"cancer detection": models, code, and papers
Deep Integrated Pipeline of Segmentation Leading to Classification for Automated Detection of Breast Cancer from Breast Ultrasound Images
Oct 26, 2021
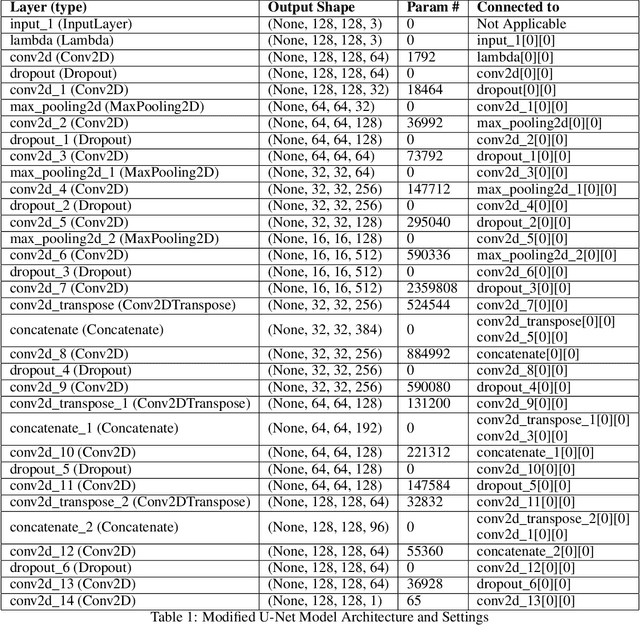
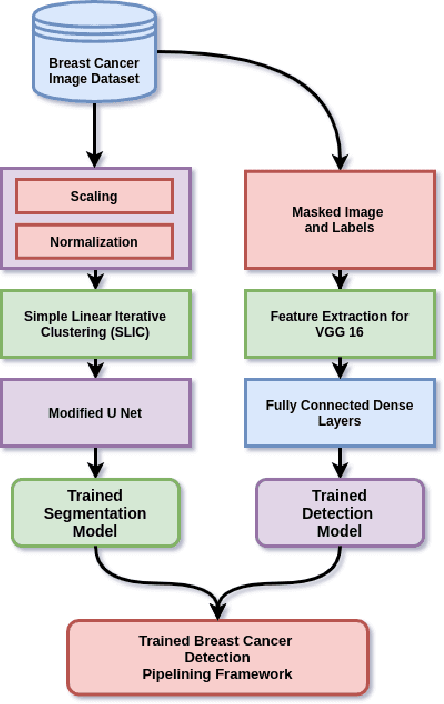
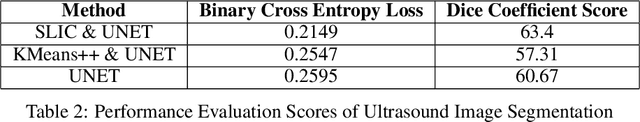
Breast cancer has become a symbol of tremendous concern in the modern world, as it is one of the major causes of cancer mortality worldwide. In this concern, many people are frequently screening for breast cancer in order to be identified early and avert mortality from the disease by receiving treatment. Breast Ultrasonography Images are frequently utilized by doctors to diagnose breast cancer at an early stage. However, the complex artifacts and heavily noised Breast Ultrasonography Images make detecting Breast Cancer a tough challenge. Furthermore, the ever-increasing number of patients being screened for Breast Cancer necessitates the use of automated Computer Aided Technology for high accuracy diagnosis at a cheap cost and in a short period of time. The current progress of Artificial Intelligence (AI) in the fields of Medical Image Analysis and Health Care is a boon to humanity. In this study, we have proposed a compact integrated automated pipelining framework which integrates ultrasonography image preprocessing with Simple Linear Iterative Clustering (SLIC) to tackle the complex artifact of Breast Ultrasonography Images complementing semantic segmentation with Modified U-Net leading to Breast Tumor classification with robust feature extraction using a transfer learning approach with pretrained VGG 16 model and densely connected neural network architecture. The proposed automated pipeline can be effectively implemented to assist medical practitioners in making more accurate and timely diagnoses of breast cancer.
Using convolutional neural networks for the classification of breast cancer images
Aug 31, 2021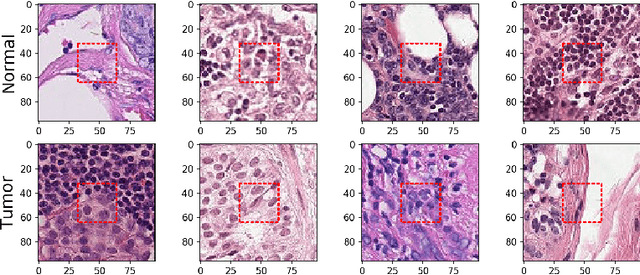
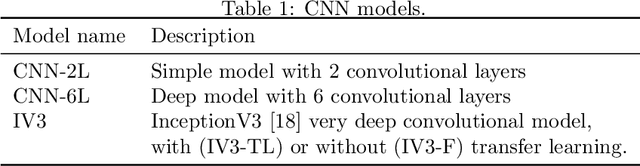

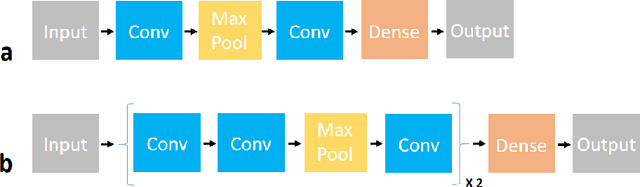
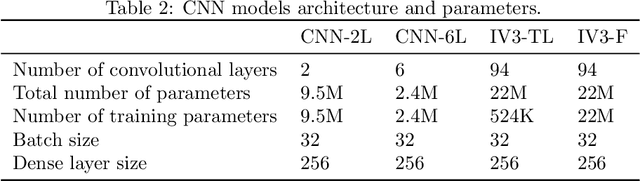
In this study I compare different architectures of convolutional neural networks and different hardware acceleration devices for the detection of breast cancer metastasis tissue from microscopic images of sentinel lymph nodes. Convolutional models with increasing depth are trained and tested on a public data set of more than 300,000 images of lymph node tissue, on three different hardware acceleration cards, using an off-the-shelf deep learning framework. The impact of transfer learning, data augmentation and hyperparameters fine-tuning are also tested. Hardware acceleration device performance can improve training time by a factor of five to seven, depending on the model used. On the other hand, increasing convolutional depth will augment the training time by a factor of four to six times, depending on the acceleration device used. Increasing the depth of the model, as could be expected, clearly improves performance, while data augmentation and transfer learning do not. Fine-tuning the hyperparameters of the model notably improves the results, with the best model showing a performance comparable to state-of-the-art models.
Breast Cancer Detection Using Multilevel Thresholding
Nov 03, 2009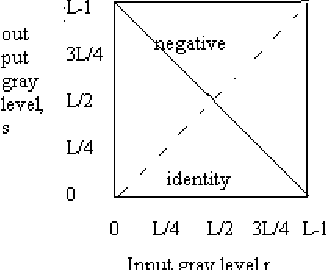
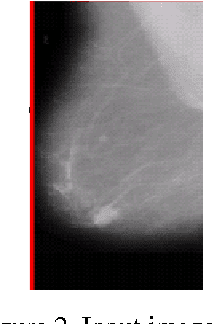

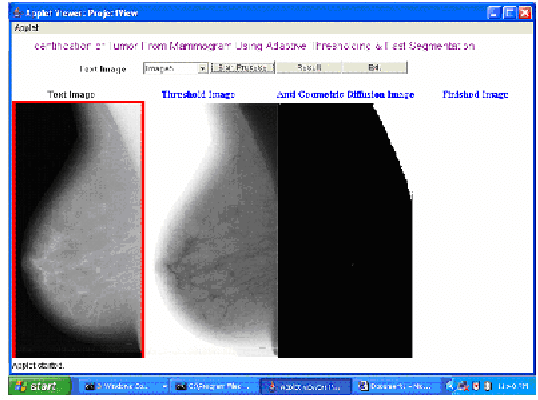
This paper presents an algorithm which aims to assist the radiologist in identifying breast cancer at its earlier stages. It combines several image processing techniques like image negative, thresholding and segmentation techniques for detection of tumor in mammograms. The algorithm is verified by using mammograms from Mammographic Image Analysis Society. The results obtained by applying these techniques are described.
* 5 pages IEEE format, International Journal of Computer Science and Information Security, IJCSIS 2009, ISSN 1947 5500, Impact Factor 0.423, http://sites.google.com/site/ijcsis/
Anatomical and Diagnostic Bayesian Segmentation in Prostate MRI $-$Should Different Clinical Objectives Mandate Different Loss Functions?
Oct 25, 2021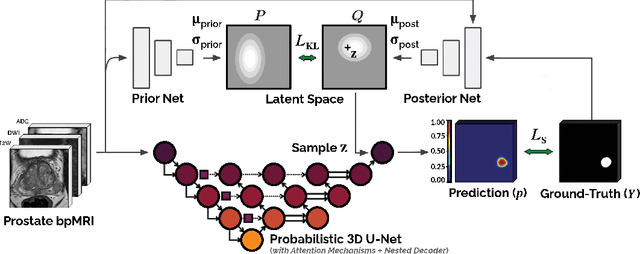
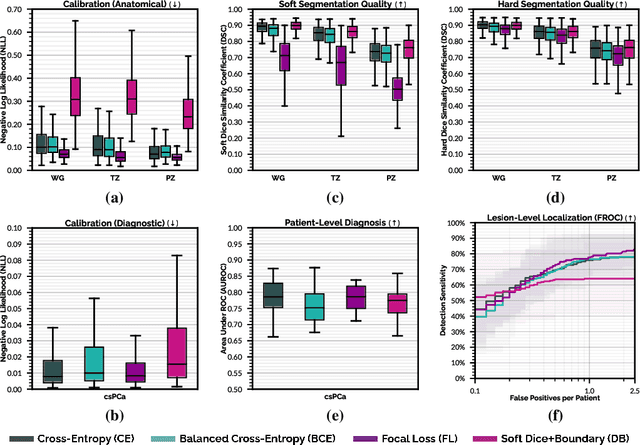
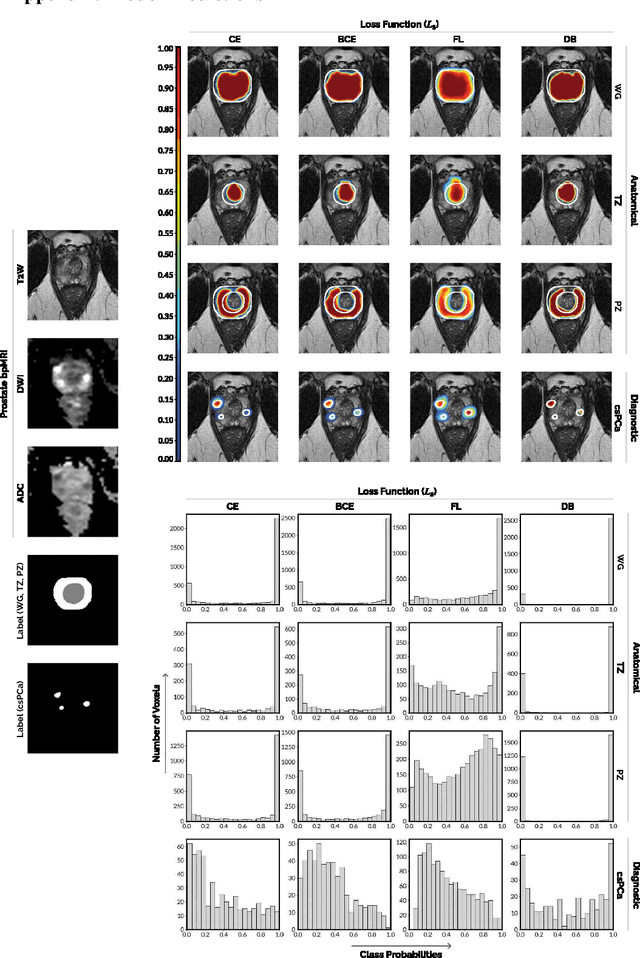
We hypothesize that probabilistic voxel-level classification of anatomy and malignancy in prostate MRI, although typically posed as near-identical segmentation tasks via U-Nets, require different loss functions for optimal performance due to inherent differences in their clinical objectives. We investigate distribution, region and boundary-based loss functions for both tasks across 200 patient exams from the publicly-available ProstateX dataset. For evaluation, we conduct a thorough comparative analysis of model predictions and calibration, measured with respect to multi-class volume segmentation of the prostate anatomy (whole-gland, transitional zone, peripheral zone), as well as, patient-level diagnosis and lesion-level detection of clinically significant prostate cancer. Notably, we find that distribution-based loss functions (in particular, focal loss) are well-suited for diagnostic or panoptic segmentation tasks such as lesion detection, primarily due to their implicit property of inducing better calibration. Meanwhile, (with the exception of focal loss) both distribution and region/boundary-based loss functions perform equally well for anatomical or semantic segmentation tasks, such as quantification of organ shape, size and boundaries.
B-Spline CNNs on Lie Groups
Sep 26, 2019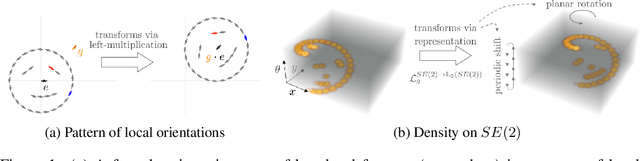
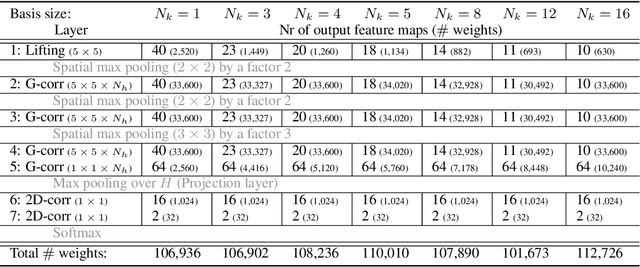
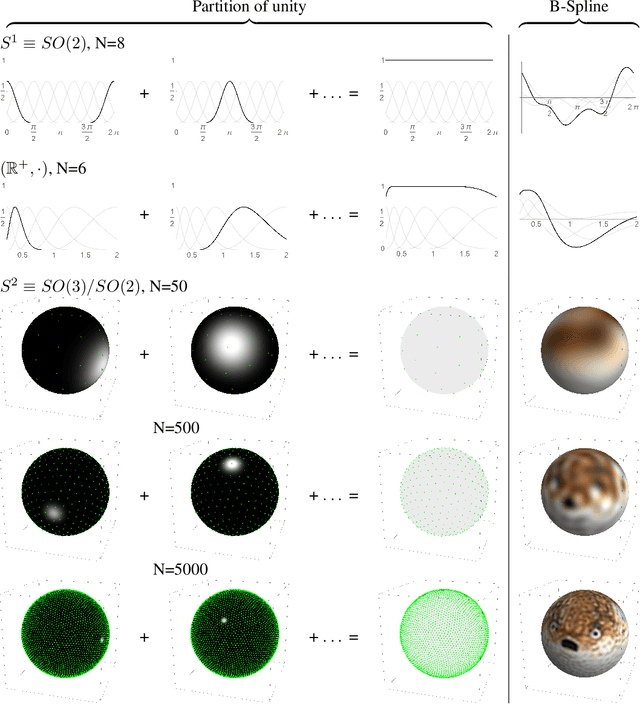
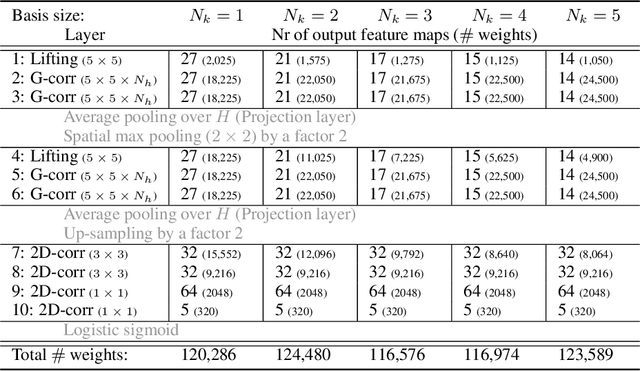
Group convolutional neural networks (G-CNNs) can be used to improve classical CNNs by equipping them with the geometric structure of groups. Central in the success of G-CNNs is the lifting of feature maps to higher dimensional disentangled representations, in which data characteristics are effectively learned, geometric data-augmentations are made obsolete, and predictable behavior under geometric transformations (equivariance) is guaranteed via group theory. Currently, however, the practical implementations of G-CNNs are limited to either discrete groups (that leave the grid intact) or continuous compact groups such as rotations (that enable the use of Fourier theory). In this paper we lift these limitations and propose a modular framework for the design and implementation of G-CNNs for arbitrary Lie groups. In our approach the differential structure of Lie groups is used to expand convolution kernels in a generic basis of B-splines that is defined on the Lie algebra. This leads to a flexible framework that enables localized, atrous, and deformable convolutions in G-CNNs by means of respectively localized, sparse and non-uniform B-spline expansions. The impact and potential of our approach is studied on two benchmark datasets: cancer detection in histopathology slides in which rotation equivariance plays a key role and facial landmark localization in which scale equivariance is important. In both cases, G-CNN architectures outperform their classical 2D counterparts and the added value of atrous and localized group convolutions is studied in detail.
Machine Learning Characterization of Cancer Patients-Derived Extracellular Vesicles using Vibrational Spectroscopies
Aug 25, 2021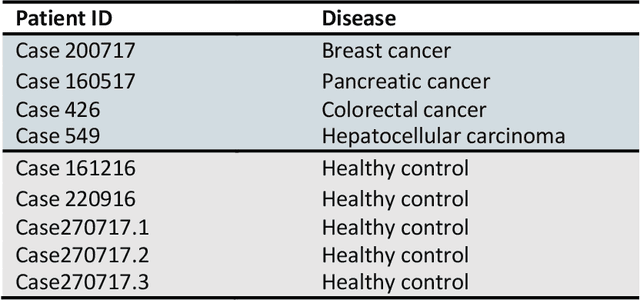
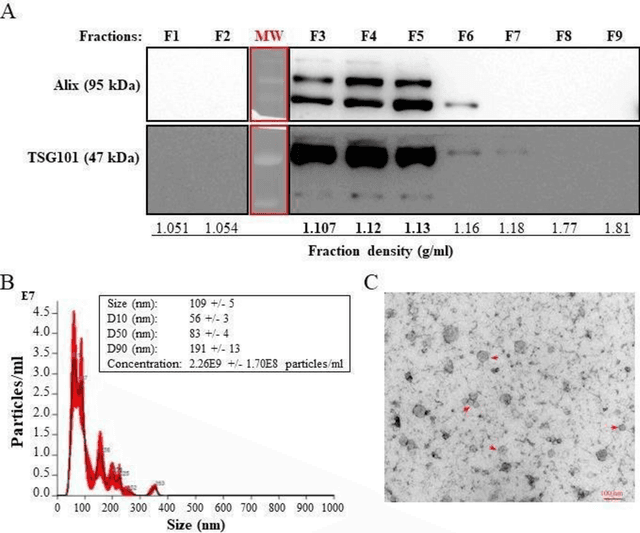
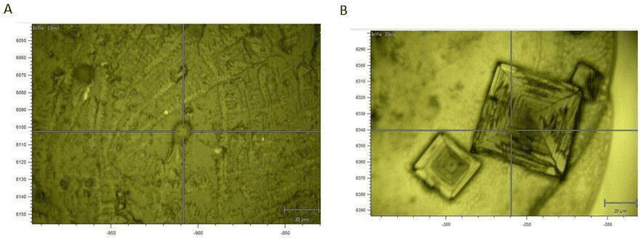
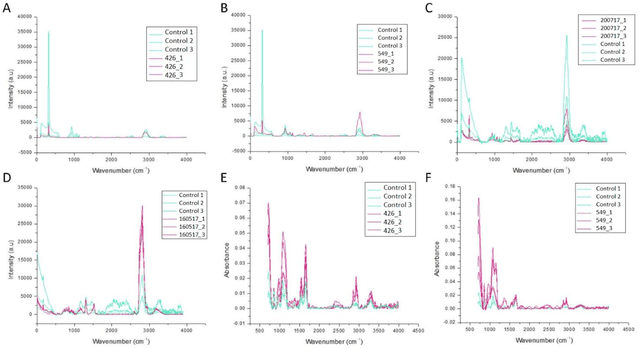
The early detection of cancer is a challenging problem in medicine. The blood sera of cancer patients are enriched with heterogeneous secretory lipid bound extracellular vesicles (EVs), which present a complex repertoire of information and biomarkers, representing their cell of origin, that are being currently studied in the field of liquid biopsy and cancer screening. Vibrational spectroscopies provide non-invasive approaches for the assessment of structural and biophysical properties in complex biological samples. In this study, multiple Raman spectroscopy measurements were performed on the EVs extracted from the blood sera of 9 patients consisting of four different cancer subtypes (colorectal cancer, hepatocellular carcinoma, breast cancer and pancreatic cancer) and five healthy patients (controls). FTIR(Fourier Transform Infrared) spectroscopy measurements were performed as a complementary approach to Raman analysis, on two of the four cancer subtypes. The AdaBoost Random Forest Classifier, Decision Trees, and Support Vector Machines (SVM) distinguished the baseline corrected Raman spectra of cancer EVs from those of healthy controls (18 spectra) with a classification accuracy of greater than 90% when reduced to a spectral frequency range of 1800 to 1940 inverse cm, and subjected to a 0.5 training/testing split. FTIR classification accuracy on 14 spectra showed an 80% classification accuracy. Our findings demonstrate that basic machine learning algorithms are powerful tools to distinguish the complex vibrational spectra of cancer patient EVs from those of healthy patients. These experimental methods hold promise as valid and efficient liquid biopsy for machine intelligence-assisted early cancer screening.
Skin Cancer Detection and Tracking using Data Synthesis and Deep Learning
Dec 04, 2016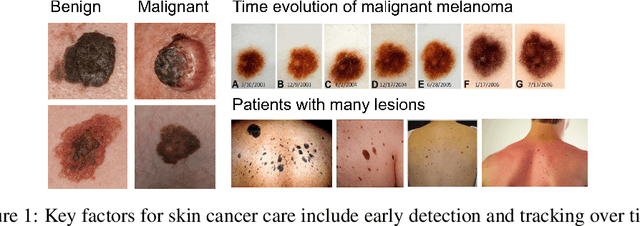
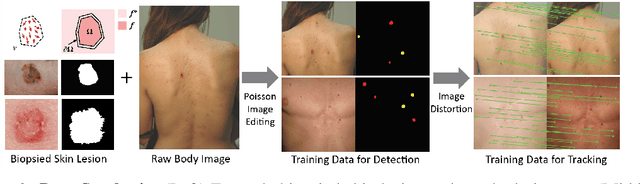
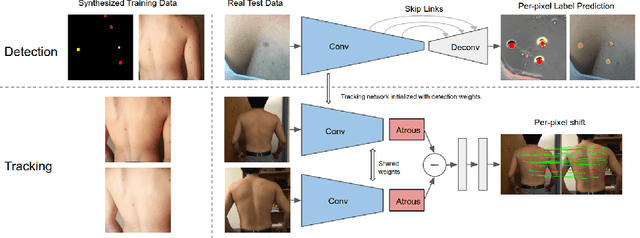
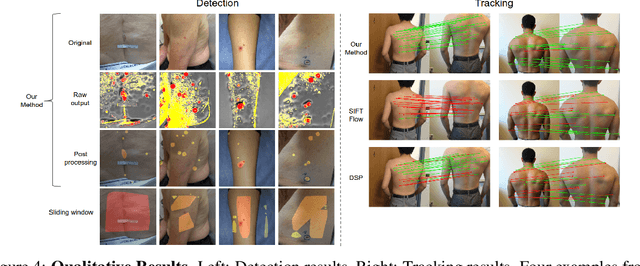
Dense object detection and temporal tracking are needed across applications domains ranging from people-tracking to analysis of satellite imagery over time. The detection and tracking of malignant skin cancers and benign moles poses a particularly challenging problem due to the general uniformity of large skin patches, the fact that skin lesions vary little in their appearance, and the relatively small amount of data available. Here we introduce a novel data synthesis technique that merges images of individual skin lesions with full-body images and heavily augments them to generate significant amounts of data. We build a convolutional neural network (CNN) based system, trained on this synthetic data, and demonstrate superior performance to traditional detection and tracking techniques. Additionally, we compare our system to humans trained with simple criteria. Our system is intended for potential clinical use to augment the capabilities of healthcare providers. While domain-specific, we believe the methods invoked in this work will be useful in applying CNNs across domains that suffer from limited data availability.
Are Quantitative Features of Lung Nodules Reproducible at Different CT Acquisition and Reconstruction Parameters?
Aug 14, 2019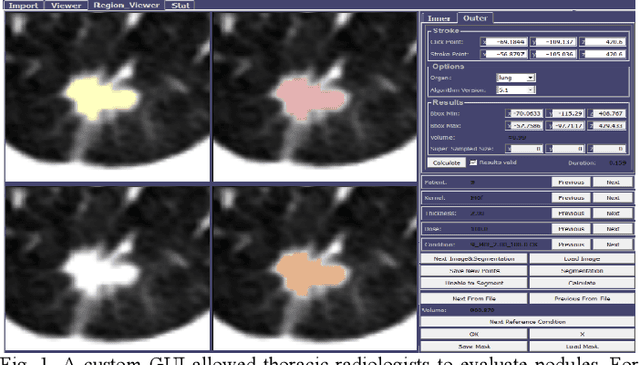
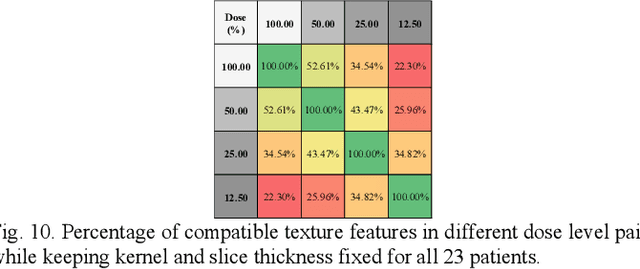
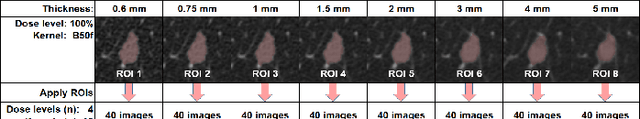
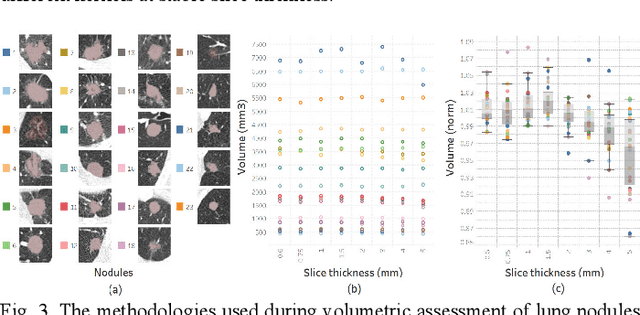
Consistency and duplicability in Computed Tomography (CT) output is essential to quantitative imaging for lung cancer detection and monitoring. This study of CT-detected lung nodules investigated the reproducibility of volume-, density-, and texture-based features (outcome variables) over routine ranges of radiation-dose, reconstruction kernel, and slice thickness. CT raw data of 23 nodules were reconstructed using 320 acquisition/reconstruction conditions (combinations of 4 doses, 10 kernels, and 8 thicknesses). Scans at 12.5%, 25%, and 50% of protocol dose were simulated; reduced-dose and full-dose data were reconstructed using conventional filtered back-projection and iterative-reconstruction kernels at a range of thicknesses (0.6-5.0 mm). Full-dose/B50f kernel reconstructions underwent expert segmentation for reference Region-Of-Interest (ROI) and nodule volume per thickness; each ROI was applied to 40 corresponding images (combinations of 4 doses and 10 kernels). Typical texture analysis metrics (including 5 histogram features, 13 Gray Level Co-occurrence Matrix, 5 Run Length Matrix, 2 Neighboring Gray-Level Dependence Matrix, and 2 Neighborhood Gray-Tone Difference Matrix) were computed per ROI. Reconstruction conditions resulting in no significant change in volume, density, or texture metrics were identified as "compatible pairs" for a given outcome variable. Our results indicate that as thickness increases, volumetric reproducibility decreases, while reproducibility of histogram- and texture-based features across different acquisition and reconstruction parameters improves. In order to achieve concomitant reproducibility of volumetric and radiomic results across studies, balanced standardization of the imaging acquisition parameters is required.
Double Encoder-Decoder Networks for Gastrointestinal Polyp Segmentation
Oct 05, 2021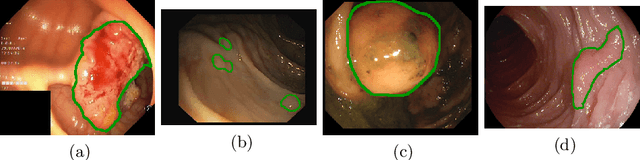
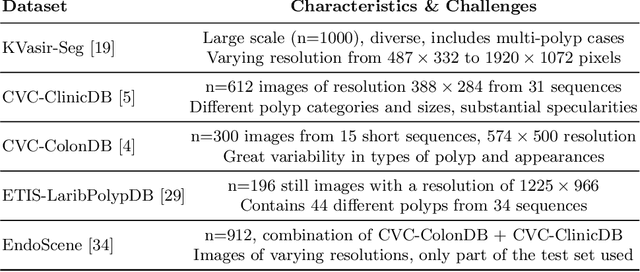

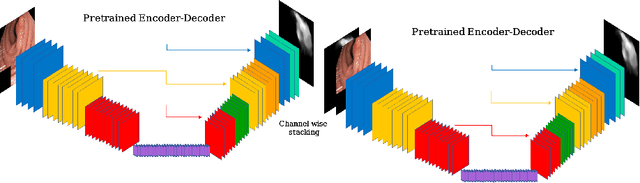
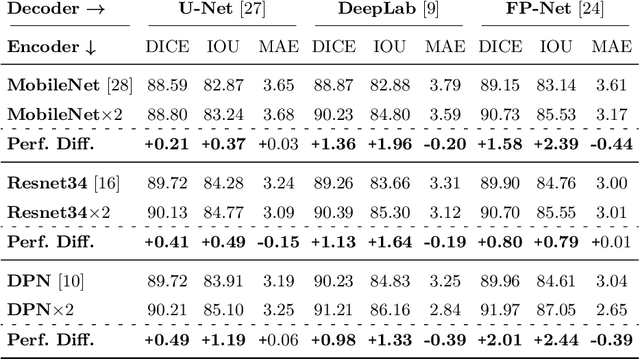
Polyps represent an early sign of the development of Colorectal Cancer. The standard procedure for their detection consists of colonoscopic examination of the gastrointestinal tract. However, the wide range of polyp shapes and visual appearances, as well as the reduced quality of this image modality, turn their automatic identification and segmentation with computational tools into a challenging computer vision task. In this work, we present a new strategy for the delineation of gastrointestinal polyps from endoscopic images based on a direct extension of common encoder-decoder networks for semantic segmentation. In our approach, two pretrained encoder-decoder networks are sequentially stacked: the second network takes as input the concatenation of the original frame and the initial prediction generated by the first network, which acts as an attention mechanism enabling the second network to focus on interesting areas within the image, thereby improving the quality of its predictions. Quantitative evaluation carried out on several polyp segmentation databases shows that double encoder-decoder networks clearly outperform their single encoder-decoder counterparts in all cases. In addition, our best double encoder-decoder combination attains excellent segmentation accuracy and reaches state-of-the-art performance results in all the considered datasets, with a remarkable boost of accuracy on images extracted from datasets not used for training.
Ensemble classifier approach in breast cancer detection and malignancy grading- A review
Apr 11, 2017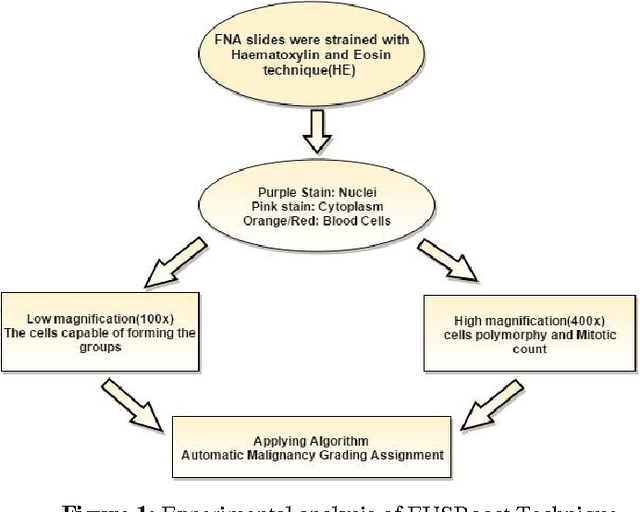



The diagnosed cases of Breast cancer is increasing annually and unfortunately getting converted into a high mortality rate. Cancer, at the early stages, is hard to detect because the malicious cells show similar properties (density) as shown by the non-malicious cells. The mortality ratio could have been minimized if the breast cancer could have been detected in its early stages. But the current systems have not been able to achieve a fully automatic system which is not just capable of detecting the breast cancer but also can detect the stage of it. Estimation of malignancy grading is important in diagnosing the degree of growth of malicious cells as well as in selecting a proper therapy for the patient. Therefore, a complete and efficient clinical decision support system is proposed which is capable of achieving breast cancer malignancy grading scheme very efficiently. The system is based on Image processing and machine learning domains. Classification Imbalance problem, a machine learning problem, occurs when instances of one class is much higher than the instances of the other class resulting in an inefficient classification of samples and hence a bad decision support system. Therefore EUSBoost, ensemble based classifier is proposed which is efficient and is able to outperform other classifiers as it takes the benefits of both-boosting algorithm with Random Undersampling techniques. Also comparison of EUSBoost with other techniques is shown in the paper.
* 10 pages,1 figure,5 tables
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge