"Time": models, code, and papers
Bayesian Neural Networks: A Min-Max Game Framework
Nov 18, 2023Bayesian neural networks use random variables to describe the neural networks rather than deterministic neural networks and are mostly trained by variational inference which updates the mean and variance at the same time. Here, we formulate the Bayesian neural networks as a minimax game problem. We do the experiments on the MNIST data set and the primary result is comparable to the existing closed-loop transcription neural network. Finally, we reveal the connections between Bayesian neural networks and closed-loop transcription neural networks, and show our framework is rather practical, and provide another view of Bayesian neural networks.
Successive Data Injection in Conditional Quantum GAN Applied to Time Series Anomaly Detection
Oct 08, 2023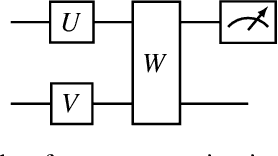
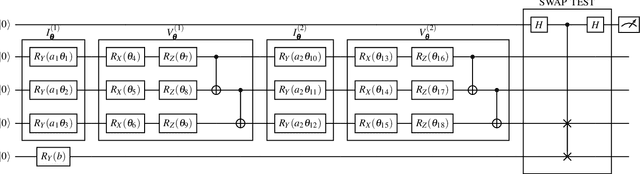
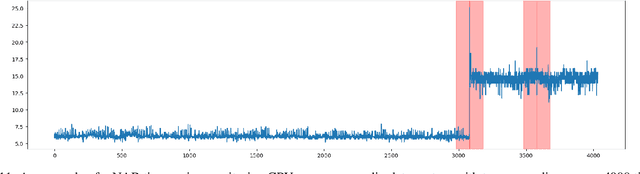
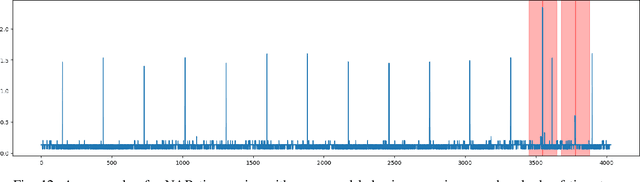
Classical GAN architectures have shown interesting results for solving anomaly detection problems in general and for time series anomalies in particular, such as those arising in communication networks. In recent years, several quantum GAN architectures have been proposed in the literature. When detecting anomalies in time series using QGANs, huge challenges arise due to the limited number of qubits compared to the size of the data. To address these challenges, we propose a new high-dimensional encoding approach, named Successive Data Injection (SuDaI). In this approach, we explore a larger portion of the quantum state than that in the conventional angle encoding, the method used predominantly in the literature, through repeated data injections into the quantum state. SuDaI encoding allows us to adapt the QGAN for anomaly detection with network data of a much higher dimensionality than with the existing known QGANs implementations. In addition, SuDaI encoding applies to other types of high-dimensional time series and can be used in contexts beyond anomaly detection and QGANs, opening up therefore multiple fields of application.
Formal concept analysis for evaluating intrinsic dimension of a natural language
Nov 17, 2023Some results of a computational experiment for determining the intrinsic dimension of linguistic varieties for the Bengali and Russian languages are presented. At the same time, both sets of words and sets of bigrams in these languages were considered separately. The method used to solve this problem was based on formal concept analysis algorithms. It was found that the intrinsic dimensions of these languages are significantly less than the dimensions used in popular neural network models in natural language processing.
Task-Distributionally Robust Data-Free Meta-Learning
Nov 23, 2023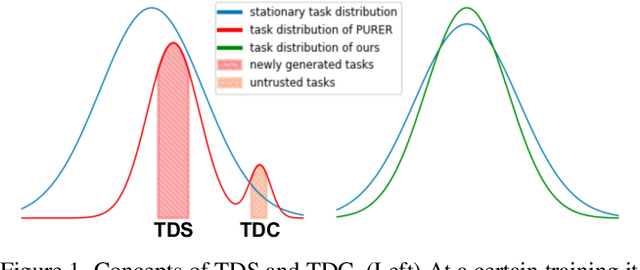
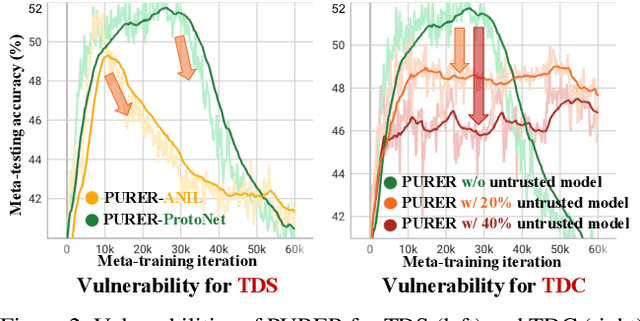
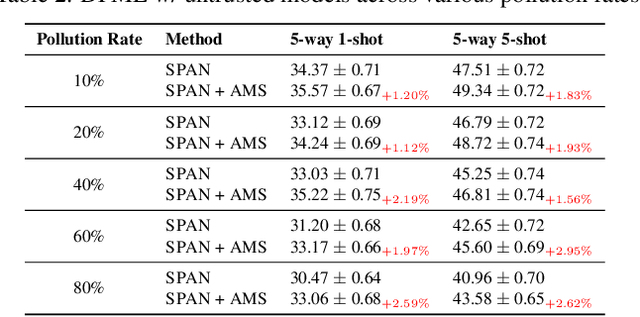
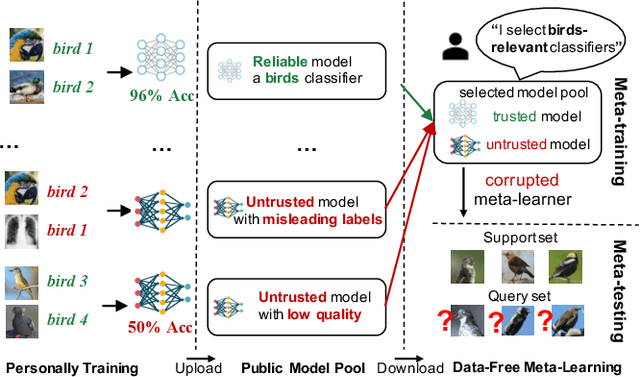
Data-Free Meta-Learning (DFML) aims to efficiently learn new tasks by leveraging multiple pre-trained models without requiring their original training data. Existing inversion-based DFML methods construct pseudo tasks from a learnable dataset, which is inversely generated from the pre-trained model pool. For the first time, we reveal two major challenges hindering their practical deployments: Task-Distribution Shift (TDS) and Task-Distribution Corruption (TDC). TDS leads to a biased meta-learner because of the skewed task distribution towards newly generated tasks. TDC occurs when untrusted models characterized by misleading labels or poor quality pollute the task distribution. To tackle these issues, we introduce a robust DFML framework that ensures task distributional robustness. We propose to meta-learn from a pseudo task distribution, diversified through task interpolation within a compact task-memory buffer. This approach reduces the meta-learner's overreliance on newly generated tasks by maintaining consistent performance across a broader range of interpolated memory tasks, thus ensuring its generalization for unseen tasks. Additionally, our framework seamlessly incorporates an automated model selection mechanism into the meta-training phase, parameterizing each model's reliability as a learnable weight. This is optimized with a policy gradient algorithm inspired by reinforcement learning, effectively addressing the non-differentiable challenge posed by model selection. Comprehensive experiments across various datasets demonstrate the framework's effectiveness in mitigating TDS and TDC, underscoring its potential to improve DFML in real-world scenarios.
Expanding the deep-learning model to diagnosis LVNC: Limitations and trade-offs
Nov 23, 2023Hyper-trabeculation or non-compaction in the left ventricle of the myocardium (LVNC) is a recently classified form of cardiomyopathy. Several methods have been proposed to quantify the trabeculae accurately in the left ventricle, but there is no general agreement in the medical community to use a particular approach. In previous work, we proposed DL-LVTQ, a deep learning approach for left ventricular trabecular quantification based on a U-Net CNN architecture. DL-LVTQ was an automatic diagnosis tool developed from a dataset of patients with the same cardiomyopathy (hypertrophic cardiomyopathy). In this work, we have extended and adapted DL-LVTQ to cope with patients with different cardiomyopathies. The dataset consists of up 379 patients in three groups with different particularities and cardiomyopathies. Patient images were taken from different scanners and hospitals. We have modified and adapted the U-Net convolutional neural network to account for the different particularities of a heterogeneous group of patients with various unclassifiable or mixed and inherited cardiomyopathies. The inclusion of new groups of patients has increased the accuracy, specificity and kappa values while maintaining the sensitivity of the automatic deep learning method proposed. Therefore, a better-prepared diagnosis tool is ready for various cardiomyopathies with different characteristics. Cardiologists have considered that 98.9% of the evaluated outputs are verified clinically for diagnosis. Therefore, the high precision to segment the different cardiac structures allows us to make a robust diagnostic system objective and faster, decreasing human error and time spent.
A Cross Attention Approach to Diagnostic Explainability using Clinical Practice Guidelines for Depression
Nov 23, 2023The lack of explainability using relevant clinical knowledge hinders the adoption of Artificial Intelligence-powered analysis of unstructured clinical dialogue. A wealth of relevant, untapped Mental Health (MH) data is available in online communities, providing the opportunity to address the explainability problem with substantial potential impact as a screening tool for both online and offline applications. We develop a method to enhance attention in popular transformer models and generate clinician-understandable explanations for classification by incorporating external clinical knowledge. Inspired by how clinicians rely on their expertise when interacting with patients, we leverage relevant clinical knowledge to model patient inputs, providing meaningful explanations for classification. This will save manual review time and engender trust. We develop such a system in the context of MH using clinical practice guidelines (CPG) for diagnosing depression, a mental health disorder of global concern. We propose an application-specific language model called ProcesS knowledge-infused cross ATtention (PSAT), which incorporates CPGs when computing attention. Through rigorous evaluation on three expert-curated datasets related to depression, we demonstrate application-relevant explainability of PSAT. PSAT also surpasses the performance of nine baseline models and can provide explanations where other baselines fall short. We transform a CPG resource focused on depression, such as the Patient Health Questionnaire (e.g. PHQ-9) and related questions, into a machine-readable ontology using SNOMED-CT. With this resource, PSAT enhances the ability of models like GPT-3.5 to generate application-relevant explanations.
AR Visualization System for Ship Detection and Recognition Based on AI
Nov 21, 2023Augmented reality technology has been widely used in industrial design interaction, exhibition guide, information retrieval and other fields. The combination of artificial intelligence and augmented reality technology has also become a future development trend. This project is an AR visualization system for ship detection and recognition based on AI, which mainly includes three parts: artificial intelligence module, Unity development module and Hololens2AR module. This project is based on R3Det algorithm to complete the detection and recognition of ships in remote sensing images. The recognition rate of model detection trained on RTX 2080Ti can reach 96%. Then, the 3D model of the ship is obtained by ship categories and information and generated in the virtual scene. At the same time, voice module and UI interaction module are added. Finally, we completed the deployment of the project on Hololens2 through MRTK. The system realizes the fusion of computer vision and augmented reality technology, which maps the results of object detection to the AR field, and makes a brave step toward the future technological trend and intelligent application.
minimax: Efficient Baselines for Autocurricula in JAX
Nov 21, 2023Unsupervised environment design (UED) is a form of automatic curriculum learning for training robust decision-making agents to zero-shot transfer into unseen environments. Such autocurricula have received much interest from the RL community. However, UED experiments, based on CPU rollouts and GPU model updates, have often required several weeks of training. This compute requirement is a major obstacle to rapid innovation for the field. This work introduces the minimax library for UED training on accelerated hardware. Using JAX to implement fully-tensorized environments and autocurriculum algorithms, minimax allows the entire training loop to be compiled for hardware acceleration. To provide a petri dish for rapid experimentation, minimax includes a tensorized grid-world based on MiniGrid, in addition to reusable abstractions for conducting autocurricula in procedurally-generated environments. With these components, minimax provides strong UED baselines, including new parallelized variants, which achieve over 120$\times$ speedups in wall time compared to previous implementations when training with equal batch sizes. The minimax library is available under the Apache 2.0 license at https://github.com/facebookresearch/minimax.
Regression-Based Analysis of Multimodal Single-Cell Data Integration Strategies
Nov 21, 2023Multimodal single-cell technologies enable the simultaneous collection of diverse data types from individual cells, enhancing our understanding of cellular states. However, the integration of these datatypes and modeling the interrelationships between modalities presents substantial computational and analytical challenges in disease biomarker detection and drug discovery. Established practices rely on isolated methodologies to investigate individual molecular aspects separately, often resulting in inaccurate analyses. To address these obstacles, distinct Machine Learning Techniques are leveraged, each of its own kind to model the co-variation of DNA to RNA, and finally to surface proteins in single cells during hematopoietic stem cell development, which simplifies understanding of underlying cellular mechanisms and immune responses. Experiments conducted on a curated subset of a 300,000-cell time course dataset, highlights the exceptional performance of Echo State Networks, boasting a remarkable state-of-the-art correlation score of 0.94 and 0.895 on Multi-omic and CiteSeq datasets. Beyond the confines of this study, these findings hold promise for advancing comprehension of cellular differentiation and function, leveraging the potential of Machine Learning.
Infinite forecast combinations based on Dirichlet process
Nov 21, 2023Forecast combination integrates information from various sources by consolidating multiple forecast results from the target time series. Instead of the need to select a single optimal forecasting model, this paper introduces a deep learning ensemble forecasting model based on the Dirichlet process. Initially, the learning rate is sampled with three basis distributions as hyperparameters to convert the infinite mixture into a finite one. All checkpoints are collected to establish a deep learning sub-model pool, and weight adjustment and diversity strategies are developed during the combination process. The main advantage of this method is its ability to generate the required base learners through a single training process, utilizing the decaying strategy to tackle the challenge posed by the stochastic nature of gradient descent in determining the optimal learning rate. To ensure the method's generalizability and competitiveness, this paper conducts an empirical analysis using the weekly dataset from the M4 competition and explores sensitivity to the number of models to be combined. The results demonstrate that the ensemble model proposed offers substantial improvements in prediction accuracy and stability compared to a single benchmark model.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge