"cancer detection": models, code, and papers
Improved Pancreatic Tumor Detection by Utilizing Clinically-Relevant Secondary Features
Aug 06, 2022
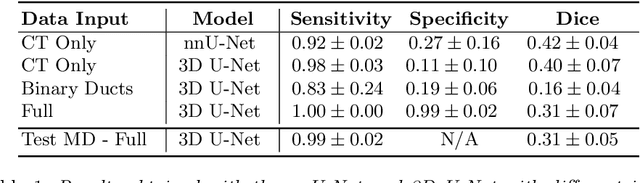
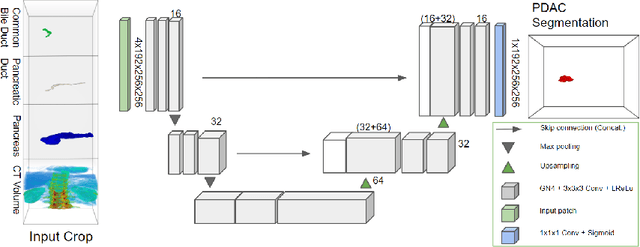

Pancreatic cancer is one of the global leading causes of cancer-related deaths. Despite the success of Deep Learning in computer-aided diagnosis and detection (CAD) methods, little attention has been paid to the detection of Pancreatic Cancer. We propose a method for detecting pancreatic tumor that utilizes clinically-relevant features in the surrounding anatomical structures, thereby better aiming to exploit the radiologist's knowledge compared to other, conventional deep learning approaches. To this end, we collect a new dataset consisting of 99 cases with pancreatic ductal adenocarcinoma (PDAC) and 97 control cases without any pancreatic tumor. Due to the growth pattern of pancreatic cancer, the tumor may not be always visible as a hypodense lesion, therefore experts refer to the visibility of secondary external features that may indicate the presence of the tumor. We propose a method based on a U-Net-like Deep CNN that exploits the following external secondary features: the pancreatic duct, common bile duct and the pancreas, along with a processed CT scan. Using these features, the model segments the pancreatic tumor if it is present. This segmentation for classification and localization approach achieves a performance of 99% sensitivity (one case missed) and 99% specificity, which realizes a 5% increase in sensitivity over the previous state-of-the-art method. The model additionally provides location information with reasonable accuracy and a shorter inference time compared to previous PDAC detection methods. These results offer a significant performance improvement and highlight the importance of incorporating the knowledge of the clinical expert when developing novel CAD methods.
EGFR Mutation Prediction of Lung Biopsy Images using Deep Learning
Aug 26, 2022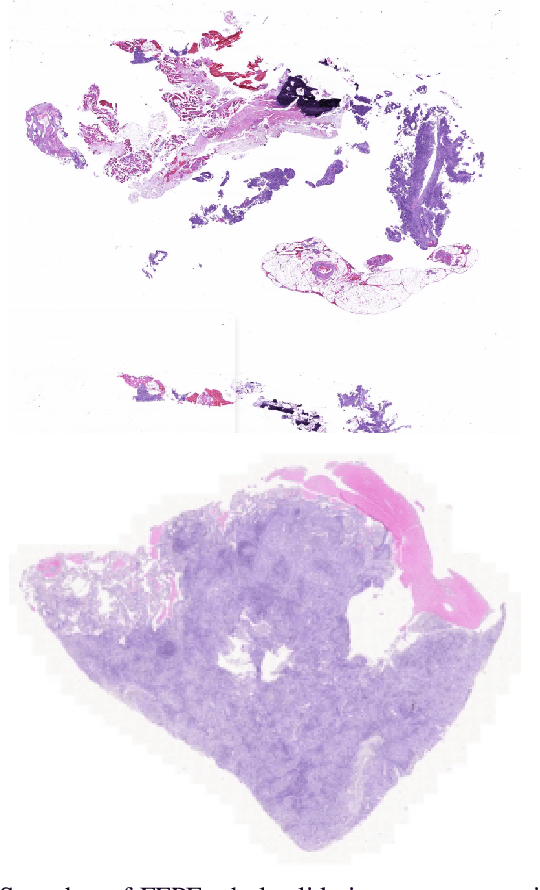
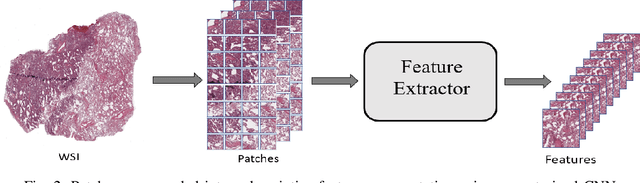
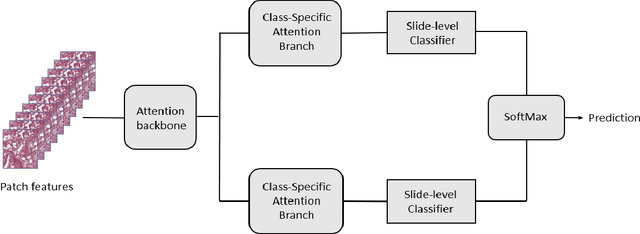
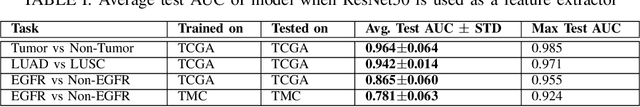
The standard diagnostic procedures for targeted therapies in lung cancer treatment involve histological subtyping and subsequent detection of key driver mutations, such as EGFR. Even though molecular profiling can uncover the driver mutation, the process is often expensive and time-consuming. Deep learning-oriented image analysis offers a more economical alternative for discovering driver mutations directly from whole slide images (WSIs). In this work, we used customized deep learning pipelines with weak supervision to identify the morphological correlates of EGFR mutation from hematoxylin and eosin-stained WSIs, in addition to detecting tumor and histologically subtyping it. We demonstrate the effectiveness of our pipeline by conducting rigorous experiments and ablation studies on two lung cancer datasets - TCGA and a private dataset from India. With our pipeline, we achieved an average area under the curve (AUC) of 0.964 for tumor detection, and 0.942 for histological subtyping between adenocarcinoma and squamous cell carcinoma on the TCGA dataset. For EGFR detection, we achieved an average AUC of 0.864 on the TCGA dataset and 0.783 on the dataset from India. Our key learning points include the following. Firstly, there is no particular advantage of using a feature extractor layers trained on histology, if one is going to fine-tune the feature extractor on the target dataset. Secondly, selecting patches with high cellularity, presumably capturing tumor regions, is not always helpful, as the sign of a disease class may be present in the tumor-adjacent stroma.
Embedded Deep Regularized Block HSIC Thermomics for Early Diagnosis of Breast Cancer
Jun 03, 2021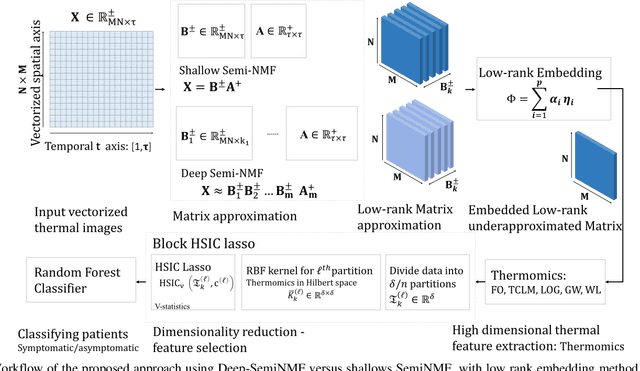
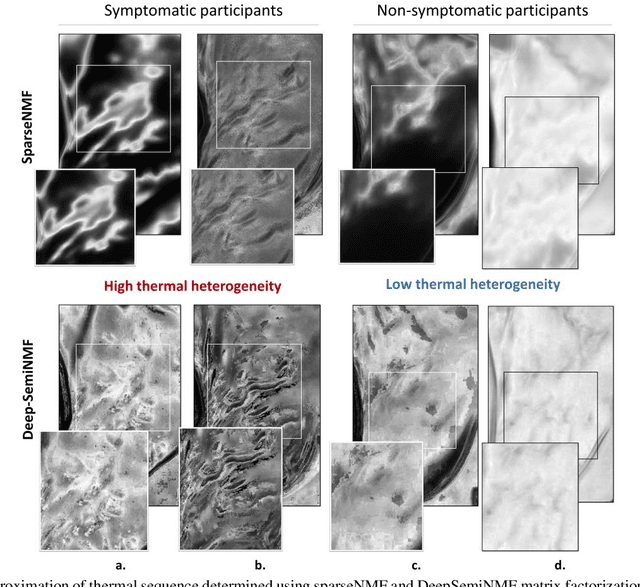
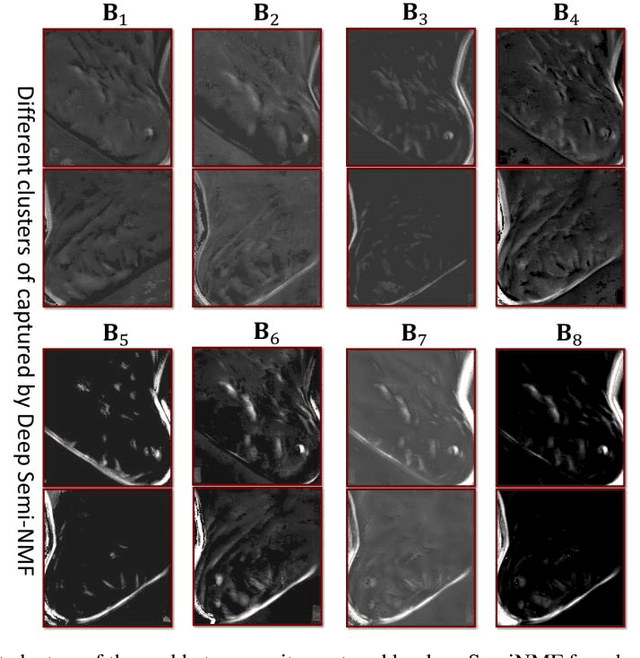


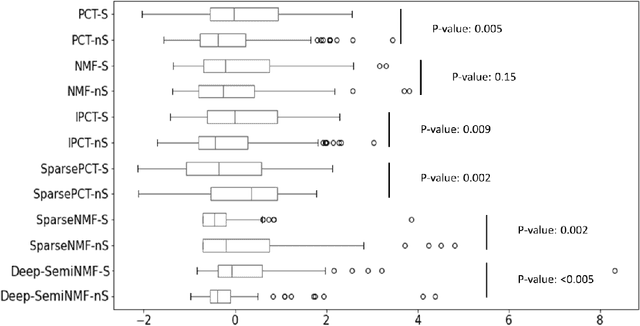
Thermography has been used extensively as a complementary diagnostic tool in breast cancer detection. Among thermographic methods matrix factorization (MF) techniques show an unequivocal capability to detect thermal patterns corresponding to vasodilation in cancer cases. One of the biggest challenges in such techniques is selecting the best representation of the thermal basis. In this study, an embedding method is proposed to address this problem and Deep-semi-nonnegative matrix factorization (Deep-SemiNMF) for thermography is introduced, then tested for 208 breast cancer screening cases. First, we apply Deep-SemiNMF to infrared images to extract low-rank thermal representations for each case. Then, we embed low-rank bases to obtain one basis for each patient. After that, we extract 300 thermal imaging features, called thermomics, to decode imaging information for the automatic diagnostic model. We reduced the dimensionality of thermomics by spanning them onto Hilbert space using RBF kernel and select the three most efficient features using the block Hilbert Schmidt Independence Criterion Lasso (block HSIC Lasso). The preserved thermal heterogeneity successfully classified asymptomatic versus symptomatic patients applying a random forest model (cross-validated accuracy of 71.36% (69.42%-73.3%)).
* Authors version. arXiv admin note: text overlap with arXiv:2010.06784
Detecting and analysing spontaneous oral cancer speech in the wild
Jul 28, 2020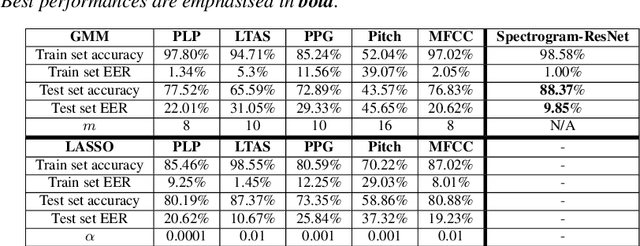
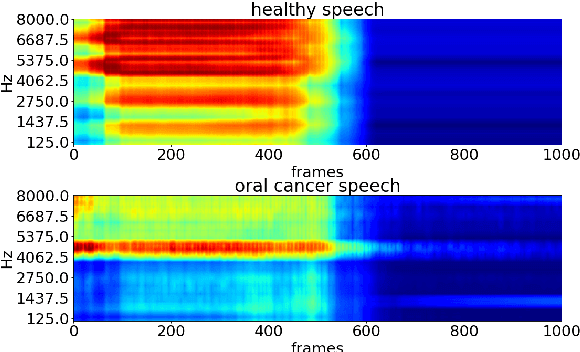
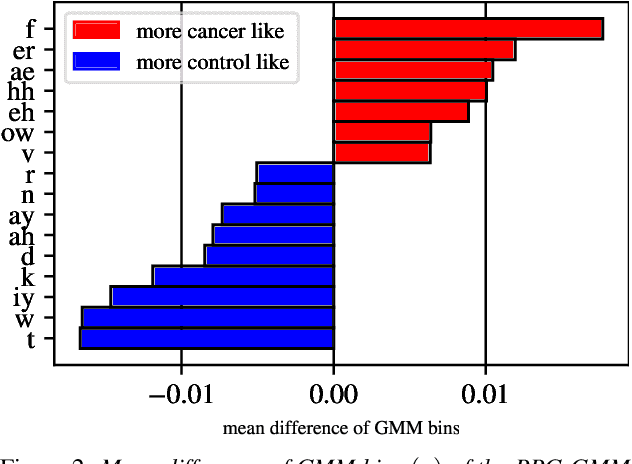
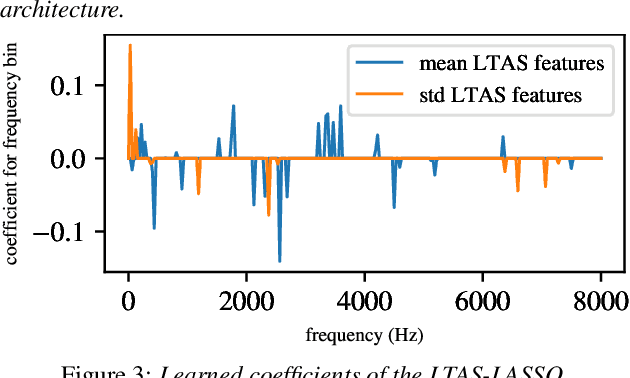
Oral cancer speech is a disease which impacts more than half a million people worldwide every year. Analysis of oral cancer speech has so far focused on read speech. In this paper, we 1) present and 2) analyse a three-hour long spontaneous oral cancer speech dataset collected from YouTube. 3) We set baselines for an oral cancer speech detection task on this dataset. The analysis of these explainable machine learning baselines shows that sibilants and stop consonants are the most important indicators for spontaneous oral cancer speech detection.
Proposing method to Increase the detection accuracy of stomach cancer based on colour and lint features of tongue using CNN and SVM
Nov 18, 2020
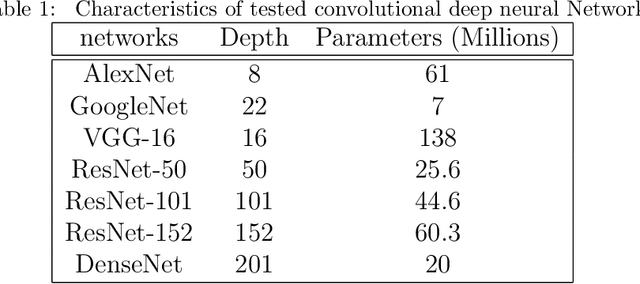
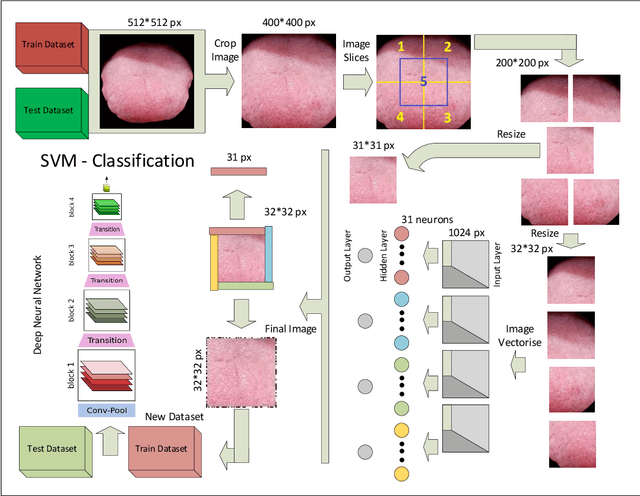
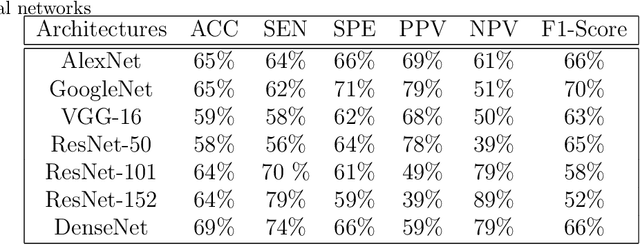
Today, gastric cancer is one of the diseases which affected many people's life. Early detection and accuracy are the main and crucial challenges in finding this kind of cancer. In this paper, a method to increase the accuracy of the diagnosis of detecting cancer using lint and colour features of tongue based on deep convolutional neural networks and support vector machine is proposed. In the proposed method, the region of tongue is first separated from the face image by {deep RCNN} \color{black} Recursive Convolutional Neural Network (R-CNN) \color{black}. After the necessary preprocessing, the images to the convolutional neural network are provided and the training and test operations are triggered. The results show that the proposed method is correctly able to identify the area of the tongue as well as the patient's person from the non-patient. Based on experiments, the DenseNet network has the highest accuracy compared to other deep architectures. The experimental results show that the accuracy of this network for gastric cancer detection reaches 91% which shows the superiority of method in comparison to the state-of-the-art methods.
$\text{O}^2$PF: Oversampling via Optimum-Path Forest for Breast Cancer Detection
Jan 14, 2021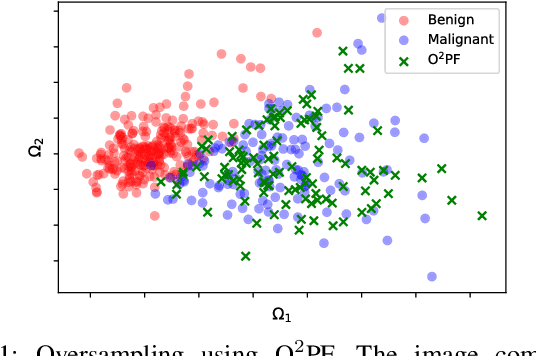
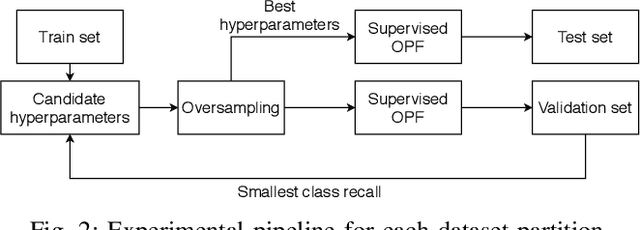
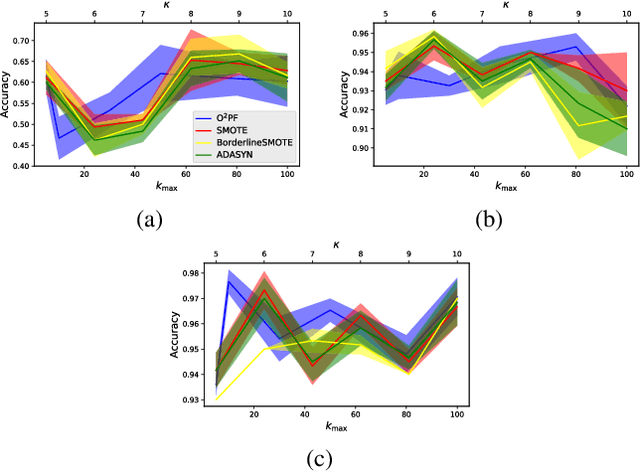
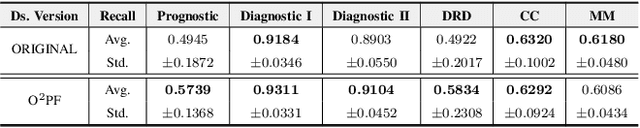
Breast cancer is among the most deadly diseases, distressing mostly women worldwide. Although traditional methods for detection have presented themselves as valid for the task, they still commonly present low accuracies and demand considerable time and effort from professionals. Therefore, a computer-aided diagnosis (CAD) system capable of providing early detection becomes hugely desirable. In the last decade, machine learning-based techniques have been of paramount importance in this context, since they are capable of extracting essential information from data and reasoning about it. However, such approaches still suffer from imbalanced data, specifically on medical issues, where the number of healthy people samples is, in general, considerably higher than the number of patients. Therefore this paper proposes the $\text{O}^2$PF, a data oversampling method based on the unsupervised Optimum-Path Forest Algorithm. Experiments conducted over the full oversampling scenario state the robustness of the model, which is compared against three well-established oversampling methods considering three breast cancer and three general-purpose tasks for medical issues datasets.
Visual Probing and Correction of Object Recognition Models with Interactive user feedback
Dec 29, 2020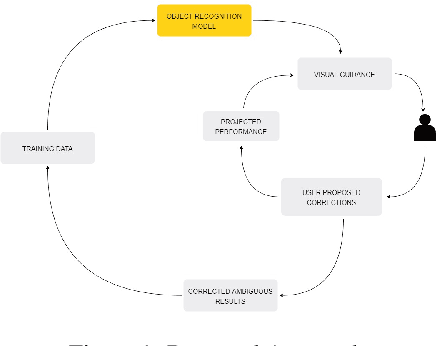
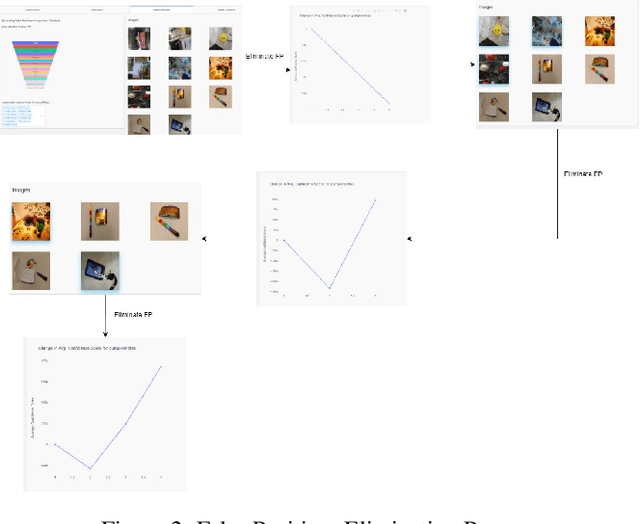
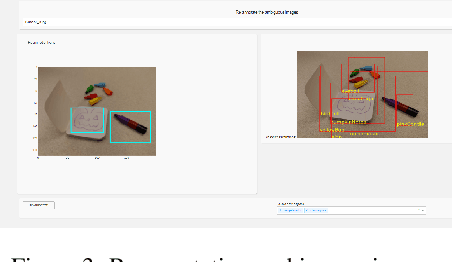
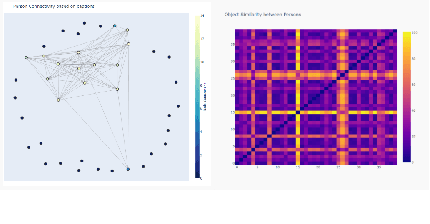
With the advent of state-of-the-art machine learning and deep learning technologies, several industries are moving towards the field. Applications of such technologies are highly diverse ranging from natural language processing to computer vision. Object recognition is one such area in the computer vision domain. Although proven to perform with high accuracy, there are still areas where such models can be improved. This is in-fact highly important in real-world use cases like autonomous driving or cancer detection, that are highly sensitive and expect such technologies to have almost no uncertainties. In this paper, we attempt to visualise the uncertainties in object recognition models and propose a correction process via user feedback. We further demonstrate our approach on the data provided by the VAST 2020 Mini-Challenge 2.
BCNet: A Deep Convolutional Neural Network for Breast Cancer Grading
Jul 11, 2021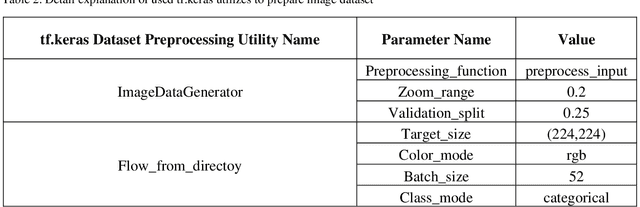
Breast cancer has become one of the most prevalent cancers by which people all over the world are affected and is posed serious threats to human beings, in a particular woman. In order to provide effective treatment or prevention of this cancer, disease diagnosis in the early stages would be of high importance. There have been various methods to detect this disorder in which using images have to play a dominant role. Deep learning has been recently adopted widely in different areas of science, especially medicine. In breast cancer detection problems, some diverse deep learning techniques have been developed on different datasets and resulted in good accuracy. In this article, we aimed to present a deep neural network model to classify histopathological images from the Databiox image dataset as the first application on this image database. Our proposed model named BCNet has taken advantage of the transfer learning approach in which VGG16 is selected from available pertained models as a feature extractor. Furthermore, to address the problem of insufficient data, we employed the data augmentation technique to expand the input dataset. All implementations in this research, ranging from pre-processing actions to depicting the diagram of the model architecture, have been carried out using tf.keras API. As a consequence of the proposed model execution, the significant validation accuracy of 88% and evaluation accuracy of 72% obtained.
Comparison of different CNNs for breast tumor classification from ultrasound images
Dec 28, 2020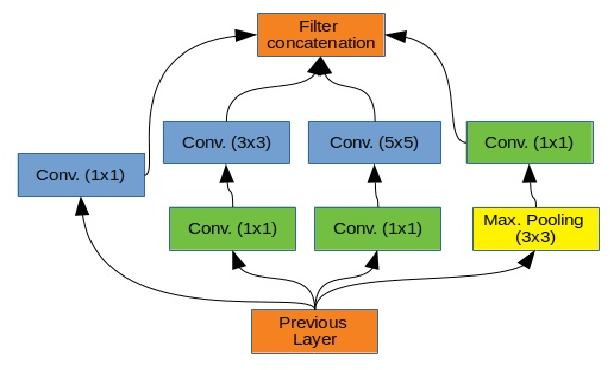
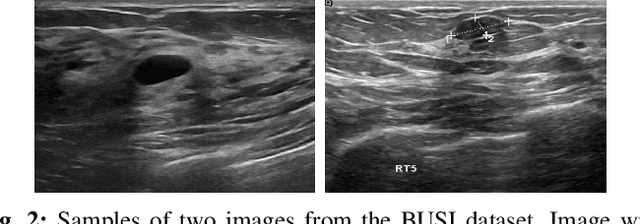


Breast cancer is one of the deadliest cancer worldwide. Timely detection could reduce mortality rates. In the clinical routine, classifying benign and malignant tumors from ultrasound (US) imaging is a crucial but challenging task. An automated method, which can deal with the variability of data is therefore needed. In this paper, we compared different Convolutional Neural Networks (CNNs) and transfer learning methods for the task of automated breast tumor classification. The architectures investigated in this study were VGG-16 and Inception V3. Two different training strategies were investigated: the first one was using pretrained models as feature extractors and the second one was to fine-tune the pre-trained models. A total of 947 images were used, 587 corresponded to US images of benign tumors and 360 with malignant tumors. 678 images were used for the training and validation process, while 269 images were used for testing the models. Accuracy and Area Under the receiver operating characteristic Curve (AUC) were used as performance metrics. The best performance was obtained by fine tuning VGG-16, with an accuracy of 0.919 and an AUC of 0.934. The obtained results open the opportunity to further investigation with a view of improving cancer detection.
Overhead MNIST: A Benchmark Satellite Dataset
Feb 08, 2021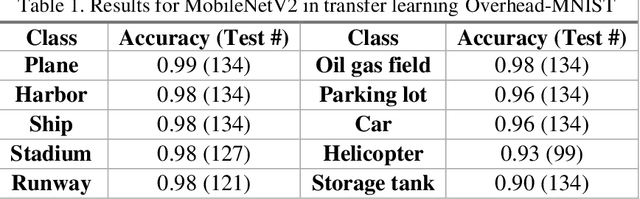

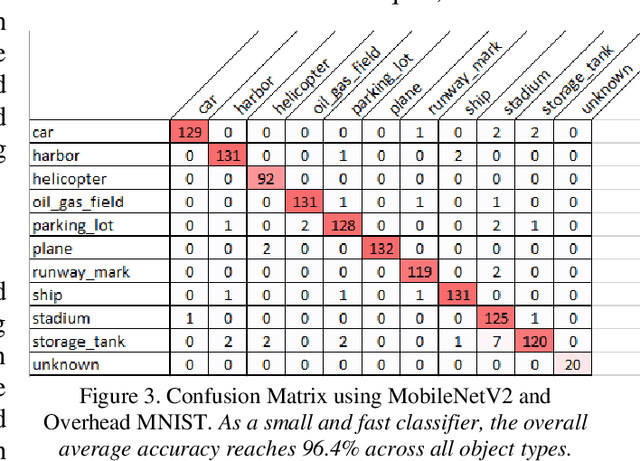
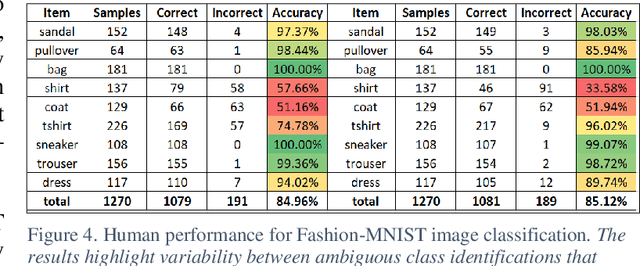
The research presents an overhead view of 10 important objects and follows the general formatting requirements of the most popular machine learning task: digit recognition with MNIST. This dataset offers a public benchmark extracted from over a million human-labelled and curated examples. The work outlines the key multi-class object identification task while matching with prior work in handwriting, cancer detection, and retail datasets. A prototype deep learning approach with transfer learning and convolutional neural networks (MobileNetV2) correctly identifies the ten overhead classes with an average accuracy of 96.7%. This model exceeds the peak human performance of 93.9%. For upgrading satellite imagery and object recognition, this new dataset benefits diverse endeavors such as disaster relief, land use management, and other traditional remote sensing tasks. The work extends satellite benchmarks with new capabilities to identify efficient and compact algorithms that might work on-board small satellites, a practical task for future multi-sensor constellations. The dataset is available on Kaggle and Github.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge