"cancer detection": models, code, and papers
Sources of performance variability in deep learning-based polyp detection
Nov 17, 2022
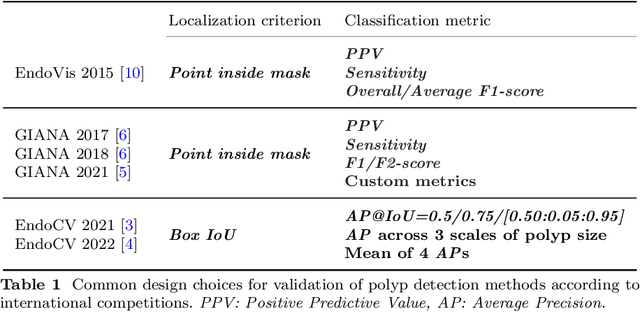
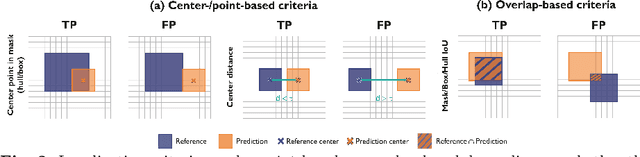
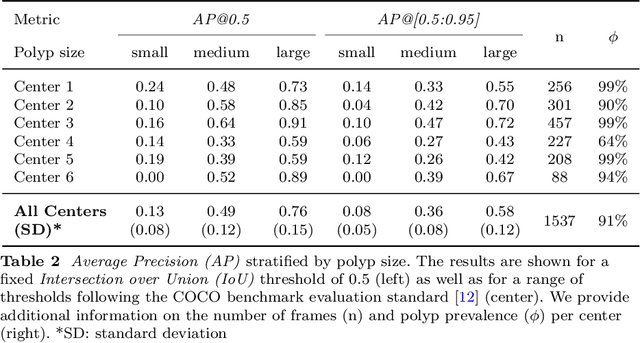
Validation metrics are a key prerequisite for the reliable tracking of scientific progress and for deciding on the potential clinical translation of methods. While recent initiatives aim to develop comprehensive theoretical frameworks for understanding metric-related pitfalls in image analysis problems, there is a lack of experimental evidence on the concrete effects of common and rare pitfalls on specific applications. We address this gap in the literature in the context of colon cancer screening. Our contribution is twofold. Firstly, we present the winning solution of the Endoscopy computer vision challenge (EndoCV) on colon cancer detection, conducted in conjunction with the IEEE International Symposium on Biomedical Imaging (ISBI) 2022. Secondly, we demonstrate the sensitivity of commonly used metrics to a range of hyperparameters as well as the consequences of poor metric choices. Based on comprehensive validation studies performed with patient data from six clinical centers, we found all commonly applied object detection metrics to be subject to high inter-center variability. Furthermore, our results clearly demonstrate that the adaptation of standard hyperparameters used in the computer vision community does not generally lead to the clinically most plausible results. Finally, we present localization criteria that correspond well to clinical relevance. Our work could be a first step towards reconsidering common validation strategies in automatic colon cancer screening applications.
CT Multi-Task Learning with a Large Image-Text (LIT) Model
Apr 03, 2023Large language models (LLM) not only empower multiple language tasks but also serve as a general interface across different spaces. Up to now, it has not been demonstrated yet how to effectively translate the successes of LLMs in the computer vision field to the medical imaging field which involves high-dimensional and multi-modal medical images. In this paper, we report a feasibility study of building a multi-task CT large image-text (LIT) model for lung cancer diagnosis by combining an LLM and a large image model (LIM). Specifically, the LLM and LIM are used as encoders to perceive multi-modal information under task-specific text prompts, which synergizes multi-source information and task-specific and patient-specific priors for optimized diagnostic performance. The key components of our LIT model and associated techniques are evaluated with an emphasis on 3D lung CT analysis. Our initial results show that the LIT model performs multiple medical tasks well, including lung segmentation, lung nodule detection, and lung cancer classification. Active efforts are in progress to develop large image-language models for superior medical imaging in diverse applications and optimal patient outcomes.
Multimodal Data Integration for Oncology in the Era of Deep Neural Networks: A Review
Mar 11, 2023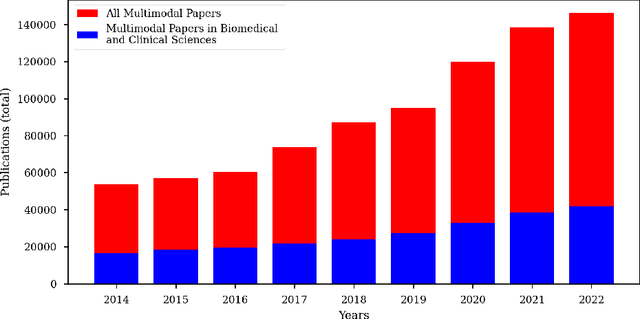
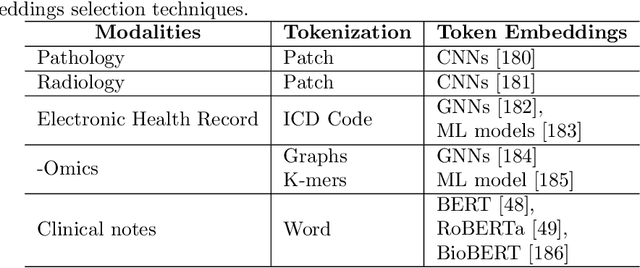
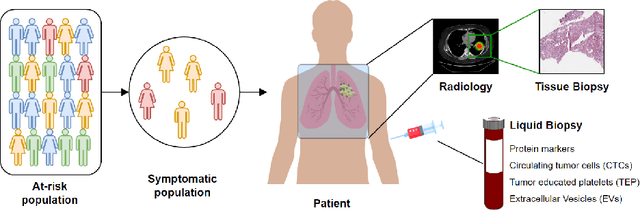
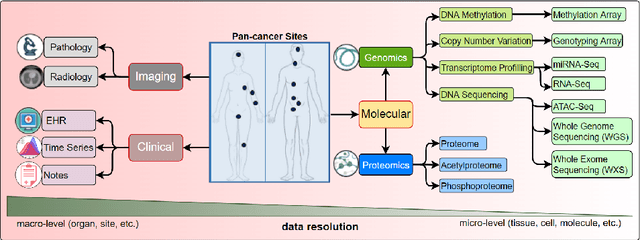
Cancer has relational information residing at varying scales, modalities, and resolutions of the acquired data, such as radiology, pathology, genomics, proteomics, and clinical records. Integrating diverse data types can improve the accuracy and reliability of cancer diagnosis and treatment. There can be disease-related information that is too subtle for humans or existing technological tools to discern visually. Traditional methods typically focus on partial or unimodal information about biological systems at individual scales and fail to encapsulate the complete spectrum of the heterogeneous nature of data. Deep neural networks have facilitated the development of sophisticated multimodal data fusion approaches that can extract and integrate relevant information from multiple sources. Recent deep learning frameworks such as Graph Neural Networks (GNNs) and Transformers have shown remarkable success in multimodal learning. This review article provides an in-depth analysis of the state-of-the-art in GNNs and Transformers for multimodal data fusion in oncology settings, highlighting notable research studies and their findings. We also discuss the foundations of multimodal learning, inherent challenges, and opportunities for integrative learning in oncology. By examining the current state and potential future developments of multimodal data integration in oncology, we aim to demonstrate the promising role that multimodal neural networks can play in cancer prevention, early detection, and treatment through informed oncology practices in personalized settings.
Improving Prostate Cancer Detection with Breast Histopathology Images
Mar 14, 2019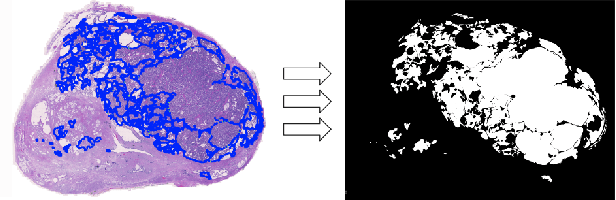
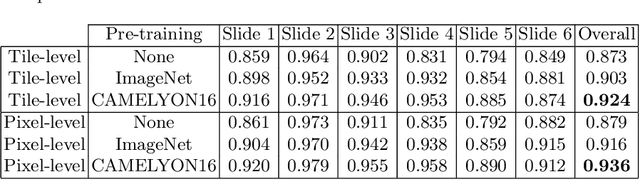
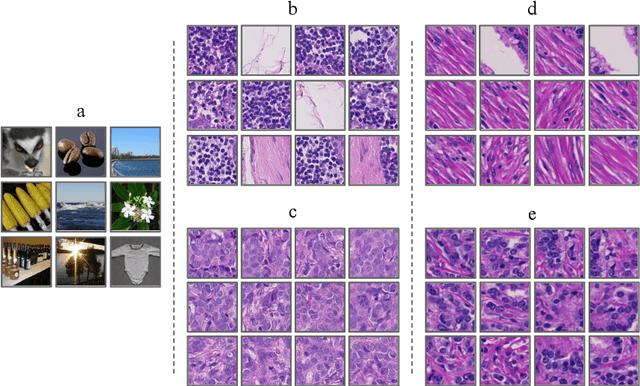
Deep neural networks have introduced significant advancements in the field of machine learning-based analysis of digital pathology images including prostate tissue images. With the help of transfer learning, classification and segmentation performance of neural network models have been further increased. However, due to the absence of large, extensively annotated, publicly available prostate histopathology datasets, several previous studies employ datasets from well-studied computer vision tasks such as ImageNet dataset. In this work, we propose a transfer learning scheme from breast histopathology images to improve prostate cancer detection performance. We validate our approach on annotated prostate whole slide images by using a publicly available breast histopathology dataset as pre-training. We show that the proposed cross-cancer approach outperforms transfer learning from ImageNet dataset.
New pyramidal hybrid textural and deep features based automatic skin cancer classification model: Ensemble DarkNet and textural feature extractor
Mar 28, 2022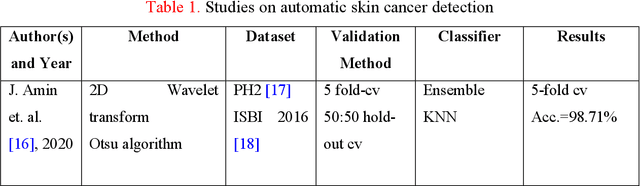
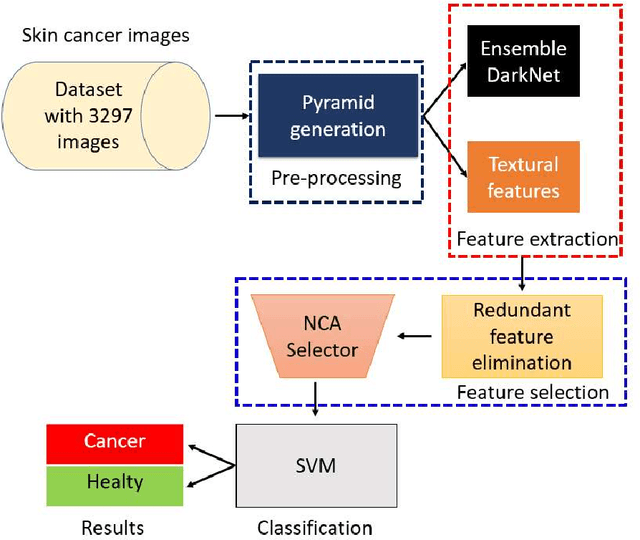
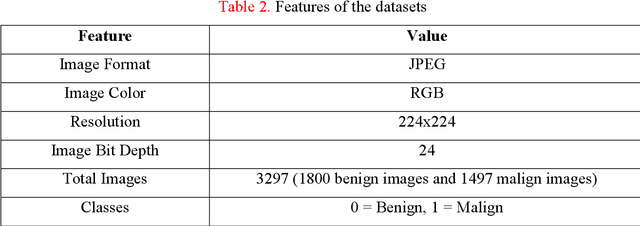
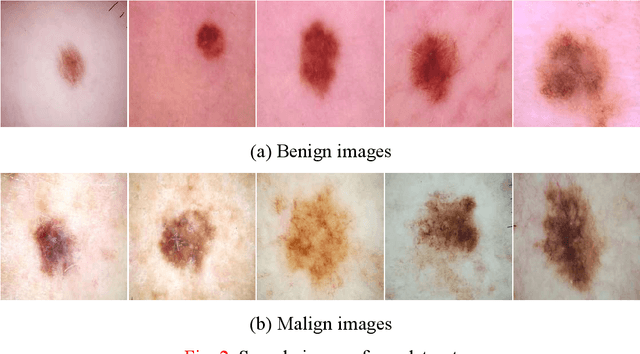
Background: Skin cancer is one of the widely seen cancer worldwide and automatic classification of skin cancer can be benefited dermatology clinics for an accurate diagnosis. Hence, a machine learning-based automatic skin cancer detection model must be developed. Material and Method: This research interests to overcome automatic skin cancer detection problem. A colored skin cancer image dataset is used. This dataset contains 3297 images with two classes. An automatic multilevel textural and deep features-based model is presented. Multilevel fuse feature generation using discrete wavelet transform (DWT), local phase quantization (LPQ), local binary pattern (LBP), pre-trained DarkNet19, and DarkNet53 are utilized to generate features of the skin cancer images, top 1000 features are selected threshold value-based neighborhood component analysis (NCA). The chosen top 1000 features are classified using the 10-fold cross-validation technique. Results: To obtain results, ten-fold cross-validation is used and 91.54% classification accuracy results are obtained by using the recommended pyramidal hybrid feature generator and NCA selector-based model. Further, various training and testing separation ratios (90:10, 80:20, 70:30, 60:40, 50:50) are used and the maximum classification rate is calculated as 95.74% using the 90:10 separation ratio. Conclusions: The findings and accuracies calculated are denoted that this model can be used in dermatology and pathology clinics to simplify the skin cancer detection process and help physicians.
Breast cancer detection using artificial intelligence techniques: A systematic literature review
Mar 08, 2022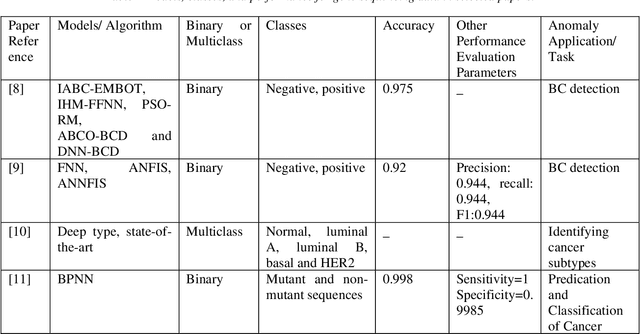
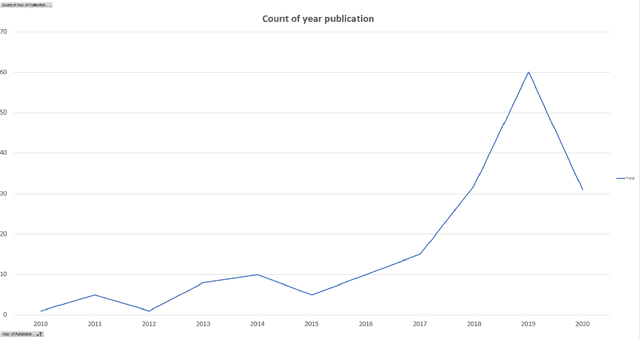
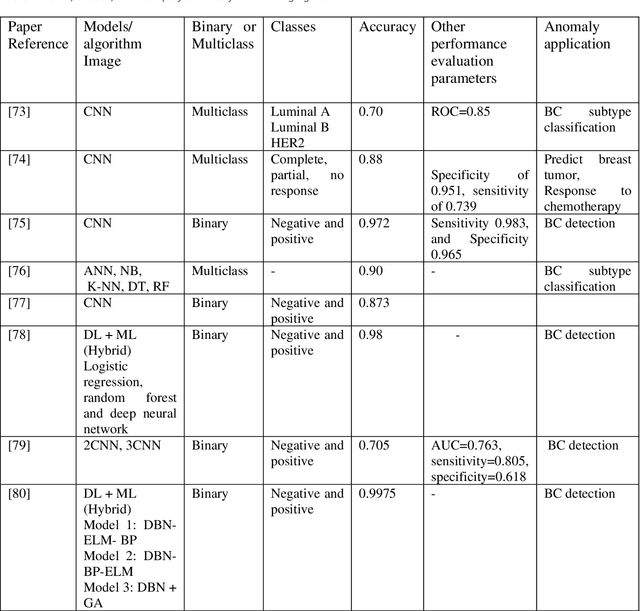
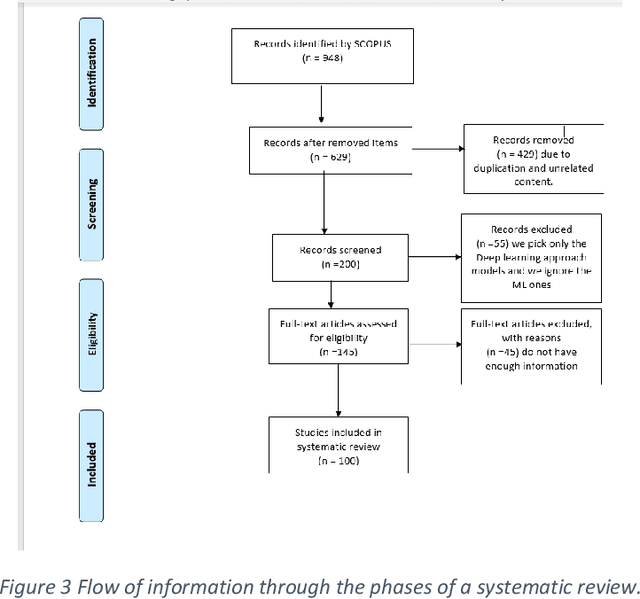
Cancer is one of the most dangerous diseases to humans, and yet no permanent cure has been developed for it. Breast cancer is one of the most common cancer types. According to the National Breast Cancer foundation, in 2020 alone, more than 276,000 new cases of invasive breast cancer and more than 48,000 non-invasive cases were diagnosed in the US. To put these figures in perspective, 64% of these cases are diagnosed early in the disease's cycle, giving patients a 99% chance of survival. Artificial intelligence and machine learning have been used effectively in detection and treatment of several dangerous diseases, helping in early diagnosis and treatment, and thus increasing the patient's chance of survival. Deep learning has been designed to analyze the most important features affecting detection and treatment of serious diseases. For example, breast cancer can be detected using genes or histopathological imaging. Analysis at the genetic level is very expensive, so histopathological imaging is the most common approach used to detect breast cancer. In this research work, we systematically reviewed previous work done on detection and treatment of breast cancer using genetic sequencing or histopathological imaging with the help of deep learning and machine learning. We also provide recommendations to researchers who will work in this field
Graph Neural Networks for Breast Cancer Data Integration
Nov 28, 2022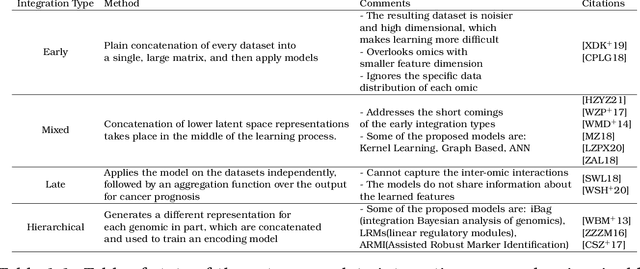
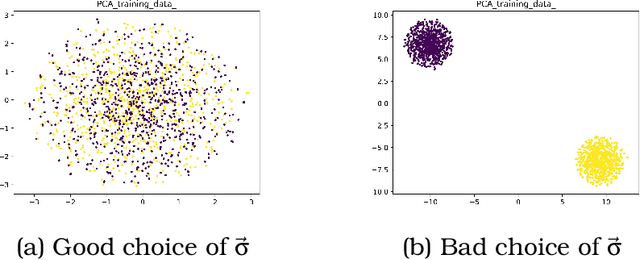
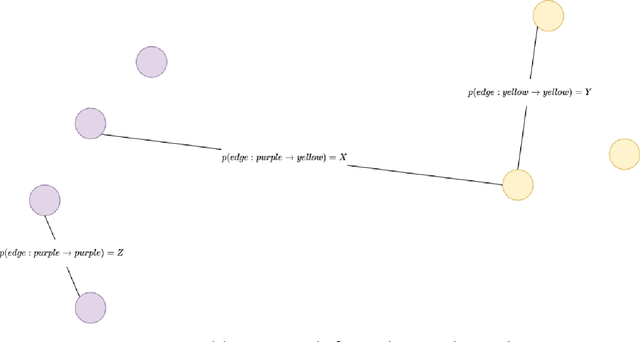
International initiatives such as METABRIC (Molecular Taxonomy of Breast Cancer International Consortium) have collected several multigenomic and clinical data sets to identify the undergoing molecular processes taking place throughout the evolution of various cancers. Numerous Machine Learning and statistical models have been designed and trained to analyze these types of data independently, however, the integration of such differently shaped and sourced information streams has not been extensively studied. To better integrate these data sets and generate meaningful representations that can ultimately be leveraged for cancer detection tasks could lead to giving well-suited treatments to patients. Hence, we propose a novel learning pipeline comprising three steps - the integration of cancer data modalities as graphs, followed by the application of Graph Neural Networks in an unsupervised setting to generate lower-dimensional embeddings from the combined data, and finally feeding the new representations on a cancer sub-type classification model for evaluation. The graph construction algorithms are described in-depth as METABRIC does not store relationships between the patient modalities, with a discussion of their influence over the quality of the generated embeddings. We also present the models used to generate the lower-latent space representations: Graph Neural Networks, Variational Graph Autoencoders and Deep Graph Infomax. In parallel, the pipeline is tested on a synthetic dataset to demonstrate that the characteristics of the underlying data, such as homophily levels, greatly influence the performance of the pipeline, which ranges between 51\% to 98\% accuracy on artificial data, and 13\% and 80\% on METABRIC. This project has the potential to improve cancer data understanding and encourages the transition of regular data sets to graph-shaped data.
Context-Aware Transformers For Spinal Cancer Detection and Radiological Grading
Jun 27, 2022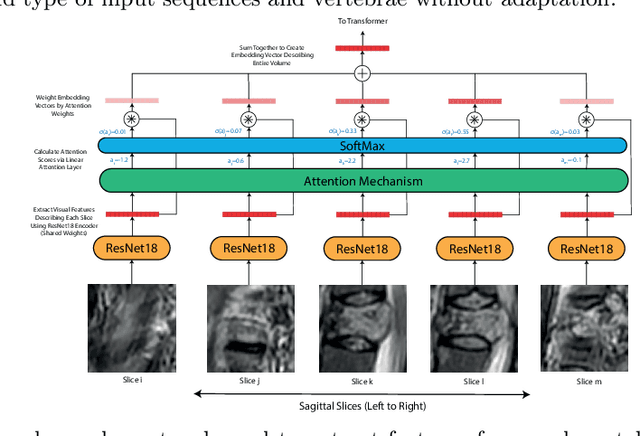
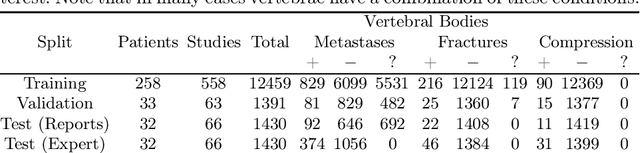

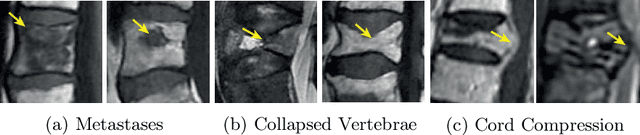
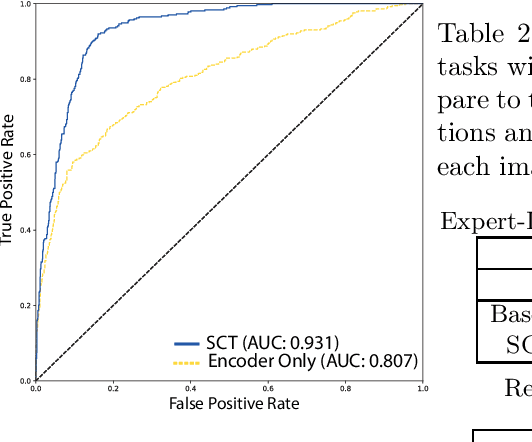
This paper proposes a novel transformer-based model architecture for medical imaging problems involving analysis of vertebrae. It considers two applications of such models in MR images: (a) detection of spinal metastases and the related conditions of vertebral fractures and metastatic cord compression, (b) radiological grading of common degenerative changes in intervertebral discs. Our contributions are as follows: (i) We propose a Spinal Context Transformer (SCT), a deep-learning architecture suited for the analysis of repeated anatomical structures in medical imaging such as vertebral bodies (VBs). Unlike previous related methods, SCT considers all VBs as viewed in all available image modalities together, making predictions for each based on context from the rest of the spinal column and all available imaging modalities. (ii) We apply the architecture to a novel and important task: detecting spinal metastases and the related conditions of cord compression and vertebral fractures/collapse from multi-series spinal MR scans. This is done using annotations extracted from free-text radiological reports as opposed to bespoke annotation. However, the resulting model shows strong agreement with vertebral-level bespoke radiologist annotations on the test set. (iii) We also apply SCT to an existing problem: radiological grading of inter-vertebral discs (IVDs) in lumbar MR scans for common degenerative changes.We show that by considering the context of vertebral bodies in the image, SCT improves the accuracy for several gradings compared to previously published model.
Transfer Learning by Cascaded Network to identify and classify lung nodules for cancer detection
Sep 24, 2020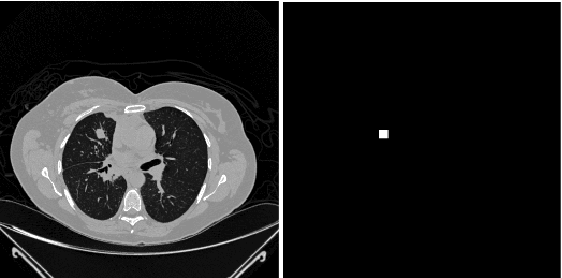

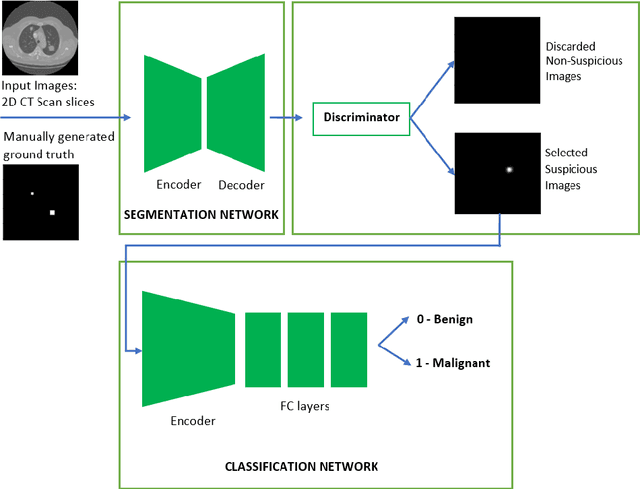
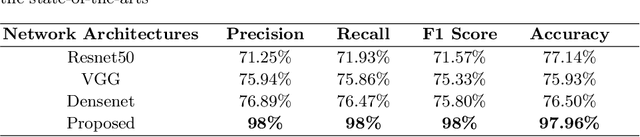
Lung cancer is one of the most deadly diseases in the world. Detecting such tumors at an early stage can be a tedious task. Existing deep learning architecture for lung nodule identification used complex architecture with large number of parameters. This study developed a cascaded architecture which can accurately segment and classify the benign or malignant lung nodules on computed tomography (CT) images. The main contribution of this study is to introduce a segmentation network where the first stage trained on a public data set can help to recognize the images which included a nodule from any data set by means of transfer learning. And the segmentation of a nodule improves the second stage to classify the nodules into benign and malignant. The proposed architecture outperformed the conventional methods with an area under curve value of 95.67\%. The experimental results showed that the classification accuracy of 97.96\% of our proposed architecture outperformed other simple and complex architectures in classifying lung nodules for lung cancer detection.
A Comprehensive Study of Data Augmentation Strategies for Prostate Cancer Detection in Diffusion-weighted MRI using Convolutional Neural Networks
Jun 01, 2020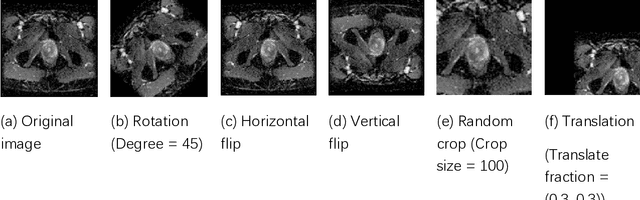
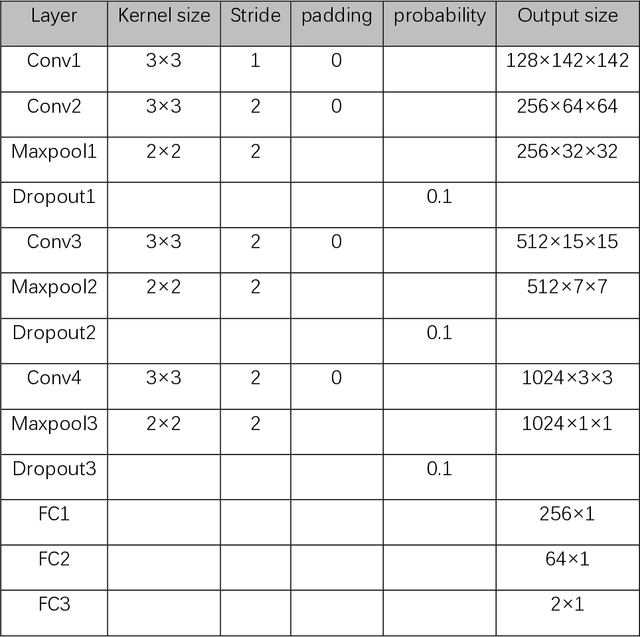
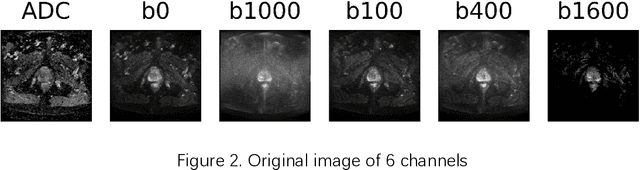
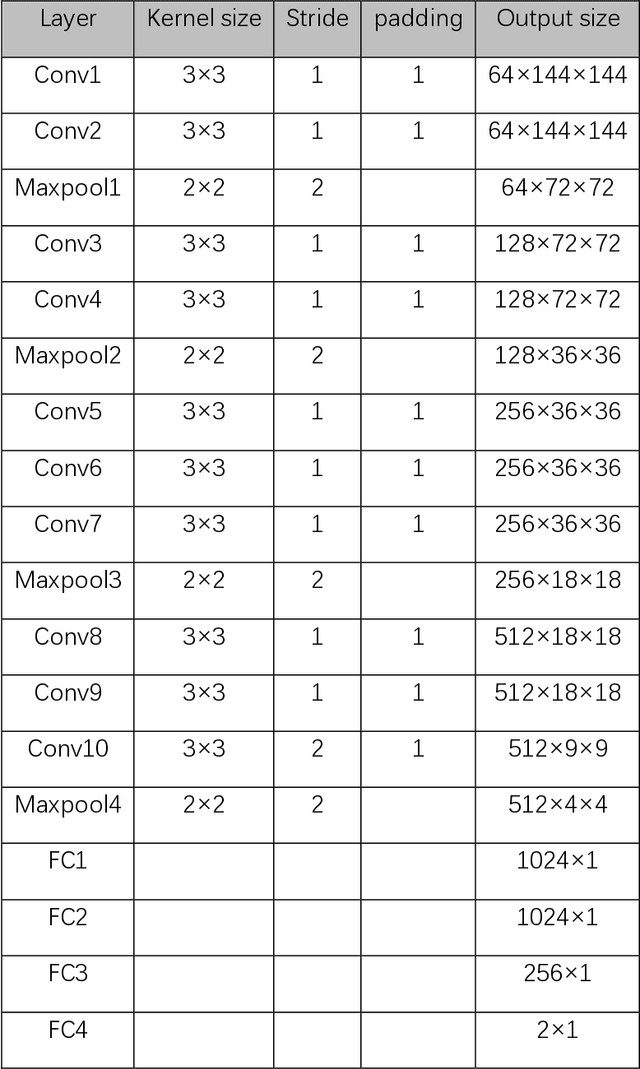
Data augmentation refers to a group of techniques whose goal is to battle limited amount of available data to improve model generalization and push sample distribution toward the true distribution. While different augmentation strategies and their combinations have been investigated for various computer vision tasks in the context of deep learning, a specific work in the domain of medical imaging is rare and to the best of our knowledge, there has been no dedicated work on exploring the effects of various augmentation methods on the performance of deep learning models in prostate cancer detection. In this work, we have statically applied five most frequently used augmentation techniques (random rotation, horizontal flip, vertical flip, random crop, and translation) to prostate Diffusion-weighted Magnetic Resonance Imaging training dataset of 217 patients separately and evaluated the effect of each method on the accuracy of prostate cancer detection. The augmentation algorithms were applied independently to each data channel and a shallow as well as a deep Convolutional Neural Network (CNN) were trained on the five augmented sets separately. We used Area Under Receiver Operating Characteristic (ROC) curve (AUC) to evaluate the performance of the trained CNNs on a separate test set of 95 patients, using a validation set of 102 patients for finetuning. The shallow network outperformed the deep network with the best 2D slice-based AUC of 0.85 obtained by the rotation method.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge