"Time": models, code, and papers
Clustering Three-Way Data with Outliers
Oct 11, 2023Matrix-variate distributions are a recent addition to the model-based clustering field, thereby making it possible to analyze data in matrix form with complex structure such as images and time series. Due to its recent appearance, there is limited literature on matrix-variate data, with even less on dealing with outliers in these models. An approach for clustering matrix-variate normal data with outliers is discussed. The approach, which uses the distribution of subset log-likelihoods, extends the OCLUST algorithm to matrix-variate normal data and uses an iterative approach to detect and trim outliers.
An On-Chip Trainable Neuron Circuit for SFQ-Based Spiking Neural Networks
Oct 11, 2023We present an on-chip trainable neuron circuit. Our proposed circuit suits bio-inspired spike-based time-dependent data computation for training spiking neural networks (SNN). The thresholds of neurons can be increased or decreased depending on the desired application-specific spike generation rate. This mechanism provides us with a flexible design and scalable circuit structure. We demonstrate the trainable neuron structure under different operating scenarios. The circuits are designed and optimized for the MIT LL SFQ5ee fabrication process. Margin values for all parameters are above 25\% with a 3GHz throughput for a 16-input neuron.
DOMINO: Domain-invariant Hyperdimensional Classification for Multi-Sensor Time Series Data
Aug 18, 2023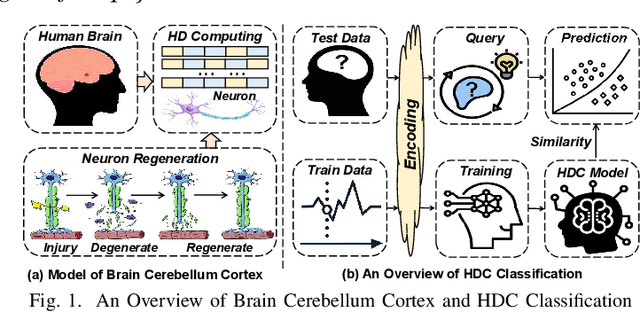
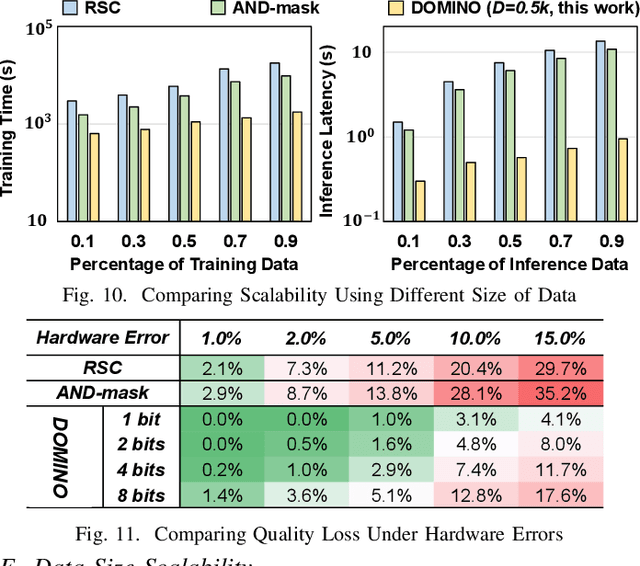
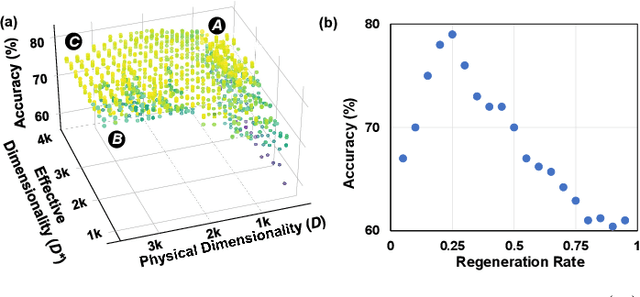
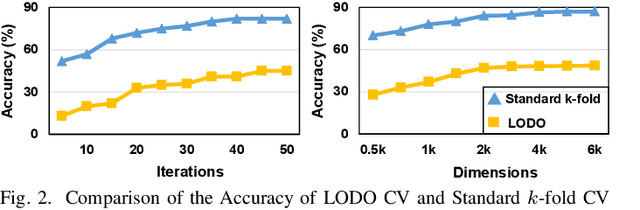
With the rapid evolution of the Internet of Things, many real-world applications utilize heterogeneously connected sensors to capture time-series information. Edge-based machine learning (ML) methodologies are often employed to analyze locally collected data. However, a fundamental issue across data-driven ML approaches is distribution shift. It occurs when a model is deployed on a data distribution different from what it was trained on, and can substantially degrade model performance. Additionally, increasingly sophisticated deep neural networks (DNNs) have been proposed to capture spatial and temporal dependencies in multi-sensor time series data, requiring intensive computational resources beyond the capacity of today's edge devices. While brain-inspired hyperdimensional computing (HDC) has been introduced as a lightweight solution for edge-based learning, existing HDCs are also vulnerable to the distribution shift challenge. In this paper, we propose DOMINO, a novel HDC learning framework addressing the distribution shift problem in noisy multi-sensor time-series data. DOMINO leverages efficient and parallel matrix operations on high-dimensional space to dynamically identify and filter out domain-variant dimensions. Our evaluation on a wide range of multi-sensor time series classification tasks shows that DOMINO achieves on average 2.04% higher accuracy than state-of-the-art (SOTA) DNN-based domain generalization techniques, and delivers 16.34x faster training and 2.89x faster inference. More importantly, DOMINO performs notably better when learning from partially labeled and highly imbalanced data, providing 10.93x higher robustness against hardware noises than SOTA DNNs.
CarExpert: Leveraging Large Language Models for In-Car Conversational Question Answering
Oct 14, 2023Large language models (LLMs) have demonstrated remarkable performance by following natural language instructions without fine-tuning them on domain-specific tasks and data. However, leveraging LLMs for domain-specific question answering suffers from severe limitations. The generated answer tends to hallucinate due to the training data collection time (when using off-the-shelf), complex user utterance and wrong retrieval (in retrieval-augmented generation). Furthermore, due to the lack of awareness about the domain and expected output, such LLMs may generate unexpected and unsafe answers that are not tailored to the target domain. In this paper, we propose CarExpert, an in-car retrieval-augmented conversational question-answering system leveraging LLMs for different tasks. Specifically, CarExpert employs LLMs to control the input, provide domain-specific documents to the extractive and generative answering components, and controls the output to ensure safe and domain-specific answers. A comprehensive empirical evaluation exhibits that CarExpert outperforms state-of-the-art LLMs in generating natural, safe and car-specific answers.
Hypernetwork-based Meta-Learning for Low-Rank Physics-Informed Neural Networks
Oct 14, 2023In various engineering and applied science applications, repetitive numerical simulations of partial differential equations (PDEs) for varying input parameters are often required (e.g., aircraft shape optimization over many design parameters) and solvers are required to perform rapid execution. In this study, we suggest a path that potentially opens up a possibility for physics-informed neural networks (PINNs), emerging deep-learning-based solvers, to be considered as one such solver. Although PINNs have pioneered a proper integration of deep-learning and scientific computing, they require repetitive time-consuming training of neural networks, which is not suitable for many-query scenarios. To address this issue, we propose a lightweight low-rank PINNs containing only hundreds of model parameters and an associated hypernetwork-based meta-learning algorithm, which allows efficient approximation of solutions of PDEs for varying ranges of PDE input parameters. Moreover, we show that the proposed method is effective in overcoming a challenging issue, known as "failure modes" of PINNs.
ARM: Refining Multivariate Forecasting with Adaptive Temporal-Contextual Learning
Oct 14, 2023Long-term time series forecasting (LTSF) is important for various domains but is confronted by challenges in handling the complex temporal-contextual relationships. As multivariate input models underperforming some recent univariate counterparts, we posit that the issue lies in the inefficiency of existing multivariate LTSF Transformers to model series-wise relationships: the characteristic differences between series are often captured incorrectly. To address this, we introduce ARM: a multivariate temporal-contextual adaptive learning method, which is an enhanced architecture specifically designed for multivariate LTSF modelling. ARM employs Adaptive Univariate Effect Learning (AUEL), Random Dropping (RD) training strategy, and Multi-kernel Local Smoothing (MKLS), to better handle individual series temporal patterns and correctly learn inter-series dependencies. ARM demonstrates superior performance on multiple benchmarks without significantly increasing computational costs compared to vanilla Transformer, thereby advancing the state-of-the-art in LTSF. ARM is also generally applicable to other LTSF architecture beyond vanilla Transformer.
Contrastive Self-Supervised Learning for Spatio-Temporal Analysis of Lung Ultrasound Videos
Oct 14, 2023Self-supervised learning (SSL) methods have shown promise for medical imaging applications by learning meaningful visual representations, even when the amount of labeled data is limited. Here, we extend state-of-the-art contrastive learning SSL methods to 2D+time medical ultrasound video data by introducing a modified encoder and augmentation method capable of learning meaningful spatio-temporal representations, without requiring constraints on the input data. We evaluate our method on the challenging clinical task of identifying lung consolidations (an important pathological feature) in ultrasound videos. Using a multi-center dataset of over 27k lung ultrasound videos acquired from over 500 patients, we show that our method can significantly improve performance on downstream localization and classification of lung consolidation. Comparisons against baseline models trained without SSL show that the proposed methods are particularly advantageous when the size of labeled training data is limited (e.g., as little as 5% of the training set).
Active Learning Framework for Cost-Effective TCR-Epitope Binding Affinity Prediction
Oct 16, 2023T cell receptors (TCRs) are critical components of adaptive immune systems, responsible for responding to threats by recognizing epitope sequences presented on host cell surface. Computational prediction of binding affinity between TCRs and epitope sequences using machine/deep learning has attracted intense attention recently. However, its success is hindered by the lack of large collections of annotated TCR-epitope pairs. Annotating their binding affinity requires expensive and time-consuming wet-lab evaluation. To reduce annotation cost, we present ActiveTCR, a framework that incorporates active learning and TCR-epitope binding affinity prediction models. Starting with a small set of labeled training pairs, ActiveTCR iteratively searches for unlabeled TCR-epitope pairs that are ''worth'' for annotation. It aims to maximize performance gains while minimizing the cost of annotation. We compared four query strategies with a random sampling baseline and demonstrated that ActiveTCR reduces annotation costs by approximately 40%. Furthermore, we showed that providing ground truth labels of TCR-epitope pairs to query strategies can help identify and reduce more than 40% redundancy among already annotated pairs without compromising model performance, enabling users to train equally powerful prediction models with less training data. Our work is the first systematic investigation of data optimization for TCR-epitope binding affinity prediction.
Detecting Speech Abnormalities with a Perceiver-based Sequence Classifier that Leverages a Universal Speech Model
Oct 16, 2023We propose a Perceiver-based sequence classifier to detect abnormalities in speech reflective of several neurological disorders. We combine this classifier with a Universal Speech Model (USM) that is trained (unsupervised) on 12 million hours of diverse audio recordings. Our model compresses long sequences into a small set of class-specific latent representations and a factorized projection is used to predict different attributes of the disordered input speech. The benefit of our approach is that it allows us to model different regions of the input for different classes and is at the same time data efficient. We evaluated the proposed model extensively on a curated corpus from the Mayo Clinic. Our model outperforms standard transformer (80.9%) and perceiver (81.8%) models and achieves an average accuracy of 83.1%. With limited task-specific data, we find that pretraining is important and surprisingly pretraining with the unrelated automatic speech recognition (ASR) task is also beneficial. Encodings from the middle layers provide a mix of both acoustic and phonetic information and achieve best prediction results compared to just using the final layer encodings (83.1% vs. 79.6%). The results are promising and with further refinements may help clinicians detect speech abnormalities without needing access to highly specialized speech-language pathologists.
DualAug: Exploiting Additional Heavy Augmentation with OOD Data Rejection
Oct 16, 2023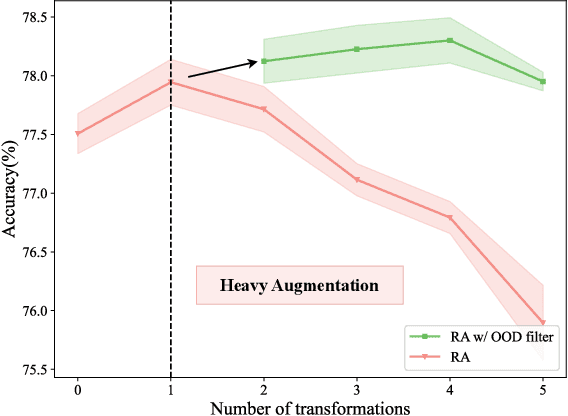
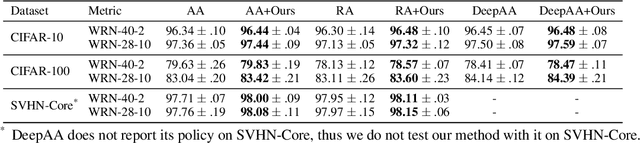

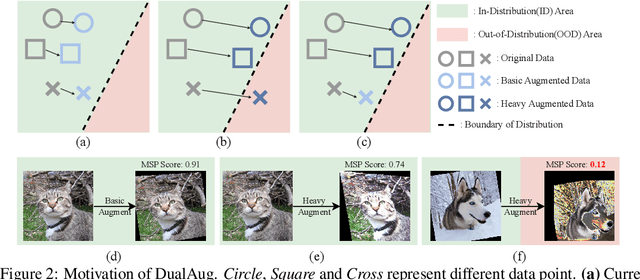
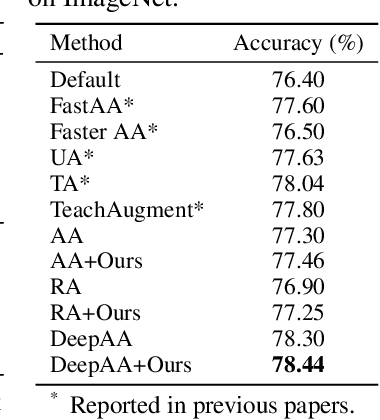
Data augmentation is a dominant method for reducing model overfitting and improving generalization. Most existing data augmentation methods tend to find a compromise in augmenting the data, \textit{i.e.}, increasing the amplitude of augmentation carefully to avoid degrading some data too much and doing harm to the model performance. We delve into the relationship between data augmentation and model performance, revealing that the performance drop with heavy augmentation comes from the presence of out-of-distribution (OOD) data. Nonetheless, as the same data transformation has different effects for different training samples, even for heavy augmentation, there remains part of in-distribution data which is beneficial to model training. Based on the observation, we propose a novel data augmentation method, named \textbf{DualAug}, to keep the augmentation in distribution as much as possible at a reasonable time and computational cost. We design a data mixing strategy to fuse augmented data from both the basic- and the heavy-augmentation branches. Extensive experiments on supervised image classification benchmarks show that DualAug improve various automated data augmentation method. Moreover, the experiments on semi-supervised learning and contrastive self-supervised learning demonstrate that our DualAug can also improve related method. Code is available at \href{https://github.com/shuguang99/DualAug}{https://github.com/shuguang99/DualAug}.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge