"cancer detection": models, code, and papers
Feasibility of Colon Cancer Detection in Confocal Laser Microscopy Images Using Convolution Neural Networks
Dec 05, 2018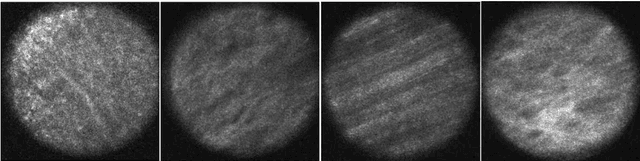
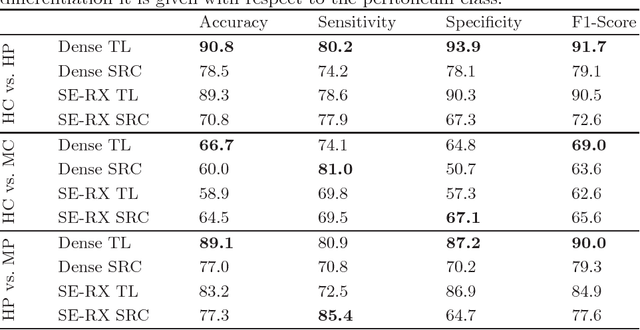
Histological evaluation of tissue samples is a typical approach to identify colorectal cancer metastases in the peritoneum. For immediate assessment, reliable and real-time in-vivo imaging would be required. For example, intraoperative confocal laser microscopy has been shown to be suitable for distinguishing organs and also malignant and benign tissue. So far, the analysis is done by human experts. We investigate the feasibility of automatic colon cancer classification from confocal laser microscopy images using deep learning models. We overcome very small dataset sizes through transfer learning with state-of-the-art architectures. We achieve an accuracy of 89.1% for cancer detection in the peritoneum which indicates viability as an intraoperative decision support system.
ESFPNet: efficient deep learning architecture for real-time lesion segmentation in autofluorescence bronchoscopic video
Jul 15, 2022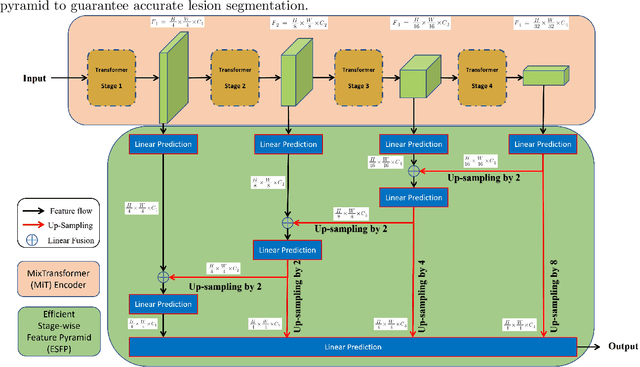
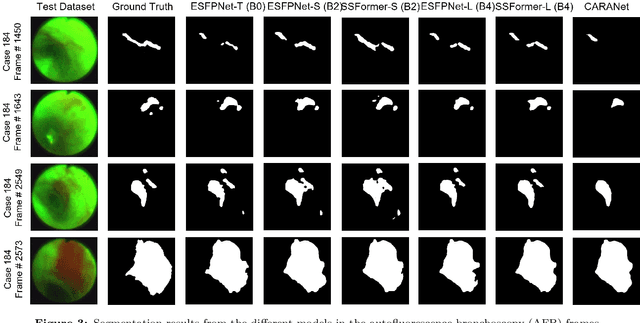

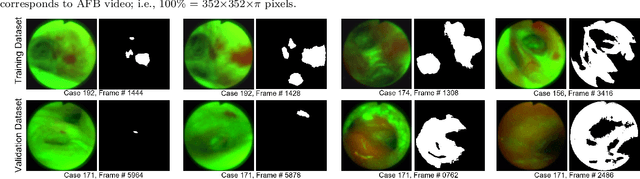

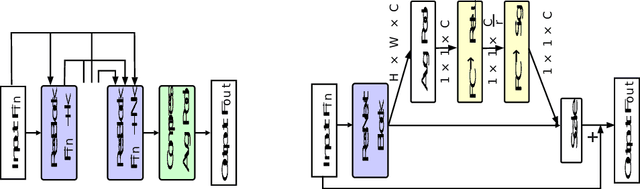
Lung cancer tends to be detected at an advanced stage, resulting in a high patient mortality rate. Thus, recent research has focused on early disease detection. Lung cancer generally first appears as lesions developing within the bronchial epithelium of the airway walls. Bronchoscopy is the procedure of choice for effective noninvasive bronchial lesion detection. In particular, autofluorescence bronchoscopy (AFB) discriminates the autofluorescence properties of normal and diseased tissue, whereby lesions appear reddish brown in AFB video frames, while normal tissue appears green. Because recent studies show AFB's ability for high lesion sensitivity, it has become a potentially pivotal method during the standard bronchoscopic airway exam for early-stage lung cancer detection. Unfortunately, manual inspection of AFB video is extremely tedious and error-prone, while limited effort has been expended toward potentially more robust automatic AFB lesion detection and segmentation. We propose a real-time deep learning architecture ESFPNet for robust detection and segmentation of bronchial lesions from an AFB video stream. The architecture features an encoder structure that exploits pretrained Mix Transformer (MiT) encoders and a stage-wise feature pyramid (ESFP) decoder structure. Results from AFB videos derived from lung cancer patient airway exams indicate that our approach gives mean Dice index and IOU values of 0.782 and 0.658, respectively, while having a processing throughput of 27 frames/sec. These values are superior to results achieved by other competing architectures that use Mix transformers or CNN-based encoders. Moreover, the superior performance on the ETIS-LaribPolypDB dataset demonstrates its potential applicability to other domains.
EBHI-Seg: A Novel Enteroscope Biopsy Histopathological Haematoxylin and Eosin Image Dataset for Image Segmentation Tasks
Dec 02, 2022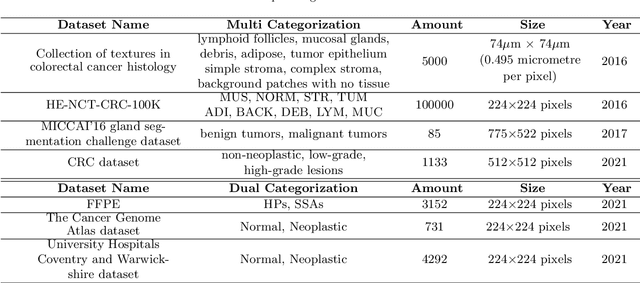
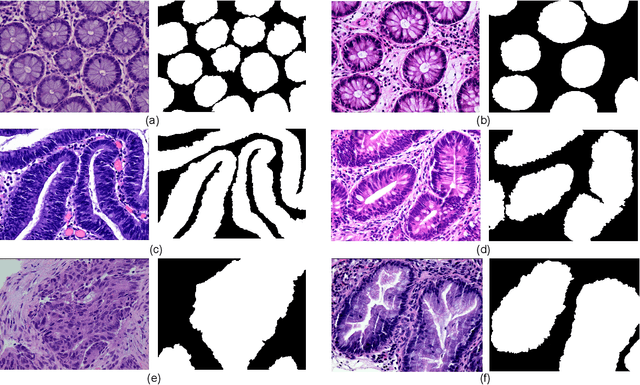
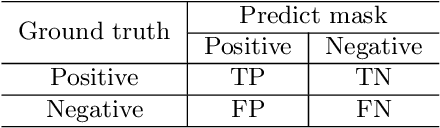
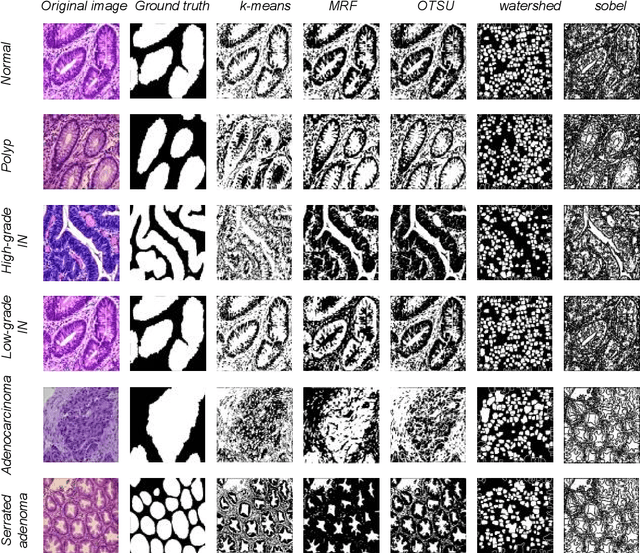
Background and Purpose: Colorectal cancer is a common fatal malignancy, the fourth most common cancer in men, and the third most common cancer in women worldwide. Timely detection of cancer in its early stages is essential for treating the disease. Currently, there is a lack of datasets for histopathological image segmentation of rectal cancer, which often hampers the assessment accuracy when computer technology is used to aid in diagnosis. Methods: This present study provided a new publicly available Enteroscope Biopsy Histopathological Hematoxylin and Eosin Image Dataset for Image Segmentation Tasks (EBHI-Seg). To demonstrate the validity and extensiveness of EBHI-Seg, the experimental results for EBHI-Seg are evaluated using classical machine learning methods and deep learning methods. Results: The experimental results showed that deep learning methods had a better image segmentation performance when utilizing EBHI-Seg. The maximum accuracy of the Dice evaluation metric for the classical machine learning method is 0.948, while the Dice evaluation metric for the deep learning method is 0.965. Conclusion: This publicly available dataset contained 5,170 images of six types of tumor differentiation stages and the corresponding ground truth images. The dataset can provide researchers with new segmentation algorithms for medical diagnosis of colorectal cancer, which can be used in the clinical setting to help doctors and patients.
Detection and Localization of Melanoma Skin Cancer in Histopathological Whole Slide Images
Feb 17, 2023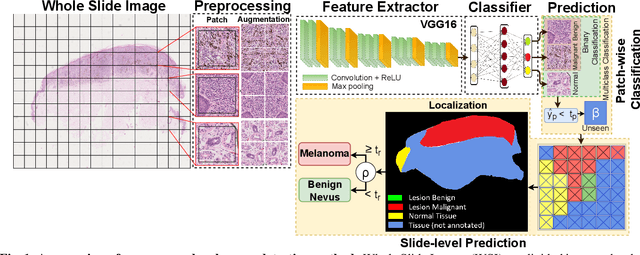
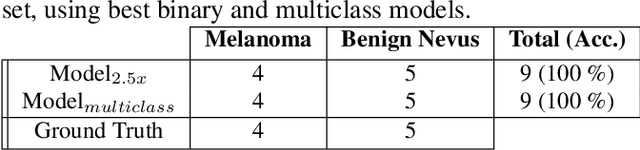
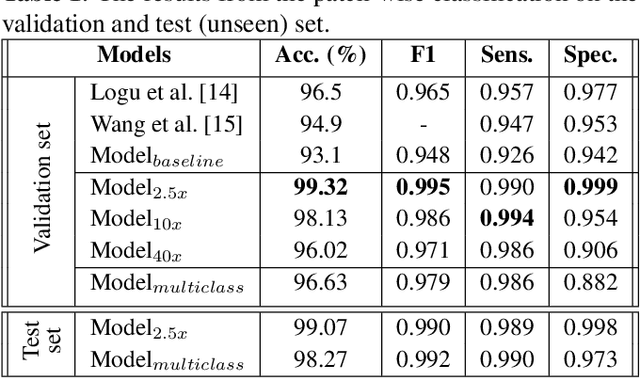
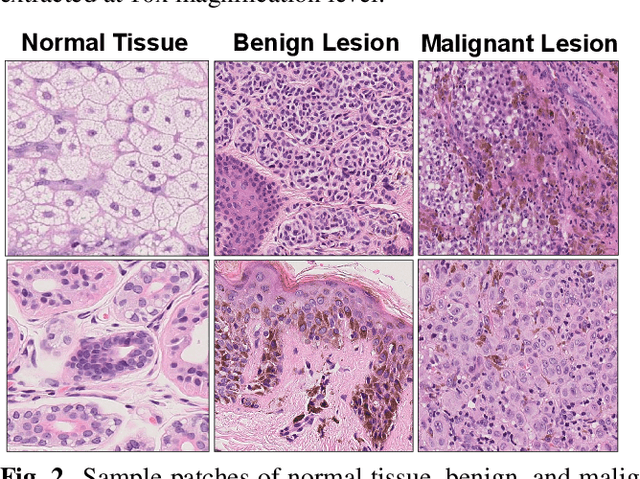

Melanoma diagnosed and treated in its early stages can increase the survival rate. A projected increase in skin cancer incidents and a dearth of dermatopathologists have emphasized the need for computational pathology (CPATH) systems. CPATH systems with deep learning (DL) models have the potential to identify the presence of melanoma by exploiting underlying morphological and cellular features. This paper proposes a DL method to detect melanoma and distinguish between normal skin and benign/malignant melanocytic lesions in Whole Slide Images (WSI). Our method detects lesions with high accuracy and localizes them on a WSI to identify potential regions of interest for pathologists. Interestingly, our DL method relies on using a single CNN network to create localization maps first and use them to perform slide-level predictions to determine patients who have melanoma. Our best model provides favorable patch-wise classification results with a 0.992 F1 score and 0.99 sensitivity on unseen data. The source code is https://github.com/RogerAmundsen/Melanoma-Diagnosis-and-Localization-from-Whole-Slide-Images-using-Convolutional-Neural-Networks.
Detection and Segmentation of Pancreas using Morphological Snakes and Deep Convolutional Neural Networks
Feb 13, 2023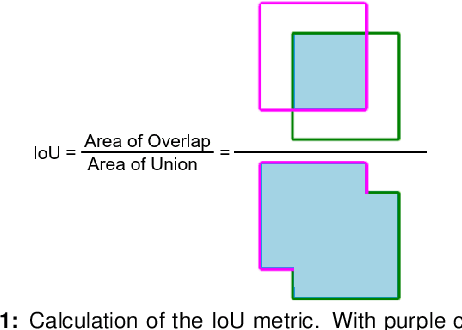
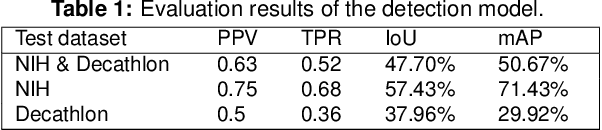
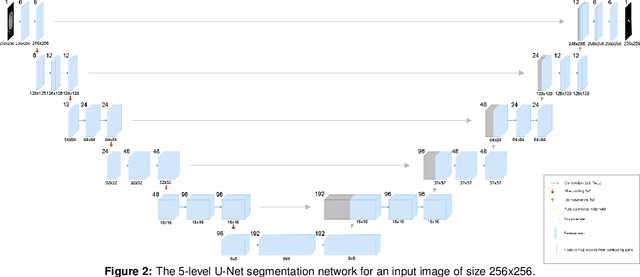
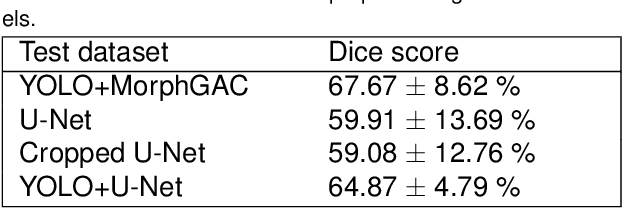
Pancreatic cancer is one of the deadliest types of cancer, with 25% of the diagnosed patients surviving for only one year and 6% of them for five. Computed tomography (CT) screening trials have played a key role in improving early detection of pancreatic cancer, which has shown significant improvement in patient survival rates. However, advanced analysis of such images often requires manual segmentation of the pancreas, which is a time-consuming task. Moreover, pancreas presents high variability in shape, while occupying only a very small area of the entire abdominal CT scans, which increases the complexity of the problem. The rapid development of deep learning can contribute to offering robust algorithms that provide inexpensive, accurate, and user-independent segmentation results that can guide the domain experts. This dissertation addresses this task by investigating a two-step approach for pancreas segmentation, by assisting the task with a prior rough localization or detection of pancreas. This rough localization of the pancreas is provided by an estimated probability map and the detection task is achieved by using the YOLOv4 deep learning algorithm. The segmentation task is tackled by a modified U-Net model applied on cropped data, as well as by using a morphological active contours algorithm. For comparison, the U-Net model was also applied on the full CT images, which provide a coarse pancreas segmentation to serve as reference. Experimental results of the detection network on the National Institutes of Health (NIH) dataset and the pancreas tumour task dataset within the Medical Segmentation Decathlon show 50.67% mean Average Precision. The best segmentation network achieved good segmentation results on the NIH dataset, reaching 67.67% Dice score.
Domain generalization in deep learning-based mass detection in mammography: A large-scale multi-center study
Jan 27, 2022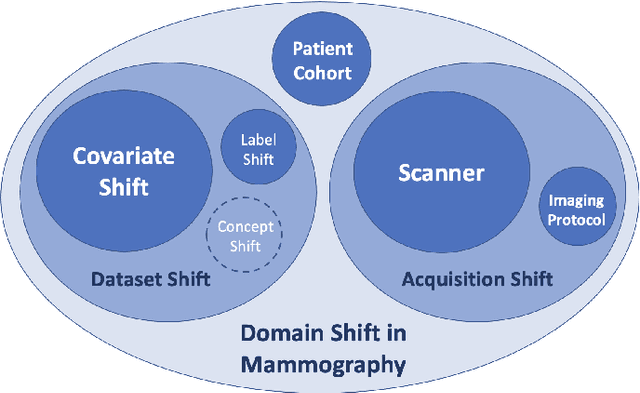
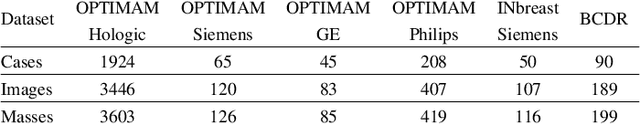

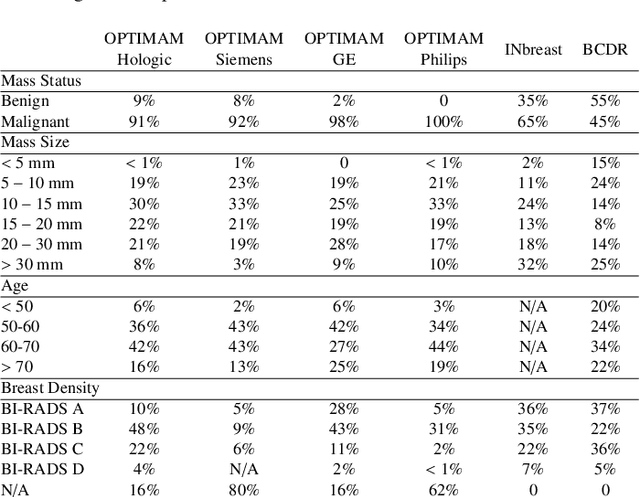
Computer-aided detection systems based on deep learning have shown great potential in breast cancer detection. However, the lack of domain generalization of artificial neural networks is an important obstacle to their deployment in changing clinical environments. In this work, we explore the domain generalization of deep learning methods for mass detection in digital mammography and analyze in-depth the sources of domain shift in a large-scale multi-center setting. To this end, we compare the performance of eight state-of-the-art detection methods, including Transformer-based models, trained in a single domain and tested in five unseen domains. Moreover, a single-source mass detection training pipeline is designed to improve the domain generalization without requiring images from the new domain. The results show that our workflow generalizes better than state-of-the-art transfer learning-based approaches in four out of five domains while reducing the domain shift caused by the different acquisition protocols and scanner manufacturers. Subsequently, an extensive analysis is performed to identify the covariate shifts with bigger effects on the detection performance, such as due to differences in patient age, breast density, mass size, and mass malignancy. Ultimately, this comprehensive study provides key insights and best practices for future research on domain generalization in deep learning-based breast cancer detection.
Improving Breast Cancer Detection using Symmetry Information with Deep Learning
Aug 17, 2018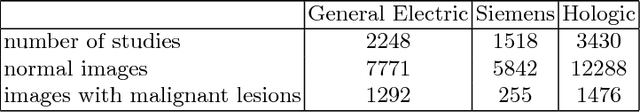
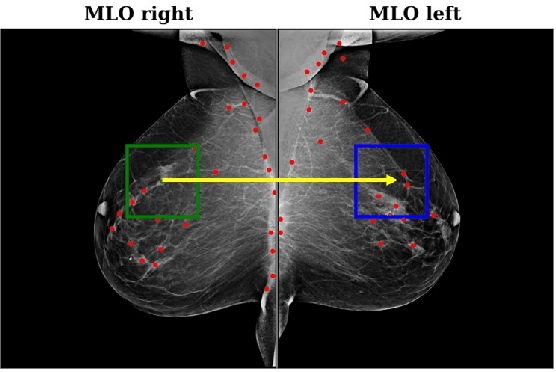
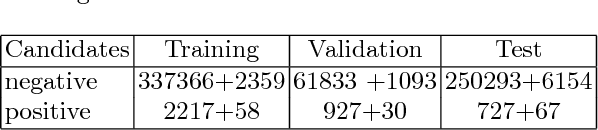
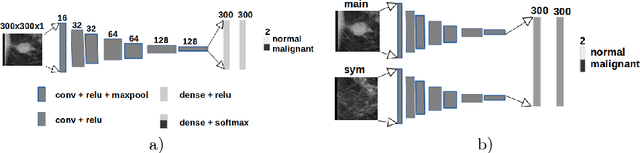
Convolutional Neural Networks (CNN) have had a huge success in many areas of computer vision and medical image analysis. However, there is still an immense potential for performance improvement in mammogram breast cancer detection Computer-Aided Detection (CAD) systems by integrating all the information that the radiologist utilizes, such as symmetry and temporal data. In this work, we proposed a patch based multi-input CNN that learns symmetrical difference to detect breast masses. The network was trained on a large-scale dataset of 28294 mammogram images. The performance was compared to a baseline architecture without symmetry context using Area Under the ROC Curve (AUC) and Competition Performance Metric (CPM). At candidate level, AUC value of 0.933 with 95% confidence interval of [0.920, 0.954] was obtained when symmetry information is incorporated in comparison with baseline architecture which yielded AUC value of 0.929 with [0.919, 0.947] confidence interval. By incorporating symmetrical information, although there was no a significant candidate level performance again (p = 0.111), we have found a compelling result at exam level with CPM value of 0.733 (p = 0.001). We believe that including temporal data, and adding benign class to the dataset could improve the detection performance.
Machine Learning Methods for Cancer Classification Using Gene Expression Data: A Review
Jan 28, 2023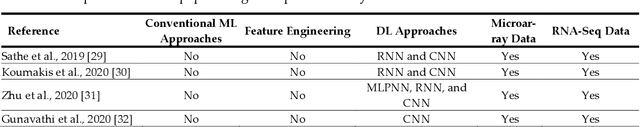
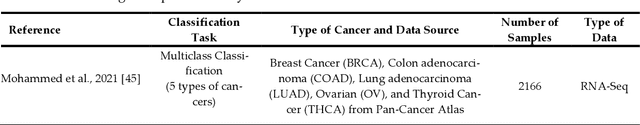
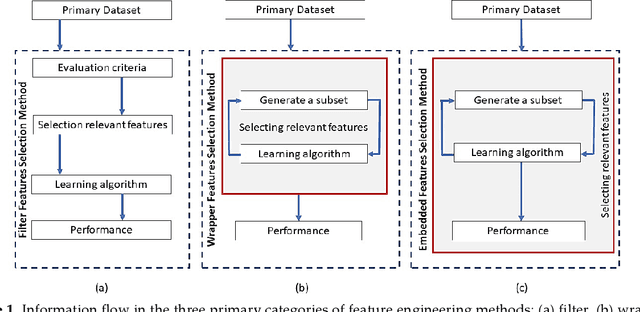
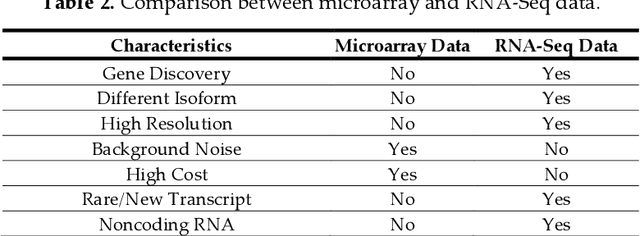

Cancer is a term that denotes a group of diseases caused by abnormal growth of cells that can spread in different parts of the body. According to the World Health Organization (WHO), cancer is the second major cause of death after cardiovascular diseases. Gene expression can play a fundamental role in the early detection of cancer, as it is indicative of the biochemical processes in tissue and cells, as well as the genetic characteristics of an organism. Deoxyribonucleic Acid (DNA) microarrays and Ribonucleic Acid (RNA)- sequencing methods for gene expression data allow quantifying the expression levels of genes and produce valuable data for computational analysis. This study reviews recent progress in gene expression analysis for cancer classification using machine learning methods. Both conventional and deep learning-based approaches are reviewed, with an emphasis on the ap-plication of deep learning models due to their comparative advantages for identifying gene patterns that are distinctive for various types of cancers. Relevant works that employ the most commonly used deep neural network architectures are covered, including multi-layer perceptrons, convolutional, recurrent, graph, and transformer networks. This survey also presents an overview of the data collection methods for gene expression analysis and lists important datasets that are commonly used for supervised machine learning for this task. Furthermore, reviewed are pertinent techniques for feature engineering and data preprocessing that are typically used to handle the high dimensionality of gene expression data, caused by a large number of genes present in data samples. The paper concludes with a discussion of future research directions for machine learning-based gene expression analysis for cancer classification.
* 29 pages, 1 figure, 11 tables
A simple normalization technique using window statistics to improve the out-of-distribution generalization in medical images
Jul 07, 2022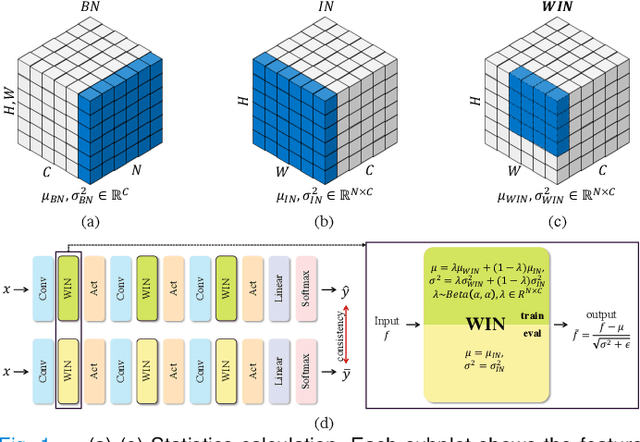
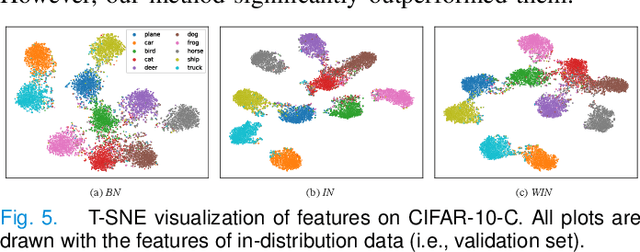
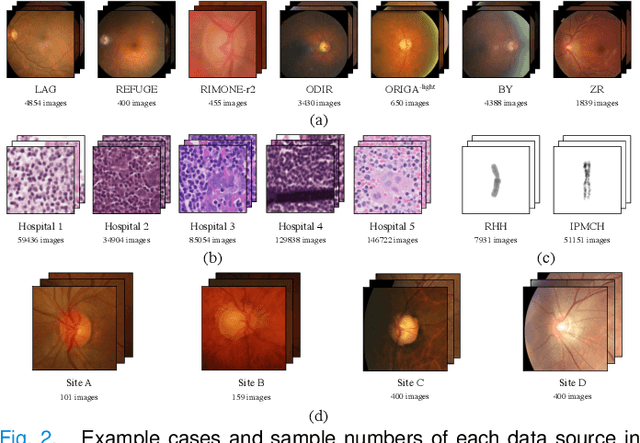

Since data scarcity and data heterogeneity are prevailing for medical images, well-trained Convolutional Neural Networks (CNNs) using previous normalization methods may perform poorly when deployed to a new site. However, a reliable model for real-world applications should be able to generalize well both on in-distribution (IND) and out-of-distribution (OOD) data (e.g., the new site data). In this study, we present a novel normalization technique called window normalization (WIN), which is a simple yet effective alternative to existing normalization methods. Specifically, WIN perturbs the normalizing statistics with the local statistics computed on a window of features. This feature-level augmentation technique regularizes the models well and improves their OOD generalization significantly. Taking its advantage, we propose a novel self-distillation method called WIN-WIN to further improve the OOD generalization in classification. WIN-WIN is easily implemented with twice forward passes and a consistency constraint, which can be a simple extension for existing methods. Extensive experimental results on various tasks (such as glaucoma detection, breast cancer detection, chromosome classification, optic disc and cup segmentation, etc.) and datasets (26 datasets) demonstrate the generality and effectiveness of our methods. The code is available at https://github.com/joe1chief/windowNormalizaion.
Multi-Head Feature Pyramid Networks for Breast Mass Detection
Feb 22, 2023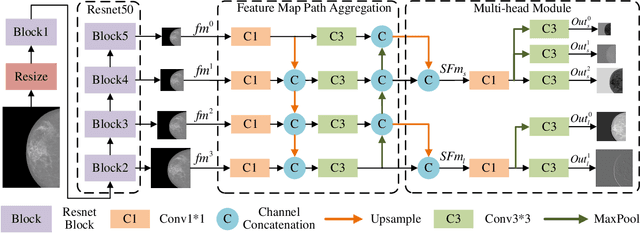
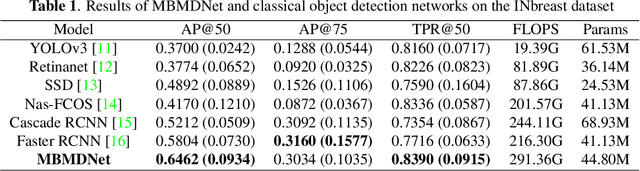
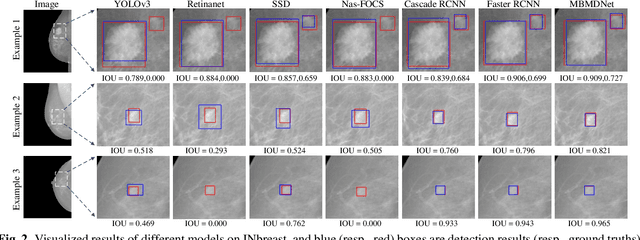
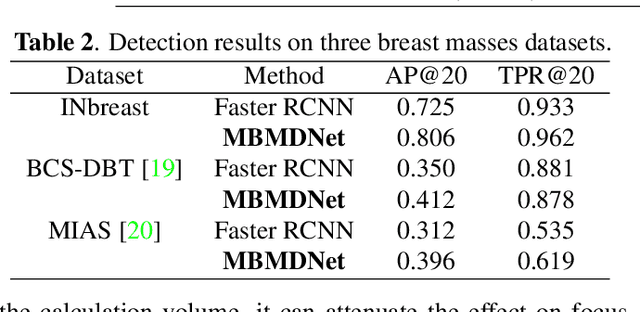
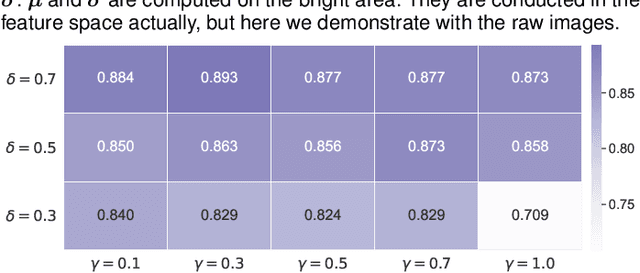
Analysis of X-ray images is one of the main tools to diagnose breast cancer. The ability to quickly and accurately detect the location of masses from the huge amount of image data is the key to reducing the morbidity and mortality of breast cancer. Currently, the main factor limiting the accuracy of breast mass detection is the unequal focus on the mass boxes, leading the network to focus too much on larger masses at the expense of smaller ones. In the paper, we propose the multi-head feature pyramid module (MHFPN) to solve the problem of unbalanced focus of target boxes during feature map fusion and design a multi-head breast mass detection network (MBMDnet). Experimental studies show that, comparing to the SOTA detection baselines, our method improves by 6.58% (in AP@50) and 5.4% (in TPR@50) on the commonly used INbreast dataset, while about 6-8% improvements (in AP@20) are also observed on the public MIAS and BCS-DBT datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge