"cancer detection": models, code, and papers
Application of Gist SVM in Cancer Detection
Mar 06, 2012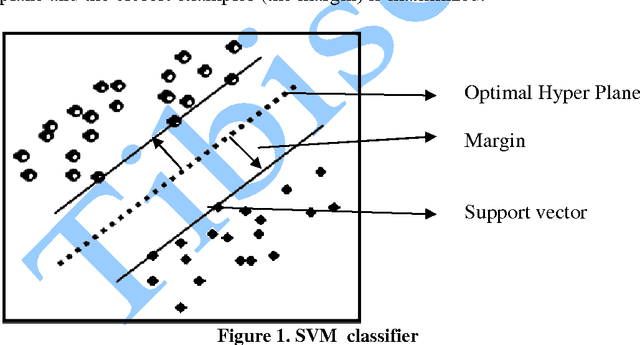
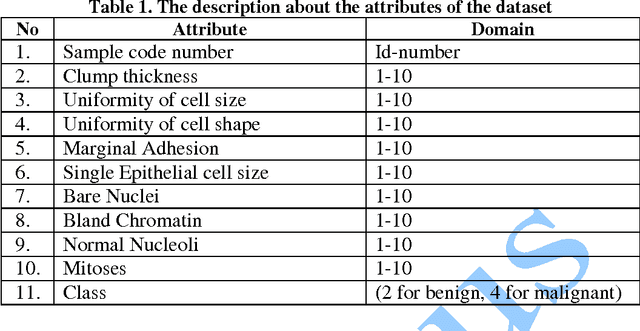

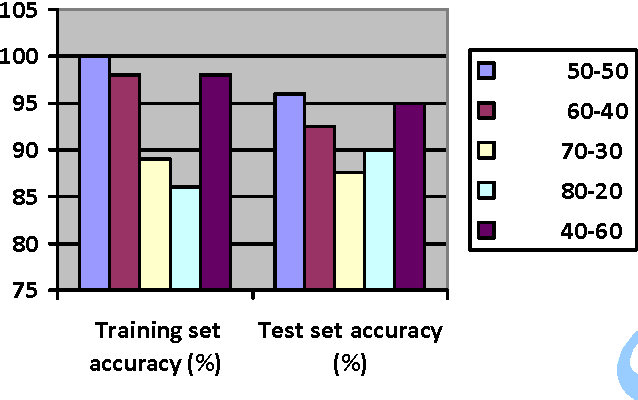
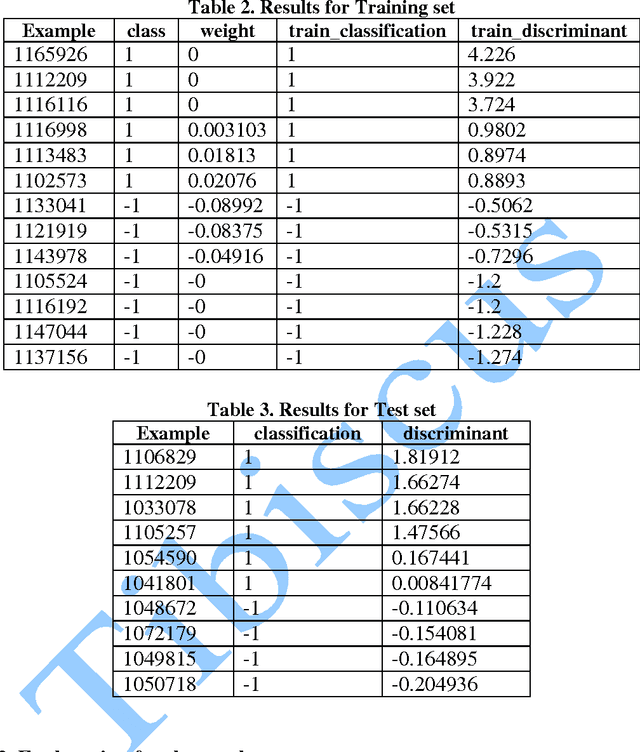
In this paper, we study the application of GIST SVM in disease prediction (detection of cancer). Pattern classification problems can be effectively solved by Support vector machines. Here we propose a classifier which can differentiate patients having benign and malignant cancer cells. To improve the accuracy of classification, we propose to determine the optimal size of the training set and perform feature selection. To find the optimal size of the training set, different sizes of training sets are experimented and the one with highest classification rate is selected. The optimal features are selected through their F-Scores.
* 10 pages
Natural vs Balanced Distribution in Deep Learning on Whole Slide Images for Cancer Detection
Dec 21, 2020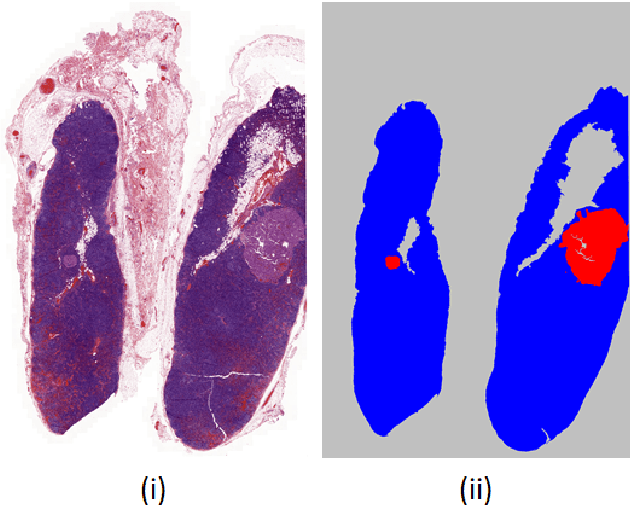
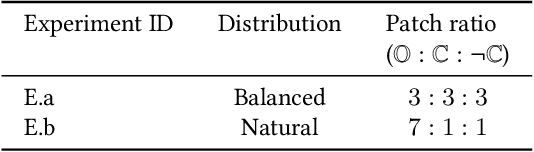
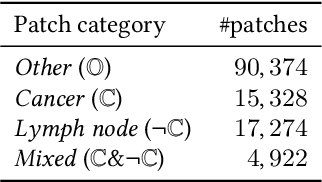


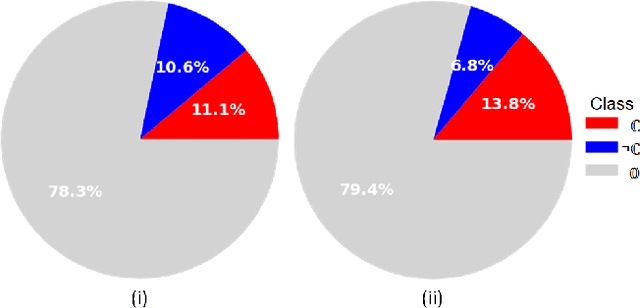
The class distribution of data is one of the factors that regulates the performance of machine learning models. However, investigations on the impact of different distributions available in the literature are very few, sometimes absent for domain-specific tasks. In this paper, we analyze the impact of natural and balanced distributions of the training set in deep learning (DL) models applied on histological images, also known as whole slide images (WSIs). WSIs are considered as the gold standard for cancer diagnosis. In recent years, researchers have turned their attention to DL models to automate and accelerate the diagnosis process. In the training of such DL models, filtering out the non-regions-of-interest from the WSIs and adopting an artificial distribution (usually, a balanced distribution) is a common trend. In our analysis, we show that keeping the WSIs data in their usual distribution (which we call natural distribution) for DL training produces fewer false positives (FPs) with comparable false negatives (FNs) than the artificially-obtained balanced distribution. We conduct an empirical comparative study with 10 random folds for each distribution, comparing the resulting average performance levels in terms of five different evaluation metrics. Experimental results show the effectiveness of the natural distribution over the balanced one across all the evaluation metrics.
Lung nodules segmentation from CT with DeepHealth toolkit
Aug 01, 2022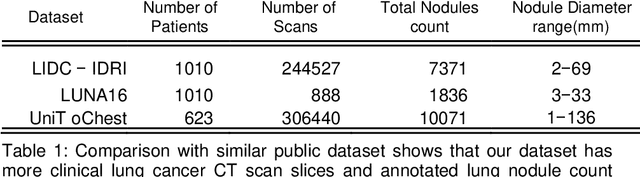
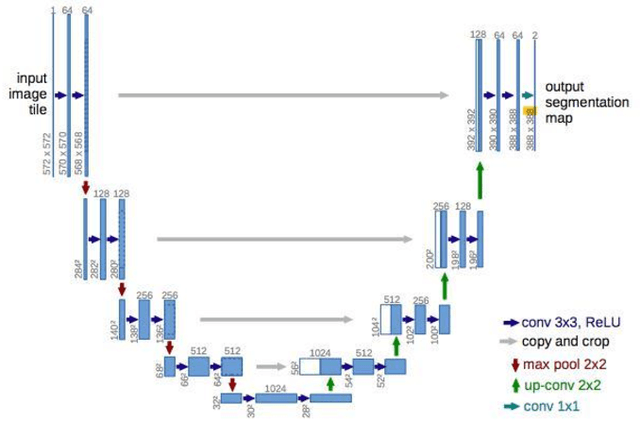

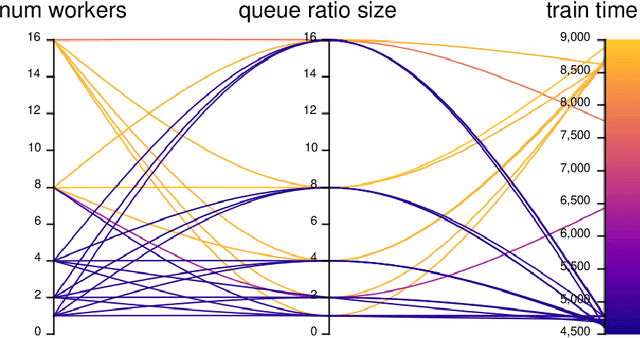
The accurate and consistent border segmentation plays an important role in the tumor volume estimation and its treatment in the field of Medical Image Segmentation. Globally, Lung cancer is one of the leading causes of death and the early detection of lung nodules is essential for the early cancer diagnosis and survival rate of patients. The goal of this study was to demonstrate the feasibility of Deephealth toolkit including PyECVL and PyEDDL libraries to precisely segment lung nodules. Experiments for lung nodules segmentation has been carried out on UniToChest using PyECVL and PyEDDL, for data pre-processing as well as neural network training. The results depict accurate segmentation of lung nodules across a wide diameter range and better accuracy over a traditional detection approach. The datasets and the code used in this paper are publicly available as a baseline reference.
Multi-modal volumetric concept activation to explain detection and classification of metastatic prostate cancer on PSMA-PET/CT
Aug 04, 2022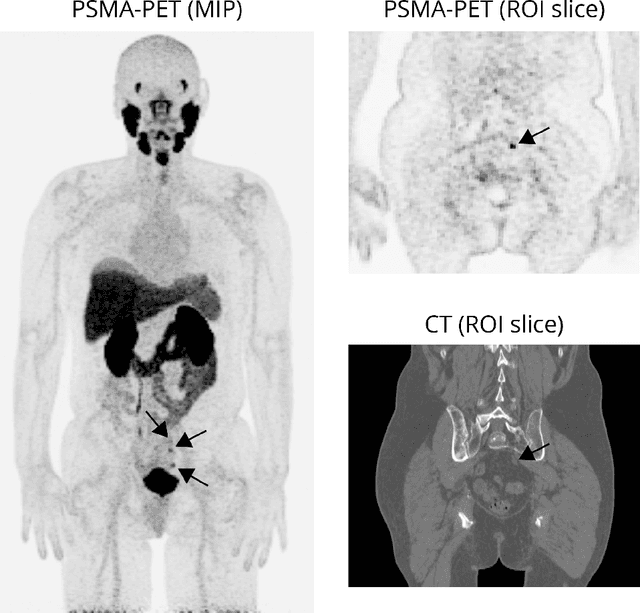
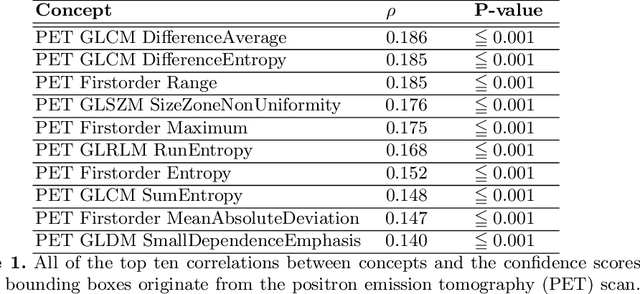
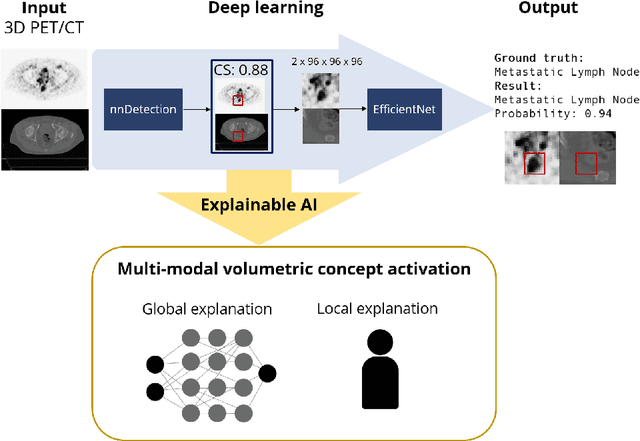
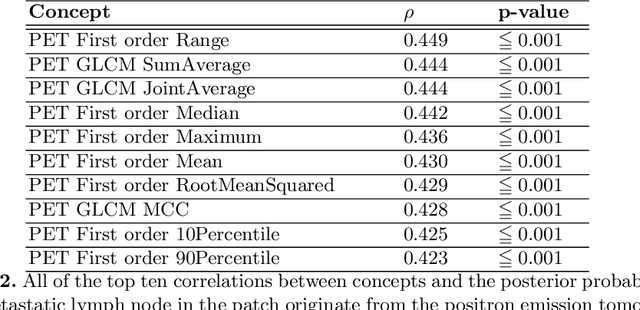
Explainable artificial intelligence (XAI) is increasingly used to analyze the behavior of neural networks. Concept activation uses human-interpretable concepts to explain neural network behavior. This study aimed at assessing the feasibility of regression concept activation to explain detection and classification of multi-modal volumetric data. Proof-of-concept was demonstrated in metastatic prostate cancer patients imaged with positron emission tomography/computed tomography (PET/CT). Multi-modal volumetric concept activation was used to provide global and local explanations. Sensitivity was 80% at 1.78 false positive per patient. Global explanations showed that detection focused on CT for anatomical location and on PET for its confidence in the detection. Local explanations showed promise to aid in distinguishing true positives from false positives. Hence, this study demonstrated feasibility to explain detection and classification of multi-modal volumetric data using regression concept activation.
Memory-aware curriculum federated learning for breast cancer classification
Jul 06, 2021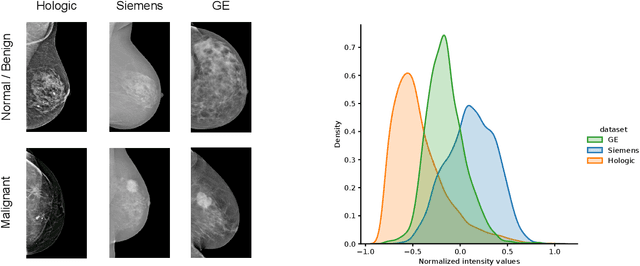
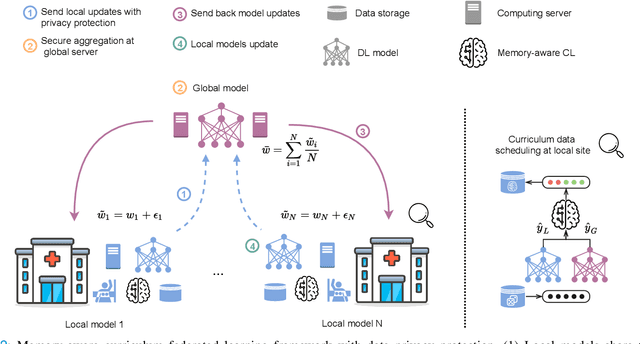
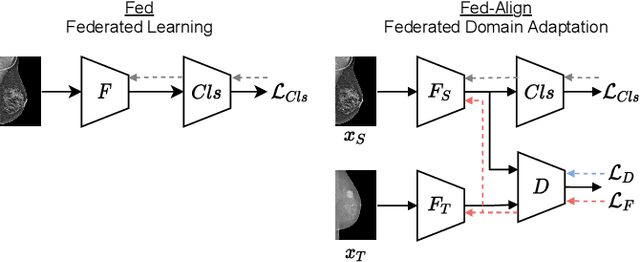
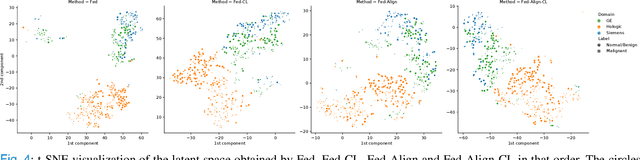
For early breast cancer detection, regular screening with mammography imaging is recommended. Routinary examinations result in datasets with a predominant amount of negative samples. A potential solution to such class-imbalance is joining forces across multiple institutions. Developing a collaborative computer-aided diagnosis system is challenging in different ways. Patient privacy and regulations need to be carefully respected. Data across institutions may be acquired from different devices or imaging protocols, leading to heterogeneous non-IID data. Also, for learning-based methods, new optimization strategies working on distributed data are required. Recently, federated learning has emerged as an effective tool for collaborative learning. In this setting, local models perform computation on their private data to update the global model. The order and the frequency of local updates influence the final global model. Hence, the order in which samples are locally presented to the optimizers plays an important role. In this work, we define a memory-aware curriculum learning method for the federated setting. Our curriculum controls the order of the training samples paying special attention to those that are forgotten after the deployment of the global model. Our approach is combined with unsupervised domain adaptation to deal with domain shift while preserving data privacy. We evaluate our method with three clinical datasets from different vendors. Our results verify the effectiveness of federated adversarial learning for the multi-site breast cancer classification. Moreover, we show that our proposed memory-aware curriculum method is beneficial to further improve classification performance. Our code is publicly available at: https://github.com/ameliajimenez/curriculum-federated-learning.
Towards Learning an Unbiased Classifier from Biased Data via Conditional Adversarial Debiasing
Mar 10, 2021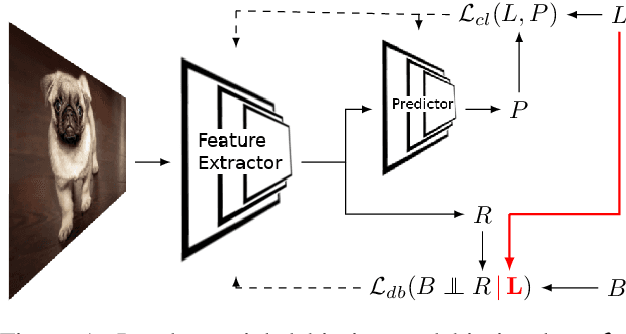
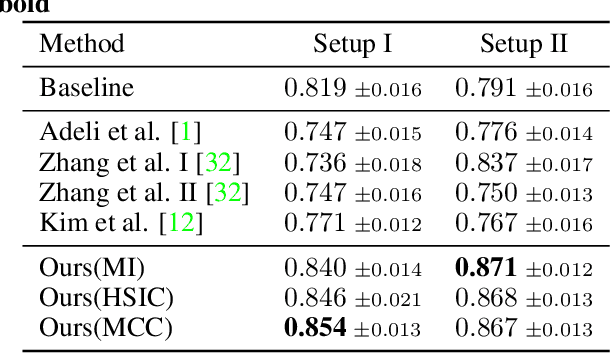
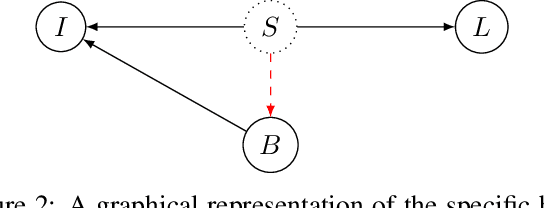
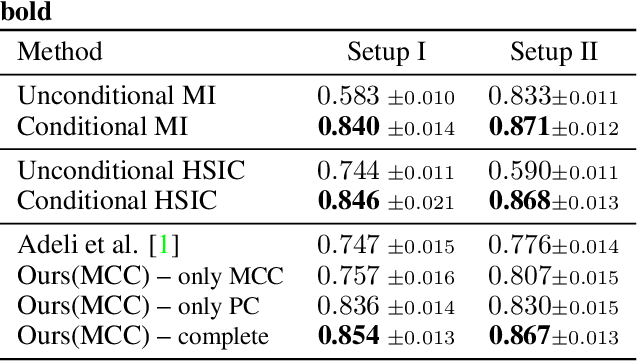
Bias in classifiers is a severe issue of modern deep learning methods, especially for their application in safety- and security-critical areas. Often, the bias of a classifier is a direct consequence of a bias in the training dataset, frequently caused by the co-occurrence of relevant features and irrelevant ones. To mitigate this issue, we require learning algorithms that prevent the propagation of bias from the dataset into the classifier. We present a novel adversarial debiasing method, which addresses a feature that is spuriously connected to the labels of training images but statistically independent of the labels for test images. Thus, the automatic identification of relevant features during training is perturbed by irrelevant features. This is the case in a wide range of bias-related problems for many computer vision tasks, such as automatic skin cancer detection or driver assistance. We argue by a mathematical proof that our approach is superior to existing techniques for the abovementioned bias. Our experiments show that our approach performs better than state-of-the-art techniques on a well-known benchmark dataset with real-world images of cats and dogs.
Distanced LSTM: Time-Distanced Gates in Long Short-Term Memory Models for Lung Cancer Detection
Sep 11, 2019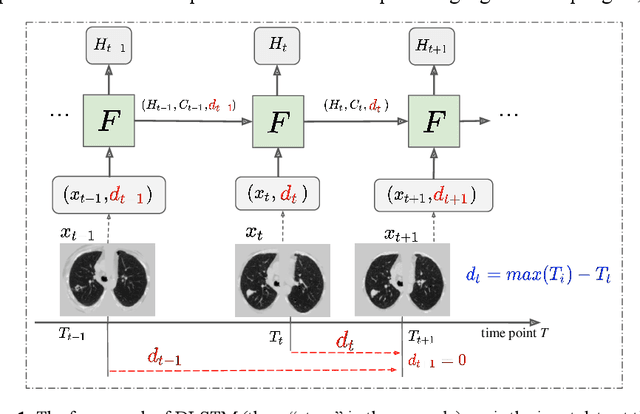
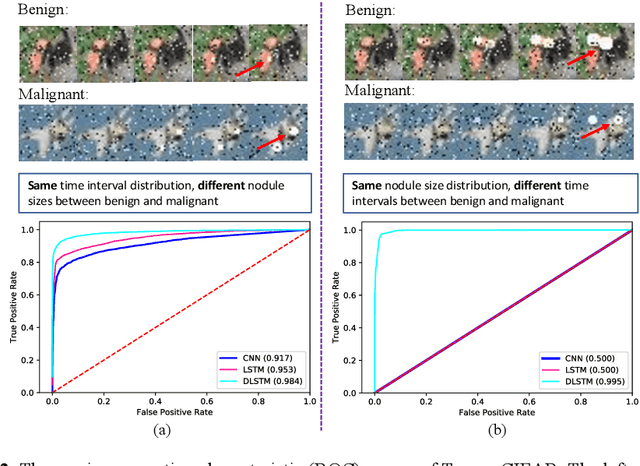
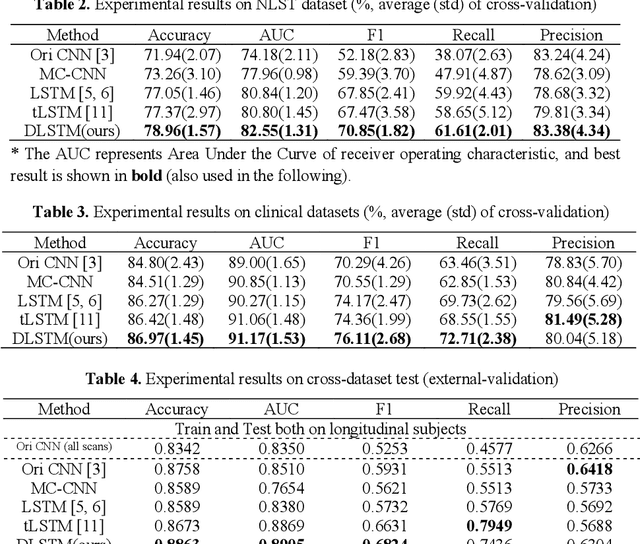
The field of lung nodule detection and cancer prediction has been rapidly developing with the support of large public data archives. Previous studies have largely focused on cross-sectional (single) CT data. Herein, we consider longitudinal data. The Long Short-Term Memory (LSTM) model addresses learning with regularly spaced time points (i.e., equal temporal intervals). However, clinical imaging follows patient needs with often heterogeneous, irregular acquisitions. To model both regular and irregular longitudinal samples, we generalize the LSTM model with the Distanced LSTM (DLSTM) for temporally varied acquisitions. The DLSTM includes a Temporal Emphasis Model (TEM) that enables learning across regularly and irregularly sampled intervals. Briefly, (1) the time intervals between longitudinal scans are modeled explicitly, (2) temporally adjustable forget and input gates are introduced for irregular temporal sampling; and (3) the latest longitudinal scan has an additional emphasis term. We evaluate the DLSTM framework in three datasets including simulated data, 1794 National Lung Screening Trial (NLST) scans, and 1420 clinically acquired data with heterogeneous and irregular temporal accession. The experiments on the first two datasets demonstrate that our method achieves competitive performance on both simulated and regularly sampled datasets (e.g. improve LSTM from 0.6785 to 0.7085 on F1 score in NLST). In external validation of clinically and irregularly acquired data, the benchmarks achieved 0.8350 (CNN feature) and 0.8380 (LSTM) on the area under the ROC curve (AUC) score, while the proposed DLSTM achieves 0.8905.
Feature Fusion of Raman Chemical Imaging and Digital Histopathology using Machine Learning for Prostate Cancer Detection
Jan 18, 2021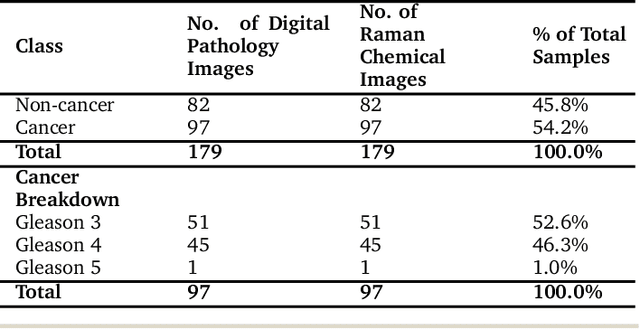
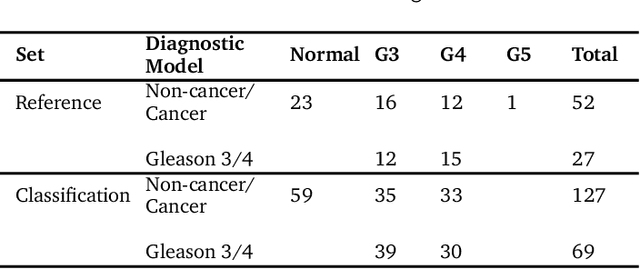
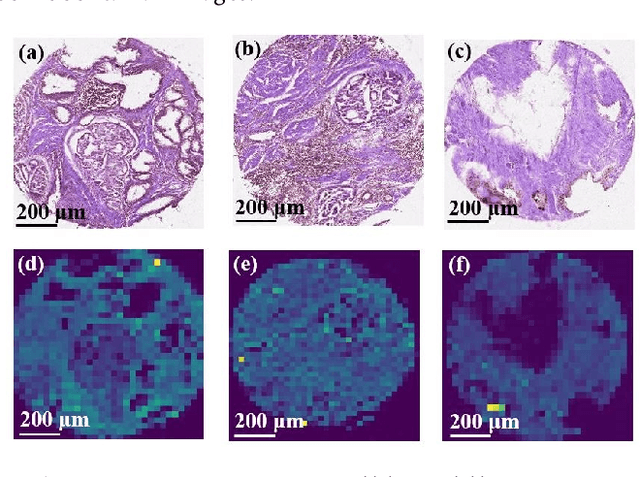
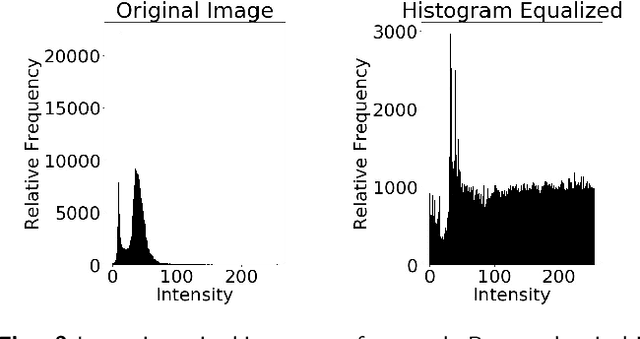

The diagnosis of prostate cancer is challenging due to the heterogeneity of its presentations, leading to the over diagnosis and treatment of non-clinically important disease. Accurate diagnosis can directly benefit a patient's quality of life and prognosis. Towards addressing this issue, we present a learning model for the automatic identification of prostate cancer. While many prostate cancer studies have adopted Raman spectroscopy approaches, none have utilised the combination of Raman Chemical Imaging (RCI) and other imaging modalities. This study uses multimodal images formed from stained Digital Histopathology (DP) and unstained RCI. The approach was developed and tested on a set of 178 clinical samples from 32 patients, containing a range of non-cancerous, Gleason grade 3 (G3) and grade 4 (G4) tissue microarray samples. For each histological sample, there is a pathologist labelled DP - RCI image pair. The hypothesis tested was whether multimodal image models can outperform single modality baseline models in terms of diagnostic accuracy. Binary non-cancer/cancer models and the more challenging G3/G4 differentiation were investigated. Regarding G3/G4 classification, the multimodal approach achieved a sensitivity of 73.8% and specificity of 88.1% while the baseline DP model showed a sensitivity and specificity of 54.1% and 84.7% respectively. The multimodal approach demonstrated a statistically significant 12.7% AUC advantage over the baseline with a value of 85.8% compared to 73.1%, also outperforming models based solely on RCI and median Raman spectra. Feature fusion of DP and RCI does not improve the more trivial task of tumour identification but does deliver an observed advantage in G3/G4 discrimination. Building on these promising findings, future work could include the acquisition of larger datasets for enhanced model generalization.
RCNN for Region of Interest Detection in Whole Slide Images
Sep 18, 2020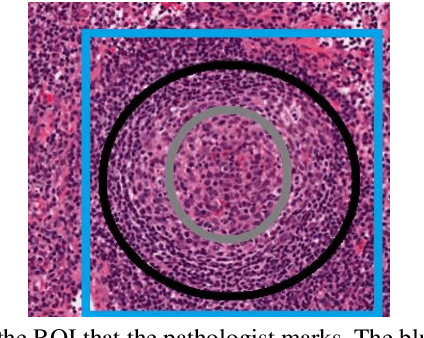
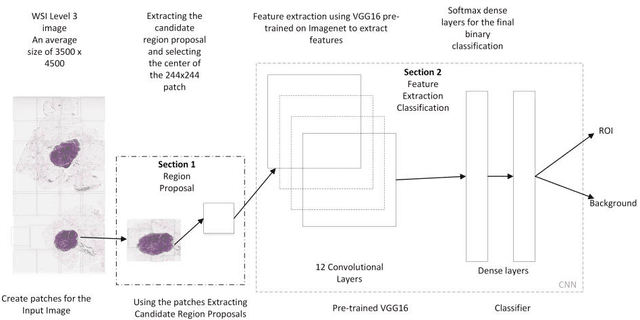
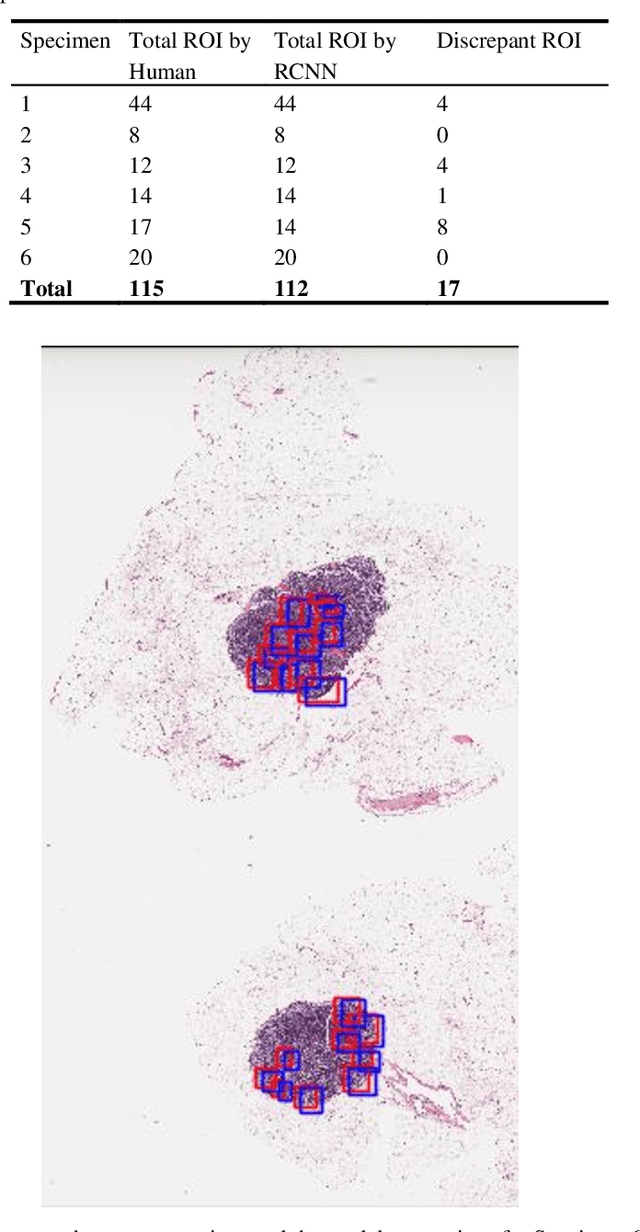
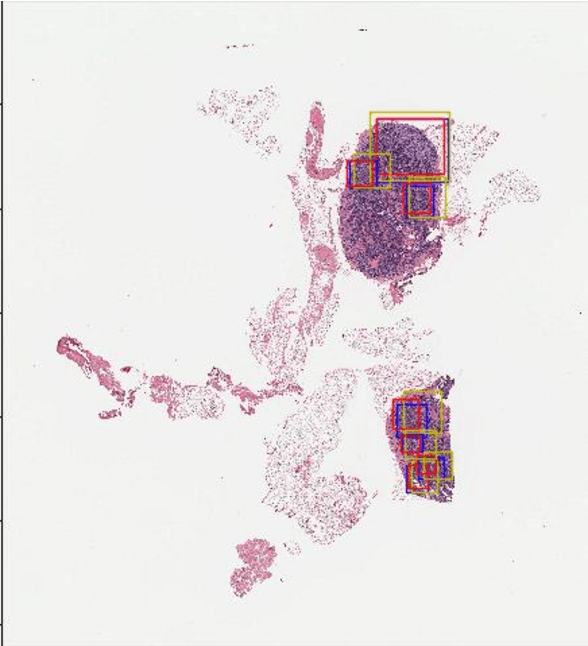
Digital pathology has attracted significant attention in recent years. Analysis of Whole Slide Images (WSIs) is challenging because they are very large, i.e., of Giga-pixel resolution. Identifying Regions of Interest (ROIs) is the first step for pathologists to analyse further the regions of diagnostic interest for cancer detection and other anomalies. In this paper, we investigate the use of RCNN, which is a deep machine learning technique, for detecting such ROIs only using a small number of labelled WSIs for training. For experimentation, we used real WSIs from a public hospital pathology service in Western Australia. We used 60 WSIs for training the RCNN model and another 12 WSIs for testing. The model was further tested on a new set of unseen WSIs. The results show that RCNN can be effectively used for ROI detection from WSIs.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge