"Time": models, code, and papers
Building Lane-Level Maps from Aerial Images
Dec 20, 2023Detecting lane lines from sensors is becoming an increasingly significant part of autonomous driving systems. However, less development has been made on high-definition lane-level mapping based on aerial images, which could automatically build and update offline maps for auto-driving systems. To this end, our work focuses on extracting fine-level detailed lane lines together with their topological structures. This task is challenging since it requires large amounts of data covering different lane types, terrain and regions. In this paper, we introduce for the first time a large-scale aerial image dataset built for lane detection, with high-quality polyline lane annotations on high-resolution images of around 80 kilometers of road. Moreover, we developed a baseline deep learning lane detection method from aerial images, called AerialLaneNet, consisting of two stages. The first stage is to produce coarse-grained results at point level, and the second stage exploits the coarse-grained results and feature to perform the vertex-matching task, producing fine-grained lanes with topology. The experiments show our approach achieves significant improvement compared with the state-of-the-art methods on our new dataset. Our code and new dataset are available at https://github.com/Jiawei-Yao0812/AerialLaneNet.
MGAug: Multimodal Geometric Augmentation in Latent Spaces of Image Deformations
Dec 20, 2023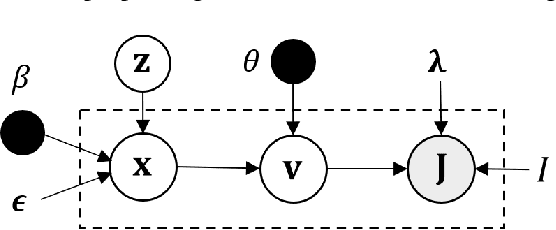
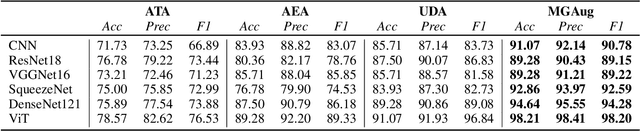
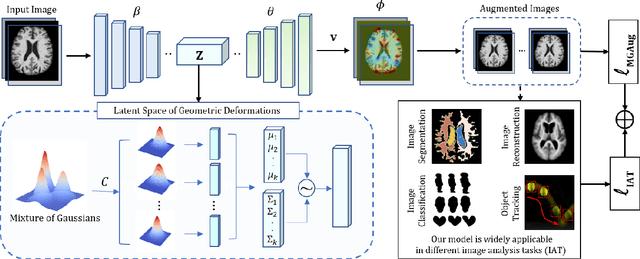
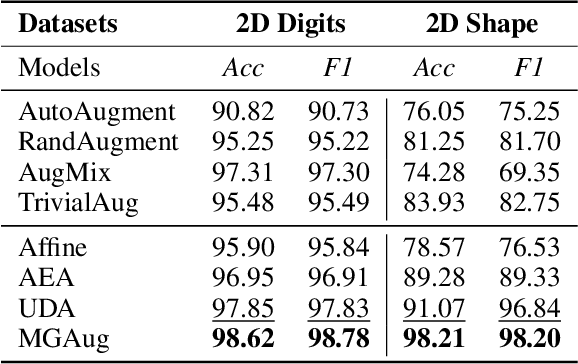
Geometric transformations have been widely used to augment the size of training images. Existing methods often assume a unimodal distribution of the underlying transformations between images, which limits their power when data with multimodal distributions occur. In this paper, we propose a novel model, Multimodal Geometric Augmentation (MGAug), that for the first time generates augmenting transformations in a multimodal latent space of geometric deformations. To achieve this, we first develop a deep network that embeds the learning of latent geometric spaces of diffeomorphic transformations (a.k.a. diffeomorphisms) in a variational autoencoder (VAE). A mixture of multivariate Gaussians is formulated in the tangent space of diffeomorphisms and serves as a prior to approximate the hidden distribution of image transformations. We then augment the original training dataset by deforming images using randomly sampled transformations from the learned multimodal latent space of VAE. To validate the efficiency of our model, we jointly learn the augmentation strategy with two distinct domain-specific tasks: multi-class classification on 2D synthetic datasets and segmentation on real 3D brain magnetic resonance images (MRIs). We also compare MGAug with state-of-the-art transformation-based image augmentation algorithms. Experimental results show that our proposed approach outperforms all baselines by significantly improved prediction accuracy. Our code is publicly available at https://github.com/tonmoy-hossain/MGAug.
MedBench: A Large-Scale Chinese Benchmark for Evaluating Medical Large Language Models
Dec 20, 2023The emergence of various medical large language models (LLMs) in the medical domain has highlighted the need for unified evaluation standards, as manual evaluation of LLMs proves to be time-consuming and labor-intensive. To address this issue, we introduce MedBench, a comprehensive benchmark for the Chinese medical domain, comprising 40,041 questions sourced from authentic examination exercises and medical reports of diverse branches of medicine. In particular, this benchmark is composed of four key components: the Chinese Medical Licensing Examination, the Resident Standardization Training Examination, the Doctor In-Charge Qualification Examination, and real-world clinic cases encompassing examinations, diagnoses, and treatments. MedBench replicates the educational progression and clinical practice experiences of doctors in Mainland China, thereby establishing itself as a credible benchmark for assessing the mastery of knowledge and reasoning abilities in medical language learning models. We perform extensive experiments and conduct an in-depth analysis from diverse perspectives, which culminate in the following findings: (1) Chinese medical LLMs underperform on this benchmark, highlighting the need for significant advances in clinical knowledge and diagnostic precision. (2) Several general-domain LLMs surprisingly possess considerable medical knowledge. These findings elucidate both the capabilities and limitations of LLMs within the context of MedBench, with the ultimate goal of aiding the medical research community.
Real-time unobtrusive sleep monitoring of in-patients with affective disorders: a feasibility study
Nov 22, 2023
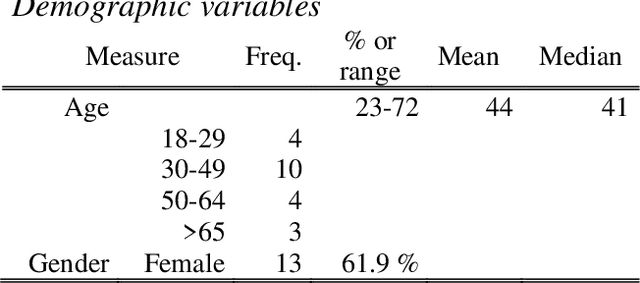
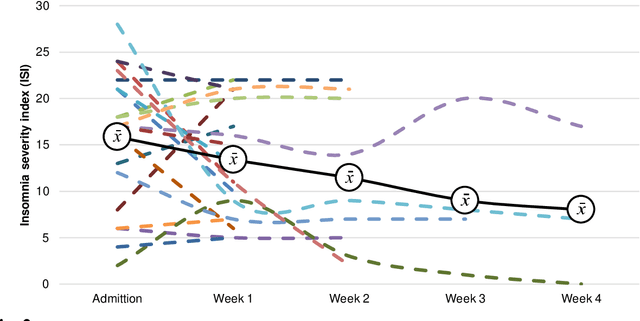
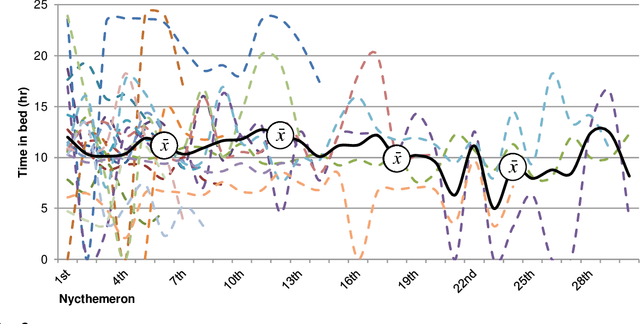
Sleep and mental health are highly related concepts, and it is an important research and clinical priority to understand their interactions. In-bed sensors using ballistocardiography provide the possibility of unobtrusive measurements of sleep. In this study, we examined the feasibility of ballistocardiography in measuring key aspects of sleep in psychiatric in-patients. Specifically, we examined a sample of patients diagnosed with depression and bipolar disorder. The subjective experiences of the researchers conducting the study are explored and descriptive analyses of patient sleep are subsequently presented. The practicalities of using the ballistocardiography device seem to be favourable. There were no apparent issues regarding data quality or data integrity. Of clinical interest, we found no link between length of stay and reduced time in bed (b = -0.06, SE = 0.03, t = -1.76, p = .08). Using ballistocardiography for measurements on in-patients with affective disorders seems to be a feasible approach.
Efficient Online Learning of Contact Force Models for Connector Insertion
Dec 14, 2023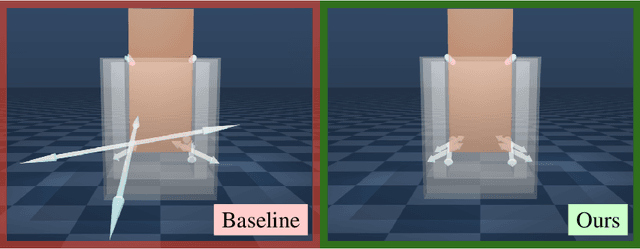
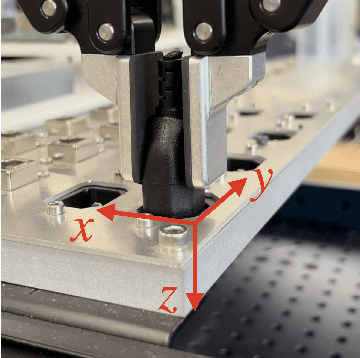
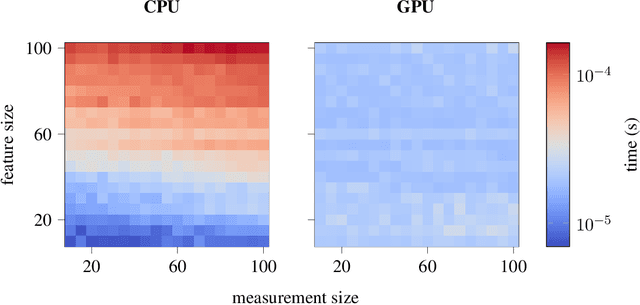
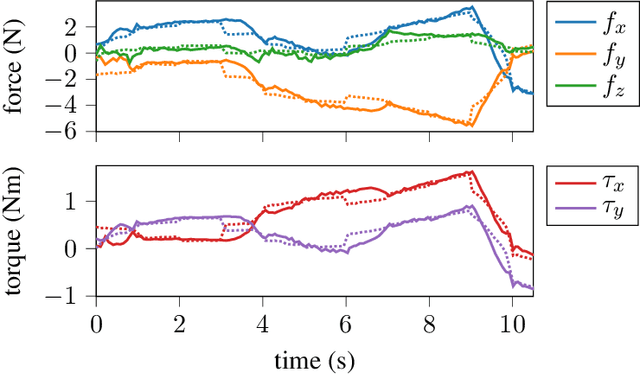

Contact-rich manipulation tasks with stiff frictional elements like connector insertion are difficult to model with rigid-body simulators. In this work, we propose a new approach for modeling these environments by learning a quasi-static contact force model instead of a full simulator. Using a feature vector that contains information about the configuration and control, we find a linear mapping adequately captures the relationship between this feature vector and the sensed contact forces. A novel Linear Model Learning (LML) algorithm is used to solve for the globally optimal mapping in real time without any matrix inversions, resulting in an algorithm that runs in nearly constant time on a GPU as the model size increases. We validate the proposed approach for connector insertion both in simulation and hardware experiments, where the learned model is combined with an optimization-based controller to achieve smooth insertions in the presence of misalignments and uncertainty. Our website featuring videos, code, and more materials is available at https://model-based-plugging.github.io/.
Towards Efficient Quantum Anomaly Detection: One-Class SVMs using Variable Subsampling and Randomized Measurements
Dec 14, 2023Quantum computing, with its potential to enhance various machine learning tasks, allows significant advancements in kernel calculation and model precision. Utilizing the one-class Support Vector Machine alongside a quantum kernel, known for its classically challenging representational capacity, notable improvements in average precision compared to classical counterparts were observed in previous studies. Conventional calculations of these kernels, however, present a quadratic time complexity concerning data size, posing challenges in practical applications. To mitigate this, we explore two distinct approaches: utilizing randomized measurements to evaluate the quantum kernel and implementing the variable subsampling ensemble method, both targeting linear time complexity. Experimental results demonstrate a substantial reduction in training and inference times by up to 95\% and 25\% respectively, employing these methods. Although unstable, the average precision of randomized measurements discernibly surpasses that of the classical Radial Basis Function kernel, suggesting a promising direction for further research in scalable, efficient quantum computing applications in machine learning.
Solving Long-run Average Reward Robust MDPs via Stochastic Games
Dec 21, 2023Markov decision processes (MDPs) provide a standard framework for sequential decision making under uncertainty. However, transition probabilities in MDPs are often estimated from data and MDPs do not take data uncertainty into account. Robust Markov decision processes (RMDPs) address this shortcoming of MDPs by assigning to each transition an uncertainty set rather than a single probability value. The goal of solving RMDPs is then to find a policy which maximizes the worst-case performance over the uncertainty sets. In this work, we consider polytopic RMDPs in which all uncertainty sets are polytopes and study the problem of solving long-run average reward polytopic RMDPs. Our focus is on computational complexity aspects and efficient algorithms. We present a novel perspective on this problem and show that it can be reduced to solving long-run average reward turn-based stochastic games with finite state and action spaces. This reduction allows us to derive several important consequences that were hitherto not known to hold for polytopic RMDPs. First, we derive new computational complexity bounds for solving long-run average reward polytopic RMDPs, showing for the first time that the threshold decision problem for them is in NP coNP and that they admit a randomized algorithm with sub-exponential expected runtime. Second, we present Robust Polytopic Policy Iteration (RPPI), a novel policy iteration algorithm for solving long-run average reward polytopic RMDPs. Our experimental evaluation shows that RPPI is much more efficient in solving long-run average reward polytopic RMDPs compared to state-of-the-art methods based on value iteration.
Hunting imaging biomarkers in pulmonary fibrosis: Benchmarks of the AIIB23 challenge
Dec 21, 2023Airway-related quantitative imaging biomarkers are crucial for examination, diagnosis, and prognosis in pulmonary diseases. However, the manual delineation of airway trees remains prohibitively time-consuming. While significant efforts have been made towards enhancing airway modelling, current public-available datasets concentrate on lung diseases with moderate morphological variations. The intricate honeycombing patterns present in the lung tissues of fibrotic lung disease patients exacerbate the challenges, often leading to various prediction errors. To address this issue, the 'Airway-Informed Quantitative CT Imaging Biomarker for Fibrotic Lung Disease 2023' (AIIB23) competition was organized in conjunction with the official 2023 International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI). The airway structures were meticulously annotated by three experienced radiologists. Competitors were encouraged to develop automatic airway segmentation models with high robustness and generalization abilities, followed by exploring the most correlated QIB of mortality prediction. A training set of 120 high-resolution computerised tomography (HRCT) scans were publicly released with expert annotations and mortality status. The online validation set incorporated 52 HRCT scans from patients with fibrotic lung disease and the offline test set included 140 cases from fibrosis and COVID-19 patients. The results have shown that the capacity of extracting airway trees from patients with fibrotic lung disease could be enhanced by introducing voxel-wise weighted general union loss and continuity loss. In addition to the competitive image biomarkers for prognosis, a strong airway-derived biomarker (Hazard ratio>1.5, p<0.0001) was revealed for survival prognostication compared with existing clinical measurements, clinician assessment and AI-based biomarkers.
Inferring the Graph of Networked Dynamical Systems under Partial Observability and Spatially Colored Noise
Dec 18, 2023In a Networked Dynamical System (NDS), each node is a system whose dynamics are coupled with the dynamics of neighboring nodes. The global dynamics naturally builds on this network of couplings and it is often excited by a noise input with nontrivial structure. The underlying network is unknown in many applications and should be inferred from observed data. We assume: i) Partial observability -- time series data is only available over a subset of the nodes; ii) Input noise -- it is correlated across distinct nodes while temporally independent, i.e., it is spatially colored. We present a feasibility condition on the noise correlation structure wherein there exists a consistent network inference estimator to recover the underlying fundamental dependencies among the observed nodes. Further, we describe a structure identification algorithm that exhibits competitive performance across distinct regimes of network connectivity, observability, and noise correlation.
Online Variational Sequential Monte Carlo
Dec 19, 2023Being the most classical generative model for serial data, state-space models (SSM) are fundamental in AI and statistical machine learning. In SSM, any form of parameter learning or latent state inference typically involves the computation of complex latent-state posteriors. In this work, we build upon the variational sequential Monte Carlo (VSMC) method, which provides computationally efficient and accurate model parameter estimation and Bayesian latent-state inference by combining particle methods and variational inference. While standard VSMC operates in the offline mode, by re-processing repeatedly a given batch of data, we distribute the approximation of the gradient of the VSMC surrogate ELBO in time using stochastic approximation, allowing for online learning in the presence of streams of data. This results in an algorithm, online VSMC, that is capable of performing efficiently, entirely on-the-fly, both parameter estimation and particle proposal adaptation. In addition, we provide rigorous theoretical results describing the algorithm's convergence properties as the number of data tends to infinity as well as numerical illustrations of its excellent convergence properties and usefulness also in batch-processing settings.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge