"cancer detection": models, code, and papers
Improving Specificity in Mammography Using Cross-correlation between Wavelet and Fourier Transform
Jan 29, 2022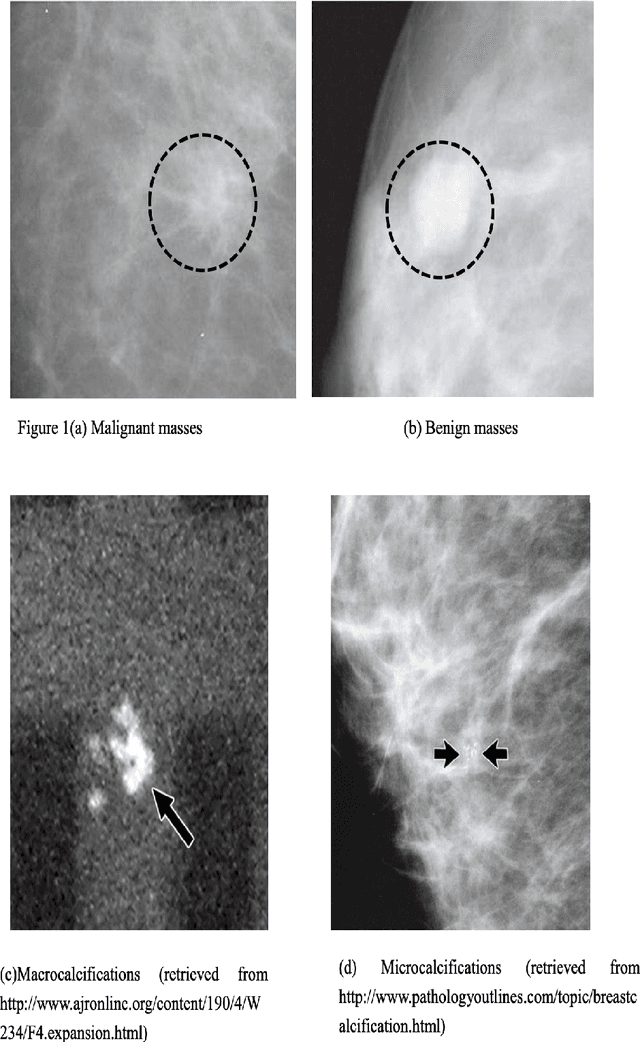
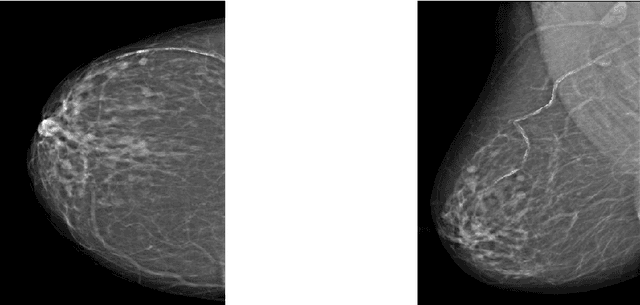
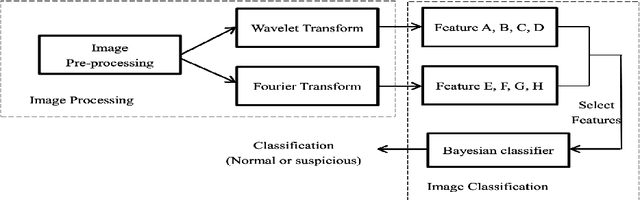
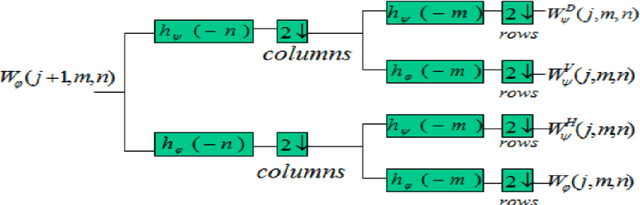
Breast cancer is in the most common malignant tumor in women. It accounted for 30% of new malignant tumor cases. Although the incidence of breast cancer remains high around the world, the mortality rate has been continuously reduced. This is mainly due to recent developments in molecular biology technology and improved level of comprehensive diagnosis and standard treatment. Early detection by mammography is an integral part of that. The most common breast abnormalities that may indicate breast cancer are masses and calcifications. Previous detection approaches usually obtain relatively high sensitivity but unsatisfactory specificity. We will investigate an approach that applies the discrete wavelet transform and Fourier transform to parse the images and extracts statistical features that characterize an image's content, such as the mean intensity and the skewness of the intensity. A naive Bayesian classifier uses these features to classify the images. We expect to achieve an optimal high specificity.
Integration of Convolutional Neural Networks for Pulmonary Nodule Malignancy Assessment in a Lung Cancer Classification Pipeline
Dec 18, 2019
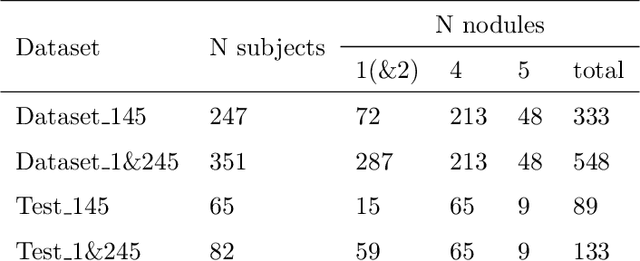
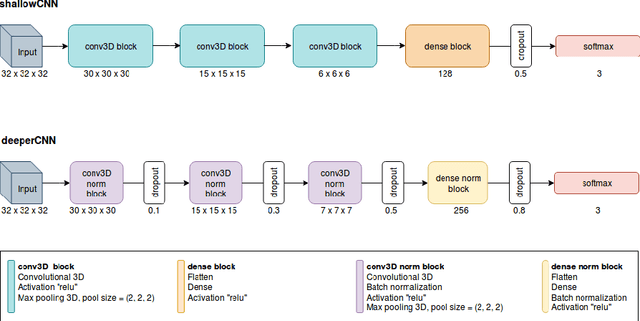
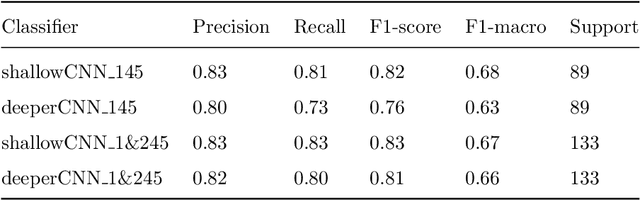
The early identification of malignant pulmonary nodules is critical for better lung cancer prognosis and less invasive chemo or radio therapies. Nodule malignancy assessment done by radiologists is extremely useful for planning a preventive intervention but is, unfortunately, a complex, time-consuming and error-prone task. This explains the lack of large datasets containing radiologists malignancy characterization of nodules. In this article, we propose to assess nodule malignancy through 3D convolutional neural networks and to integrate it in an automated end-to-end existing pipeline of lung cancer detection. For training and testing purposes we used independent subsets of the LIDC dataset. Adding the probabilities of nodules malignity in a baseline lung cancer pipeline improved its F1-weighted score by 14.7%, whereas integrating the malignancy model itself using transfer learning outperformed the baseline prediction by 11.8% of F1-weighted score. Despite the limited size of the lung cancer datasets, integrating predictive models of nodule malignancy improves prediction of lung cancer.
* 26 pages, 5 figures
Research on the Detection Method of Breast Cancer Deep Convolutional Neural Network Based on Computer Aid
Apr 23, 2021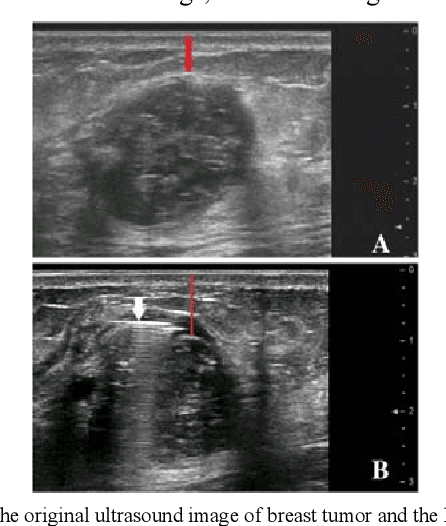
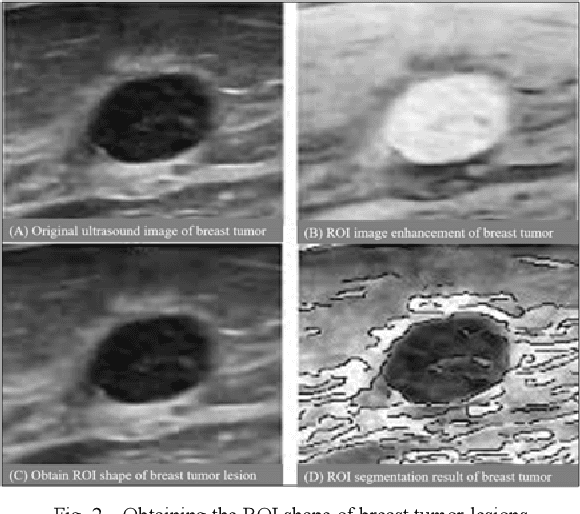
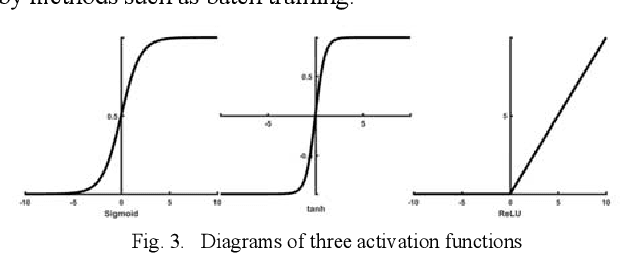
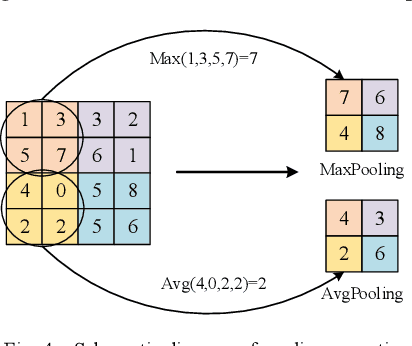
Traditional breast cancer image classification methods require manual extraction of features from medical images, which not only require professional medical knowledge, but also have problems such as time-consuming and labor-intensive and difficulty in extracting high-quality features. Therefore, the paper proposes a computer-based feature fusion Convolutional neural network breast cancer image classification and detection method. The paper pre-trains two convolutional neural networks with different structures, and then uses the convolutional neural network to automatically extract the characteristics of features, fuse the features extracted from the two structures, and finally use the classifier classifies the fused features. The experimental results show that the accuracy of this method in the classification of breast cancer image data sets is 89%, and the classification accuracy of breast cancer images is significantly improved compared with traditional methods.
Deep 3D Convolutional Neural Network for Automated Lung Cancer Diagnosis
May 04, 2019
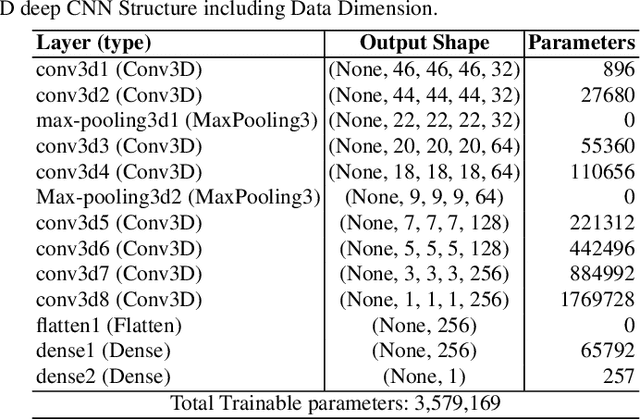
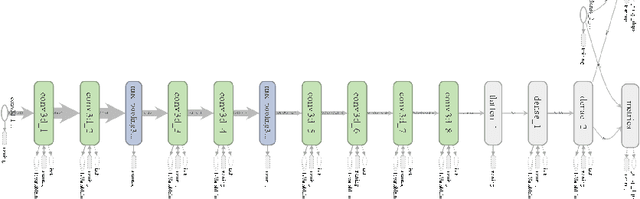
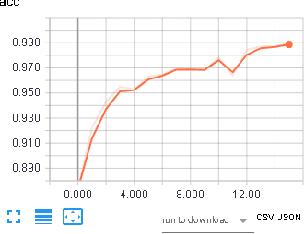
Computer Aided Diagnosis has emerged as an indispensible technique for validating the opinion of radiologists in CT interpretation. This paper presents a deep 3D Convolutional Neural Network (CNN) architecture for automated CT scan-based lung cancer detection system. It utilizes three dimensional spatial information to learn highly discriminative 3 dimensional features instead of 2D features like texture or geometric shape whick need to be generated manually. The proposed deep learning method automatically extracts the 3D features on the basis of spatio-temporal statistics.The developed model is end-to-end and is able to predict malignancy of each voxel for given input scan. Simulation results demonstrate the effectiveness of proposed 3D CNN network for classification of lung nodule in-spite of limited computational capabilities.
Skin Lesion Segmentation and Classification with Deep Learning System
Feb 16, 2019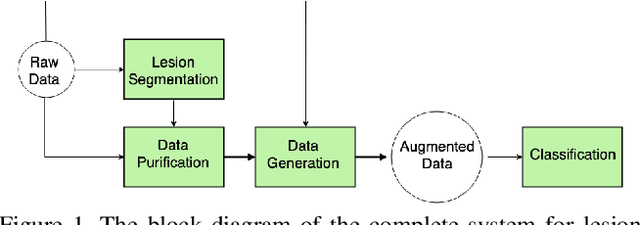
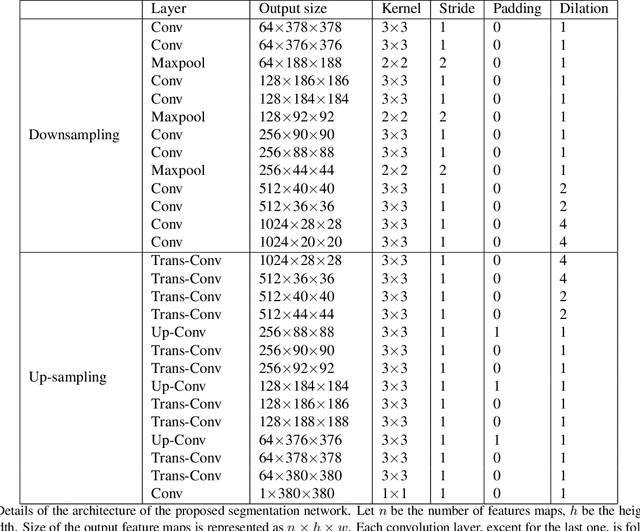
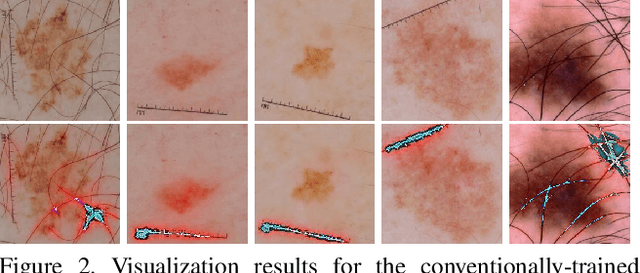
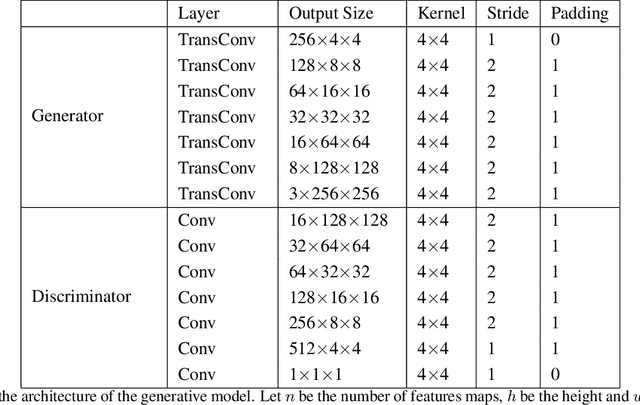
Melanoma is one of the ten most common cancers in the US. Early detection is crucial for survival, but often the cancer is diagnosed in the fatal stage. Deep learning has the potential to improve cancer detection rates, but its applicability to melanoma detection is compromised by the limitations of the available skin lesion databases, which are small, heavily imbalanced, and contain images with occlusions. We propose a complete deep learning system for lesion segmentation and classification that utilizes networks specialized in data purification and augmentation. It contains the processing unit for removing image occlusions and the data generation unit for populating scarce lesion classes, or equivalently creating virtual patients with pre-defined types of lesions. We empirically verify our approach and show superior performance over common baselines.
Interpretability methods of machine learning algorithms with applications in breast cancer diagnosis
Feb 04, 2022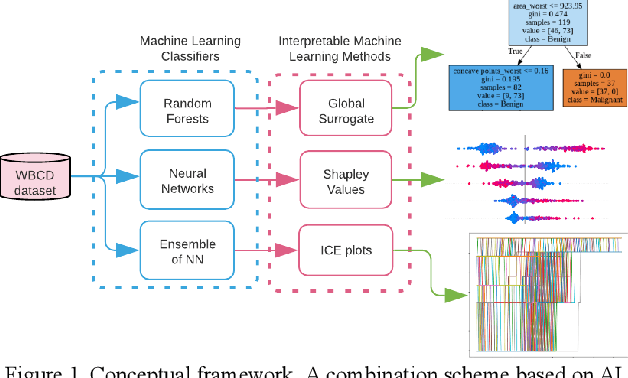
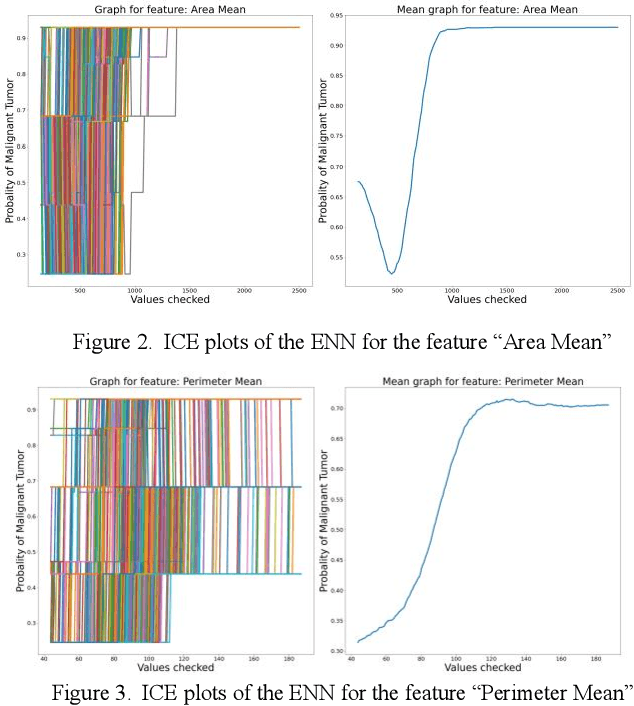
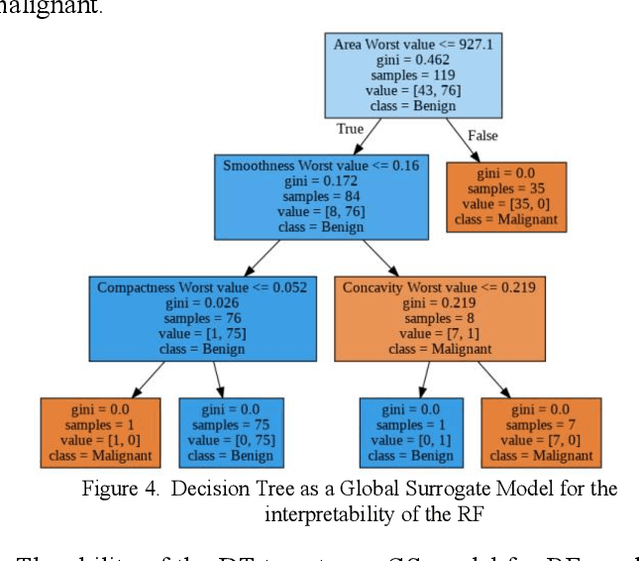
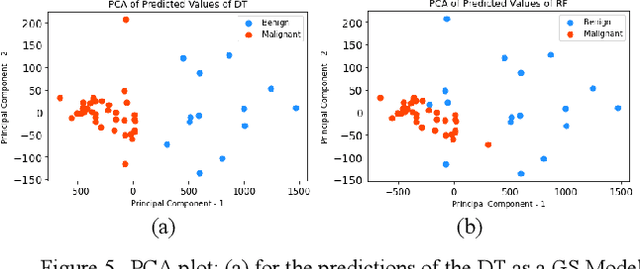
Early detection of breast cancer is a powerful tool towards decreasing its socioeconomic burden. Although, artificial intelligence (AI) methods have shown remarkable results towards this goal, their "black box" nature hinders their wide adoption in clinical practice. To address the need for AI guided breast cancer diagnosis, interpretability methods can be utilized. In this study, we used AI methods, i.e., Random Forests (RF), Neural Networks (NN) and Ensembles of Neural Networks (ENN), towards this goal and explained and optimized their performance through interpretability techniques, such as the Global Surrogate (GS) method, the Individual Conditional Expectation (ICE) plots and the Shapley values (SV). The Wisconsin Diagnostic Breast Cancer (WDBC) dataset of the open UCI repository was used for the training and evaluation of the AI algorithms. The best performance for breast cancer diagnosis was achieved by the proposed ENN (96.6% accuracy and 0.96 area under the ROC curve), and its predictions were explained by ICE plots, proving that its decisions were compliant with current medical knowledge and can be further utilized to gain new insights in the pathophysiological mechanisms of breast cancer. Feature selection based on features' importance according to the GS model improved the performance of the RF (leading the accuracy from 96.49% to 97.18% and the area under the ROC curve from 0.96 to 0.97) and feature selection based on features' importance according to SV improved the performance of the NN (leading the accuracy from 94.6% to 95.53% and the area under the ROC curve from 0.94 to 0.95). Compared to other approaches on the same dataset, our proposed models demonstrated state of the art performance while being interpretable.
Deep learning for brain metastasis detection and segmentation in longitudinal MRI data
Dec 28, 2021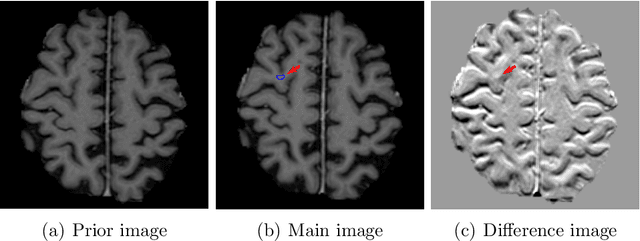
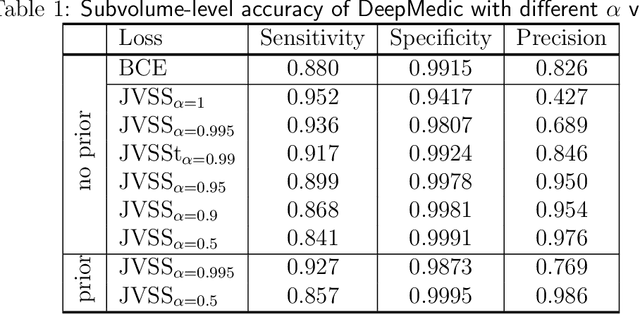
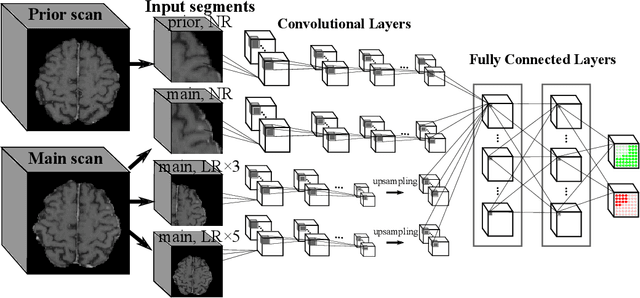
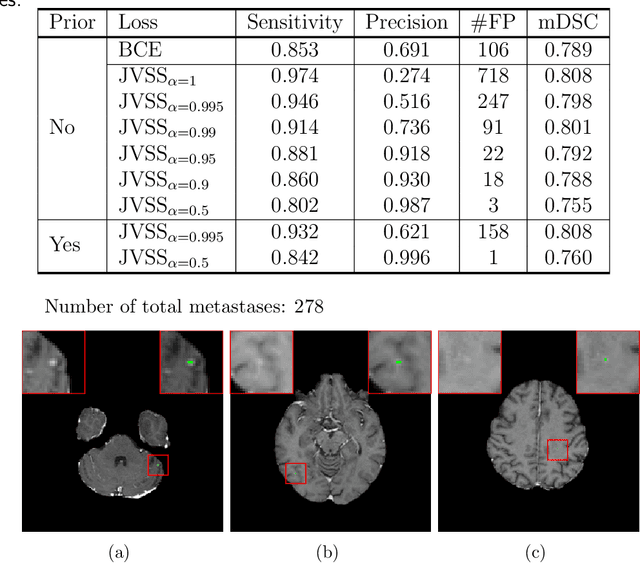
Brain metastases occur frequently in patients with metastatic cancer. Early and accurate detection of brain metastases is very essential for treatment planning and prognosis in radiation therapy. To improve brain metastasis detection performance with deep learning, a custom detection loss called volume-level sensitivity-specificity (VSS) is proposed, which rates individual metastasis detection sensitivity and specificity in (sub-)volume levels. As sensitivity and precision are always a trade-off in a metastasis level, either a high sensitivity or a high precision can be achieved by adjusting the weights in the VSS loss without decline in dice score coefficient for segmented metastases. To reduce metastasis-like structures being detected as false positive metastases, a temporal prior volume is proposed as an additional input of the neural network. Our proposed VSS loss improves the sensitivity of brain metastasis detection, increasing the sensitivity from 86.7% to 95.5%. Alternatively, it improves the precision from 68.8% to 97.8%. With the additional temporal prior volume, about 45% of the false positive metastases are reduced in the high sensitivity model and the precision reaches 99.6% for the high specificity model. The mean dice coefficient for all metastases is about 0.81. With the ensemble of the high sensitivity and high specificity models, on average only 1.5 false positive metastases per patient needs further check, while the majority of true positive metastases are confirmed. The ensemble learning is able to distinguish high confidence true positive metastases from metastases candidates that require special expert review or further follow-up, being particularly well-fit to the requirements of expert support in real clinical practice.
Coverage Testing of Deep Learning Models using Dataset Characterization
Nov 17, 2019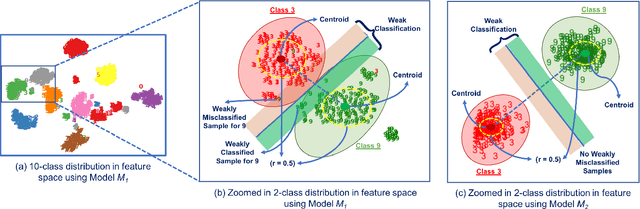
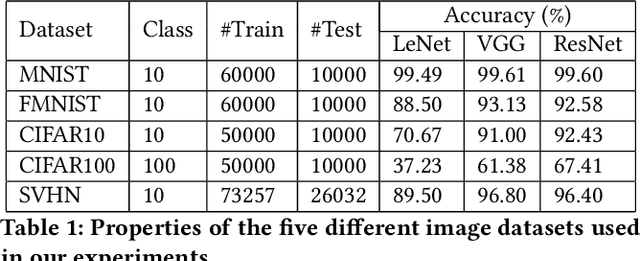
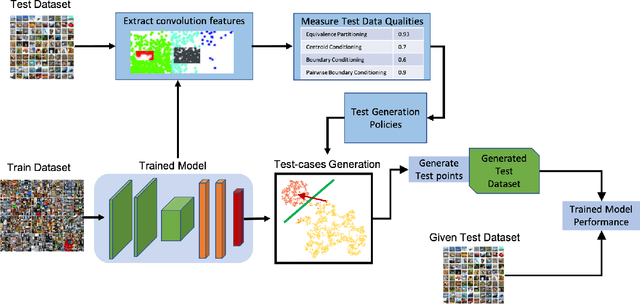
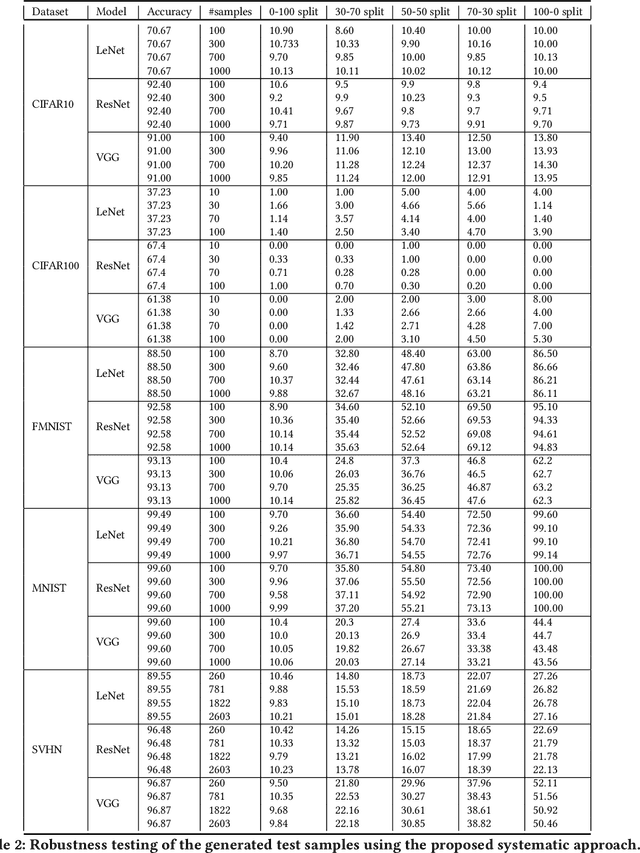
Deep Neural Networks (DNNs), with its promising performance, are being increasingly used in safety critical applications such as autonomous driving, cancer detection, and secure authentication. With growing importance in deep learning, there is a requirement for a more standardized framework to evaluate and test deep learning models. The primary challenge involved in automated generation of extensive test cases are: (i) neural networks are difficult to interpret and debug and (ii) availability of human annotators to generate specialized test points. In this research, we explain the necessity to measure the quality of a dataset and propose a test case generation system guided by the dataset properties. From a testing perspective, four different dataset quality dimensions are proposed: (i) equivalence partitioning, (ii) centroid positioning, (iii) boundary conditioning, and (iv) pair-wise boundary conditioning. The proposed system is evaluated on well known image classification datasets such as MNIST, Fashion-MNIST, CIFAR10, CIFAR100, and SVHN against popular deep learning models such as LeNet, ResNet-20, VGG-19. Further, we conduct various experiments to demonstrate the effectiveness of systematic test case generation system for evaluating deep learning models.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge