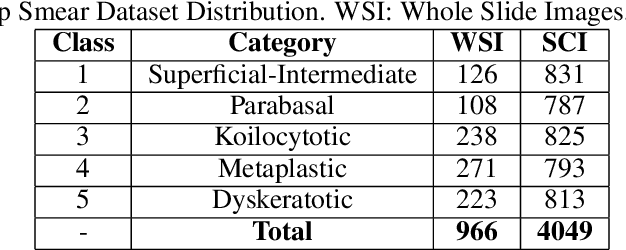
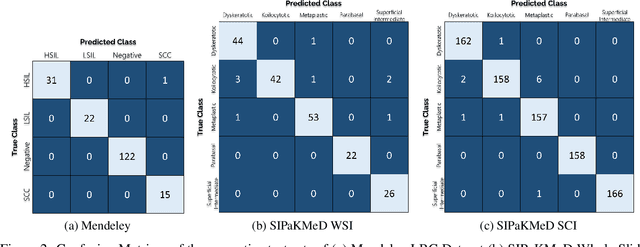
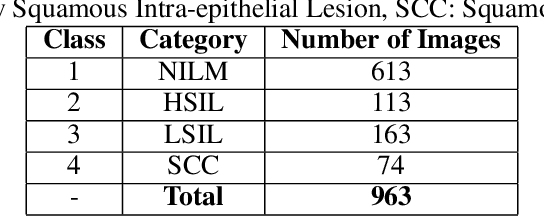
Cervical cancer is the fourth most common category of cancer, affecting more than 500,000 women annually, owing to the slow detection procedure. Early diagnosis can help in treating and even curing cancer, but the tedious, time-consuming testing process makes it impossible to conduct population-wise screening. To aid the pathologists in efficient and reliable detection, in this paper, we propose a fully automated computer-aided diagnosis tool for classifying single-cell and slide images of cervical cancer. The main concern in developing an automatic detection tool for biomedical image classification is the low availability of publicly accessible data. Ensemble Learning is a popular approach for image classification, but simplistic approaches that leverage pre-determined weights to classifiers fail to perform satisfactorily. In this research, we use the Sugeno Fuzzy Integral to ensemble the decision scores from three popular pretrained deep learning models, namely, Inception v3, DenseNet-161 and ResNet-34. The proposed Fuzzy fusion is capable of taking into consideration the confidence scores of the classifiers for each sample, and thus adaptively changing the importance given to each classifier, capturing the complementary information supplied by each, thus leading to superior classification performance. We evaluated the proposed method on three publicly available datasets, the Mendeley Liquid Based Cytology (LBC) dataset, the SIPaKMeD Whole Slide Image (WSI) dataset, and the SIPaKMeD Single Cell Image (SCI) dataset, and the results thus yielded are promising. Analysis of the approach using GradCAM-based visual representations and statistical tests, and comparison of the method with existing and baseline models in literature justify the efficacy of the approach.
Using Machine Learning to Automate Mammogram Images Analysis
Dec 06, 2020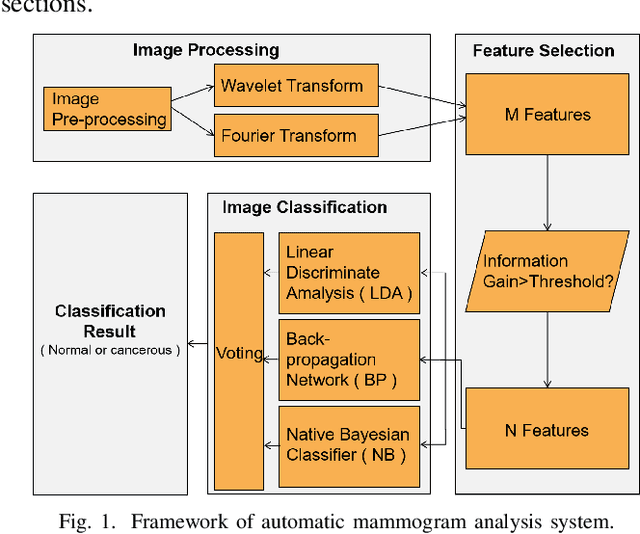
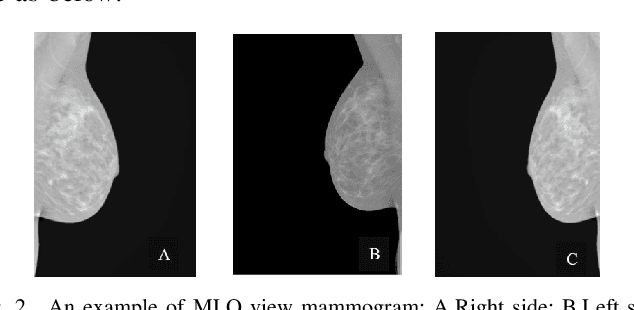
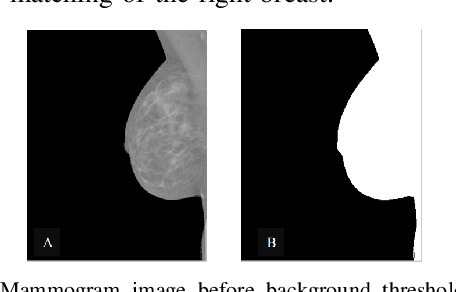
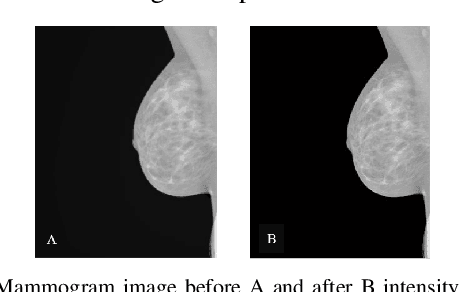
Breast cancer is the second leading cause of cancer-related death after lung cancer in women. Early detection of breast cancer in X-ray mammography is believed to have effectively reduced the mortality rate. However, a relatively high false positive rate and a low specificity in mammography technology still exist. In this work, a computer-aided automatic mammogram analysis system is proposed to process the mammogram images and automatically discriminate them as either normal or cancerous, consisting of three consecutive image processing, feature selection, and image classification stages. In designing the system, the discrete wavelet transforms (Daubechies 2, Daubechies 4, and Biorthogonal 6.8) and the Fourier cosine transform were first used to parse the mammogram images and extract statistical features. Then, an entropy-based feature selection method was implemented to reduce the number of features. Finally, different pattern recognition methods (including the Back-propagation Network, the Linear Discriminant Analysis, and the Naive Bayes Classifier) and a voting classification scheme were employed. The performance of each classification strategy was evaluated for sensitivity, specificity, and accuracy and for general performance using the Receiver Operating Curve. Our method is validated on the dataset from the Eastern Health in Newfoundland and Labrador of Canada. The experimental results demonstrated that the proposed automatic mammogram analysis system could effectively improve the classification performances.
Conditional Infilling GANs for Data Augmentation in Mammogram Classification
Aug 24, 2018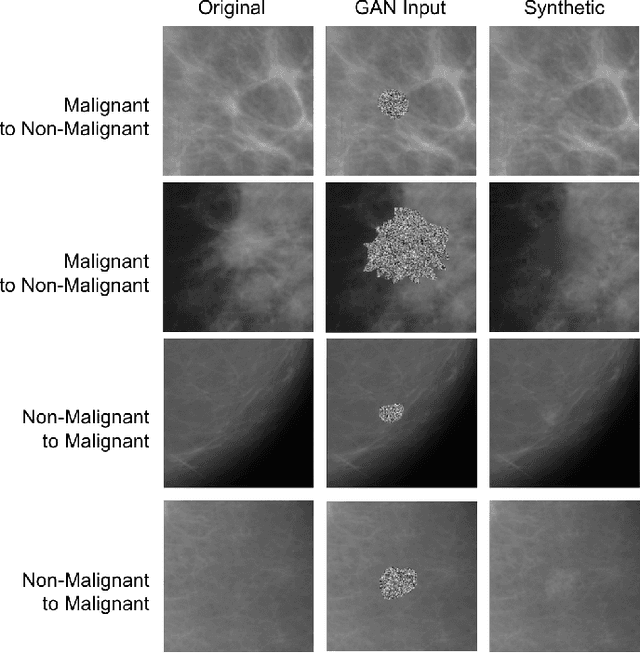
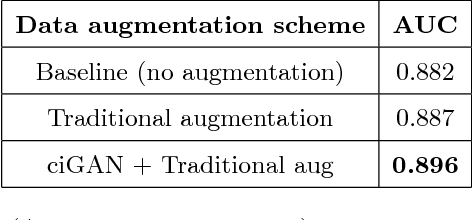
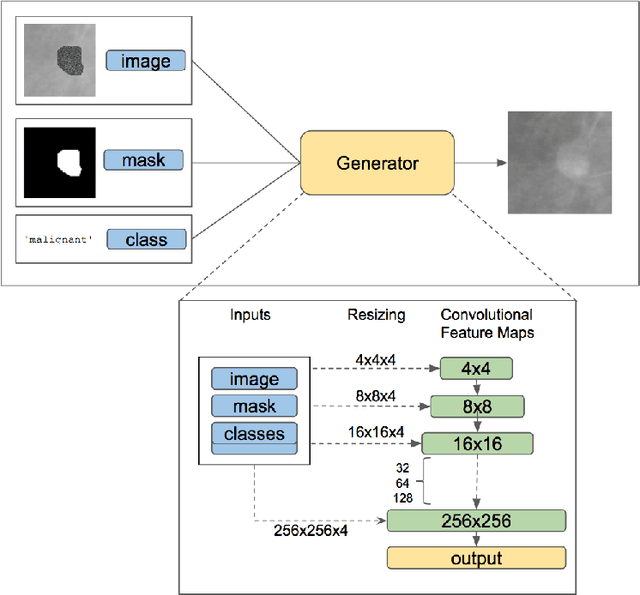
Deep learning approaches to breast cancer detection in mammograms have recently shown promising results. However, such models are constrained by the limited size of publicly available mammography datasets, in large part due to privacy concerns and the high cost of generating expert annotations. Limited dataset size is further exacerbated by substantial class imbalance since "normal" images dramatically outnumber those with findings. Given the rapid progress of generative models in synthesizing realistic images, and the known effectiveness of simple data augmentation techniques (e.g. horizontal flipping), we ask if it is possible to synthetically augment mammogram datasets using generative adversarial networks (GANs). We train a class-conditional GAN to perform contextual in-filling, which we then use to synthesize lesions onto healthy screening mammograms. First, we show that GANs are capable of generating high-resolution synthetic mammogram patches. Next, we experimentally evaluate using the augmented dataset to improve breast cancer classification performance. We observe that a ResNet-50 classifier trained with GAN-augmented training data produces a higher AUROC compared to the same model trained only on traditionally augmented data, demonstrating the potential of our approach.
Assessing domain adaptation techniques for mitosis detection in multi-scanner breast cancer histopathology images
Sep 01, 2021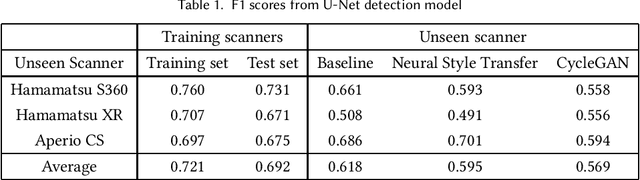
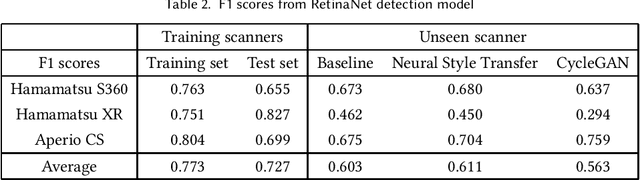
Breast cancer is the most prevalent cancer worldwide and over two million new cases are diagnosed each year. As part of the tumour grading process, histopathologists manually count how many cells are dividing, in a biological process called mitosis. Artificial intelligence (AI) methods have been developed to automatically detect mitotic figures, however these methods often perform poorly when applied to data from outside of the original (training) domain, i.e. they do not generalise well to histology images created using varied staining protocols or digitised using different scanners. Style transfer, a form of domain adaptation, provides the means to transform images from different domains to a shared visual appearance and have been adopted in various applications to mitigate the issue of domain shift. In this paper we train two mitosis detection models and two style transfer methods and evaluate the usefulness of the latter for improving mitosis detection performance in images digitised using different scanners. We found that the best of these models, U-Net without style transfer, achieved an F1-score of 0.693 on the MIDOG 2021 preliminary test set.
A Comprehensive Study on Colorectal Polyp Segmentation with ResUNet++, Conditional Random Field and Test-Time Augmentation
Jul 26, 2021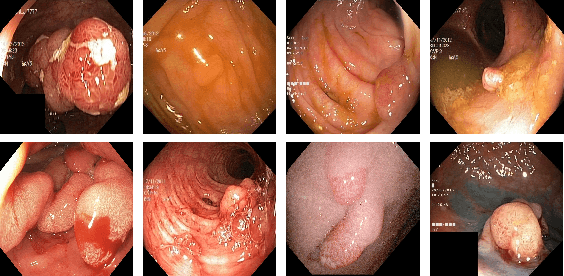
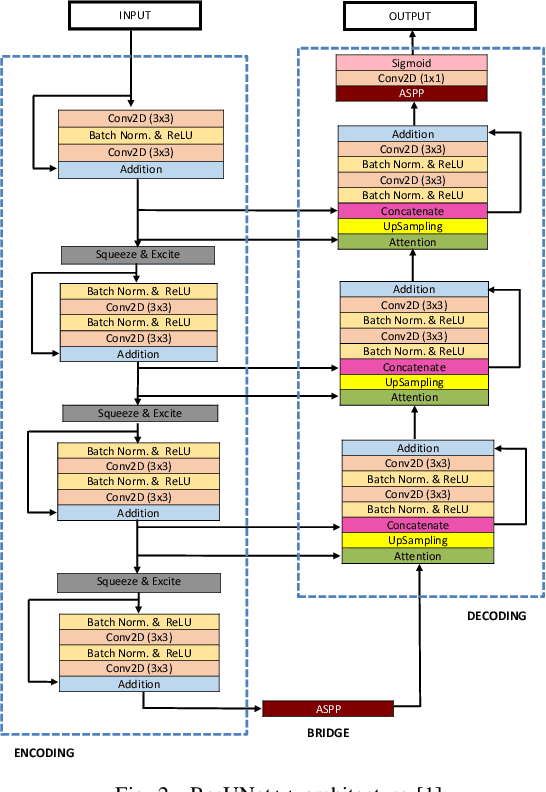
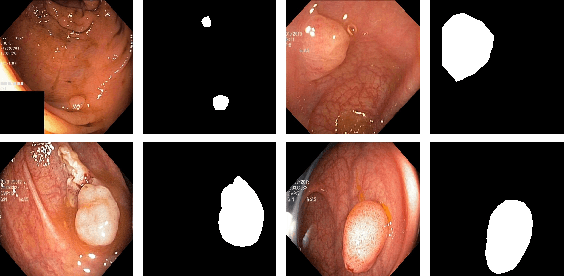
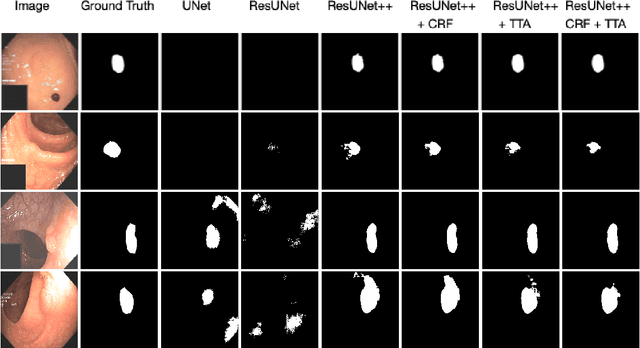
Colonoscopy is considered the gold standard for detection of colorectal cancer and its precursors. Existing examination methods are, however, hampered by high overall miss-rate, and many abnormalities are left undetected. Computer-Aided Diagnosis systems based on advanced machine learning algorithms are touted as a game-changer that can identify regions in the colon overlooked by the physicians during endoscopic examinations, and help detect and characterize lesions. In previous work, we have proposed the ResUNet++ architecture and demonstrated that it produces more efficient results compared with its counterparts U-Net and ResUNet. In this paper, we demonstrate that further improvements to the overall prediction performance of the ResUNet++ architecture can be achieved by using conditional random field and test-time augmentation. We have performed extensive evaluations and validated the improvements using six publicly available datasets: Kvasir-SEG, CVC-ClinicDB, CVC-ColonDB, ETIS-Larib Polyp DB, ASU-Mayo Clinic Colonoscopy Video Database, and CVC-VideoClinicDB. Moreover, we compare our proposed architecture and resulting model with other State-of-the-art methods. To explore the generalization capability of ResUNet++ on different publicly available polyp datasets, so that it could be used in a real-world setting, we performed an extensive cross-dataset evaluation. The experimental results show that applying CRF and TTA improves the performance on various polyp segmentation datasets both on the same dataset and cross-dataset.
Automatic Polyp Segmentation via Multi-scale Subtraction Network
Aug 11, 2021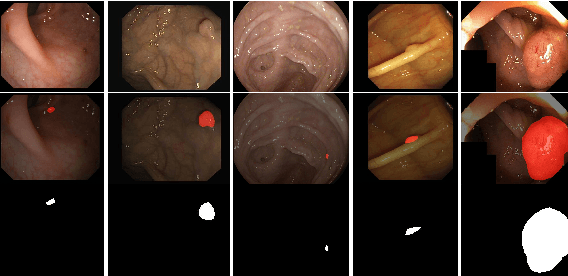
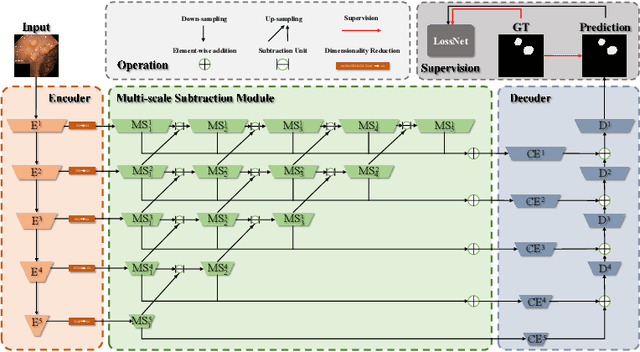
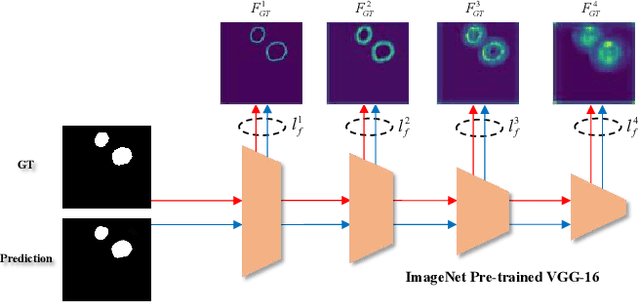
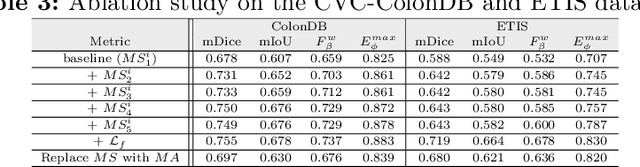
More than 90\% of colorectal cancer is gradually transformed from colorectal polyps. In clinical practice, precise polyp segmentation provides important information in the early detection of colorectal cancer. Therefore, automatic polyp segmentation techniques are of great importance for both patients and doctors. Most existing methods are based on U-shape structure and use element-wise addition or concatenation to fuse different level features progressively in decoder. However, both the two operations easily generate plenty of redundant information, which will weaken the complementarity between different level features, resulting in inaccurate localization and blurred edges of polyps. To address this challenge, we propose a multi-scale subtraction network (MSNet) to segment polyp from colonoscopy image. Specifically, we first design a subtraction unit (SU) to produce the difference features between adjacent levels in encoder. Then, we pyramidally equip the SUs at different levels with varying receptive fields, thereby obtaining rich multi-scale difference information. In addition, we build a training-free network "LossNet" to comprehensively supervise the polyp-aware features from bottom layer to top layer, which drives the MSNet to capture the detailed and structural cues simultaneously. Extensive experiments on five benchmark datasets demonstrate that our MSNet performs favorably against most state-of-the-art methods under different evaluation metrics. Furthermore, MSNet runs at a real-time speed of $\sim$70fps when processing a $352 \times 352$ image. The source code will be publicly available at \url{https://github.com/Xiaoqi-Zhao-DLUT/MSNet}. \keywords{Colorectal Cancer \and Automatic Polyp Segmentation \and Subtraction \and LossNet.}
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge