"Time": models, code, and papers
Spacecraft Collision Avoidance Challenge: design and results of a machine learning competition
Aug 07, 2020
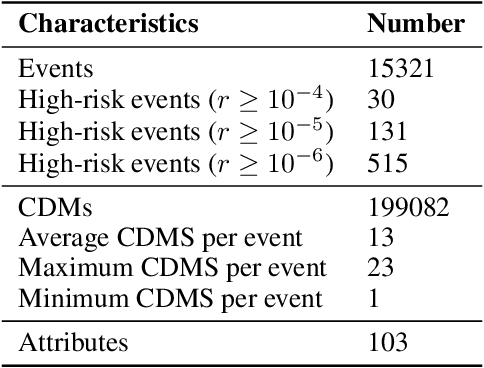
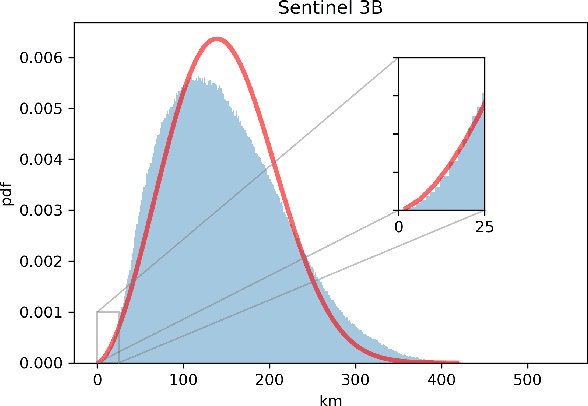

Spacecraft collision avoidance procedures have become an essential part of satellite operations. Complex and constantly updated estimates of the collision risk between orbiting objects inform the various operators who can then plan risk mitigation measures. Such measures could be aided by the development of suitable machine learning models predicting, for example, the evolution of the collision risk in time. In an attempt to study this opportunity, the European Space Agency released, in October 2019, a large curated dataset containing information about close approach events, in the form of Conjunction Data Messages (CDMs), collected from 2015 to 2019. This dataset was used in the Spacecraft Collision Avoidance Challenge, a machine learning competition where participants had to build models to predict the final collision risk between orbiting objects. This paper describes the design and results of the competition and discusses the challenges and lessons learned when applying machine learning methods to this problem domain.
Real-Time Hand Tracking Using a Sum of Anisotropic Gaussians Model
Feb 11, 2016
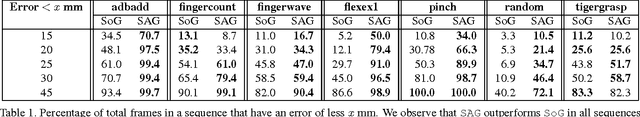
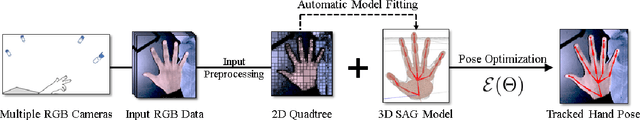
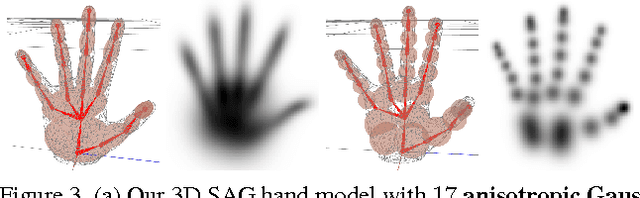
Real-time marker-less hand tracking is of increasing importance in human-computer interaction. Robust and accurate tracking of arbitrary hand motion is a challenging problem due to the many degrees of freedom, frequent self-occlusions, fast motions, and uniform skin color. In this paper, we propose a new approach that tracks the full skeleton motion of the hand from multiple RGB cameras in real-time. The main contributions include a new generative tracking method which employs an implicit hand shape representation based on Sum of Anisotropic Gaussians (SAG), and a pose fitting energy that is smooth and analytically differentiable making fast gradient based pose optimization possible. This shape representation, together with a full perspective projection model, enables more accurate hand modeling than a related baseline method from literature. Our method achieves better accuracy than previous methods and runs at 25 fps. We show these improvements both qualitatively and quantitatively on publicly available datasets.
* 8 pages, Accepted version of paper published at 3DV 2014
Regularized Flexible Activation Function Combinations for Deep Neural Networks
Aug 19, 2020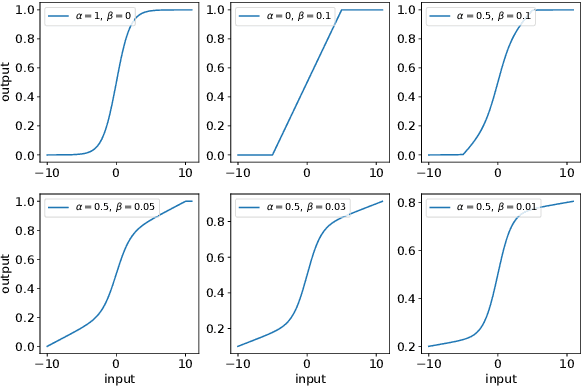
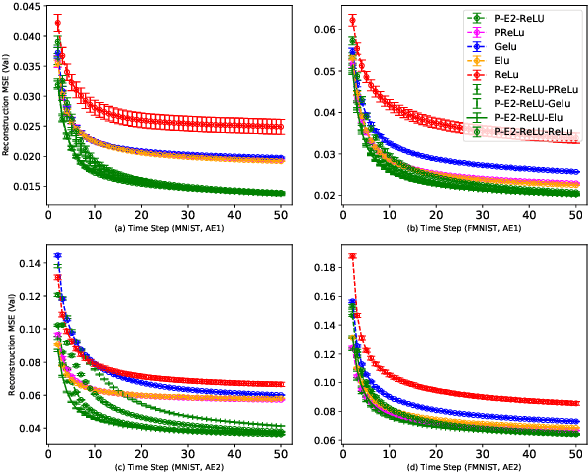
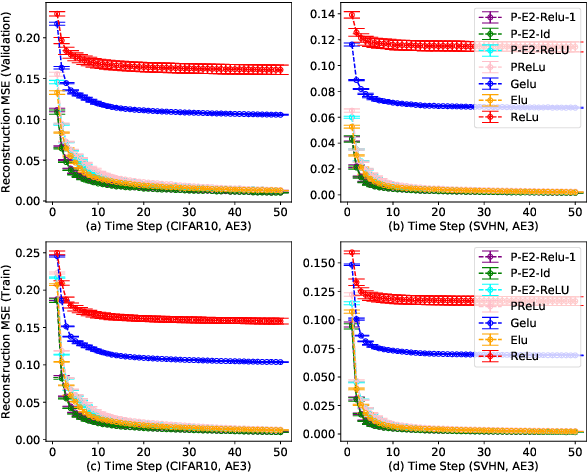
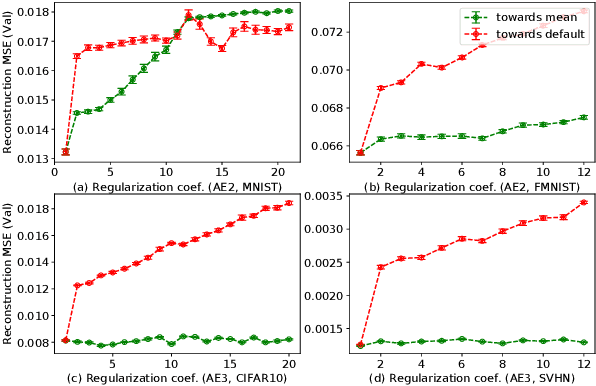
Activation in deep neural networks is fundamental to achieving non-linear mappings. Traditional studies mainly focus on finding fixed activations for a particular set of learning tasks or model architectures. The research on flexible activation is quite limited in both designing philosophy and application scenarios. In this study, three principles of choosing flexible activation components are proposed and a general combined form of flexible activation functions is implemented. Based on this, a novel family of flexible activation functions that can replace sigmoid or tanh in LSTM cells are implemented, as well as a new family by combining ReLU and ELUs. Also, two new regularisation terms based on assumptions as prior knowledge are introduced. It has been shown that LSTM models with proposed flexible activations P-Sig-Ramp provide significant improvements in time series forecasting, while the proposed P-E2-ReLU achieves better and more stable performance on lossy image compression tasks with convolutional auto-encoders. In addition, the proposed regularization terms improve the convergence, performance and stability of the models with flexible activation functions.
Application of Structural Similarity Analysis of Visually Salient Areas and Hierarchical Clustering in the Screening of Similar Wireless Capsule Endoscopic Images
Apr 01, 2020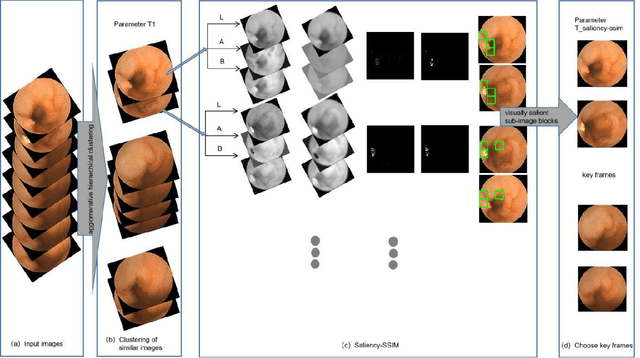
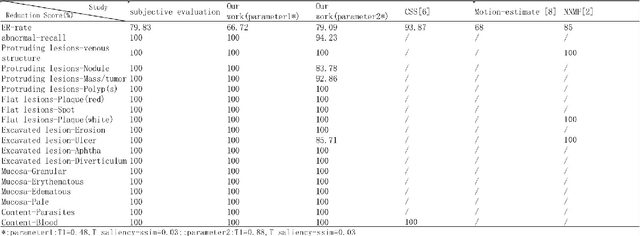
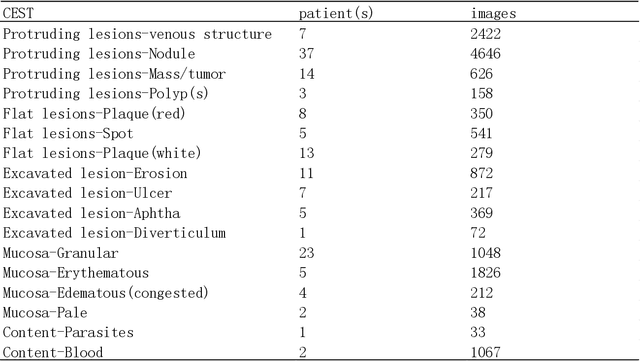

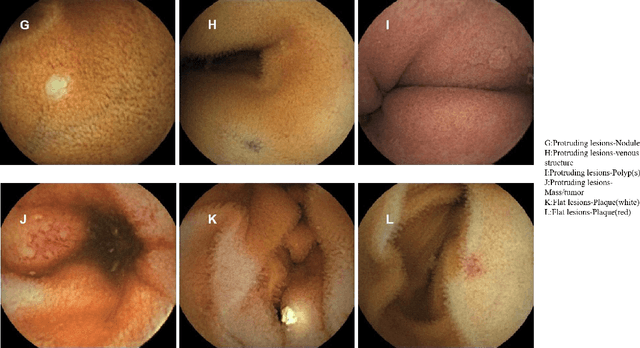
Small intestinal capsule endoscopy is the mainstream method for inspecting small intestinal lesions,but a single small intestinal capsule endoscopy will produce 60,000 - 120,000 images, the majority of which are similar and have no diagnostic value. It takes 2 - 3 hours for doctors to identify lesions from these images. This is time-consuming and increase the probability of misdiagnosis and missed diagnosis since doctors are likely to experience visual fatigue while focusing on a large number of similar images for an extended period of time.In order to solve these problems, we proposed a similar wireless capsule endoscope (WCE) image screening method based on structural similarity analysis and the hierarchical clustering of visually salient sub-image blocks. The similarity clustering of images was automatically identified by hierarchical clustering based on the hue,saturation,value (HSV) spatial color characteristics of the images,and the keyframe images were extracted based on the structural similarity of the visually salient sub-image blocks, in order to accurately identify and screen out similar small intestinal capsule endoscopic images. Subsequently, the proposed method was applied to the capsule endoscope imaging workstation. After screening out similar images in the complete data gathered by the Type I OMOM Small Intestinal Capsule Endoscope from 52 cases covering 17 common types of small intestinal lesions, we obtained a lesion recall of 100% and an average similar image reduction ratio of 76%. With similar images screened out, the average play time of the OMOM image workstation was 18 minutes, which greatly reduced the time spent by doctors viewing the images.
Tree Echo State Autoencoders with Grammars
Apr 19, 2020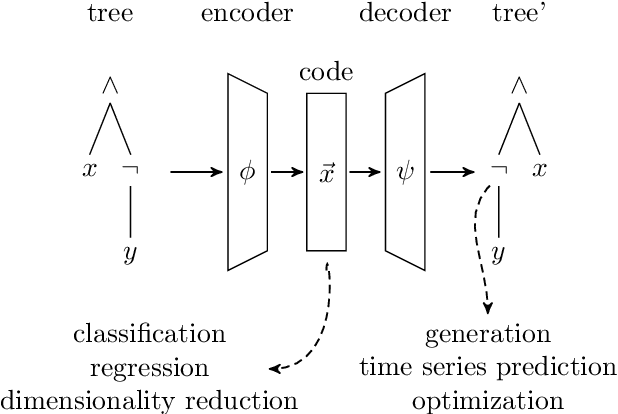
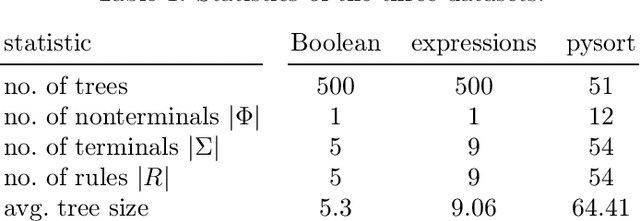

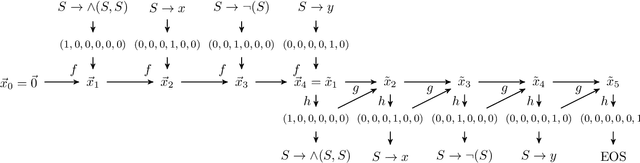
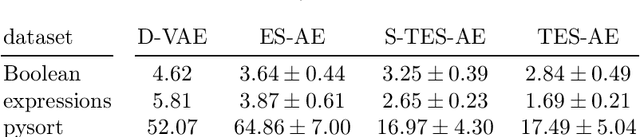
Tree data occurs in many forms, such as computer programs, chemical molecules, or natural language. Unfortunately, the non-vectorial and discrete nature of trees makes it challenging to construct functions with tree-formed output, complicating tasks such as optimization or time series prediction. Autoencoders address this challenge by mapping trees to a vectorial latent space, where tasks are easier to solve, and then mapping the solution back to a tree structure. However, existing autoencoding approaches for tree data fail to take the specific grammatical structure of tree domains into account and rely on deep learning, thus requiring large training datasets and long training times. In this paper, we propose tree echo state autoencoders (TES-AE), which are guided by a tree grammar and can be trained within seconds by virtue of reservoir computing. In our evaluation on three datasets, we demonstrate that our proposed approach is not only much faster than a state-of-the-art deep learning autoencoding approach (D-VAE) but also has less autoencoding error if little data and time is given.
Paying Per-label Attention for Multi-label Extraction from Radiology Reports
Jul 31, 2020Training medical image analysis models requires large amounts of expertly annotated data which is time-consuming and expensive to obtain. Images are often accompanied by free-text radiology reports which are a rich source of information. In this paper, we tackle the automated extraction of structured labels from head CT reports for imaging of suspected stroke patients, using deep learning. Firstly, we propose a set of 31 labels which correspond to radiographic findings (e.g. hyperdensity) and clinical impressions (e.g. haemorrhage) related to neurological abnormalities. Secondly, inspired by previous work, we extend existing state-of-the-art neural network models with a label-dependent attention mechanism. Using this mechanism and simple synthetic data augmentation, we are able to robustly extract many labels with a single model, classified according to the radiologist's reporting (positive, uncertain, negative). This approach can be used in further research to effectively extract many labels from medical text.
Detecting malicious PDF using CNN
Jul 24, 2020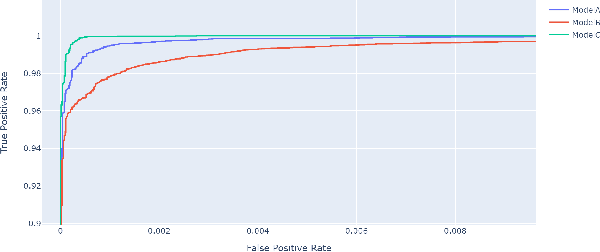
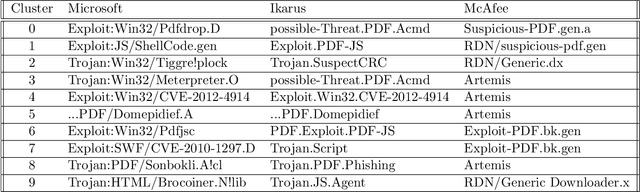
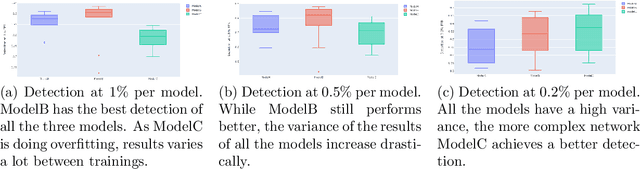
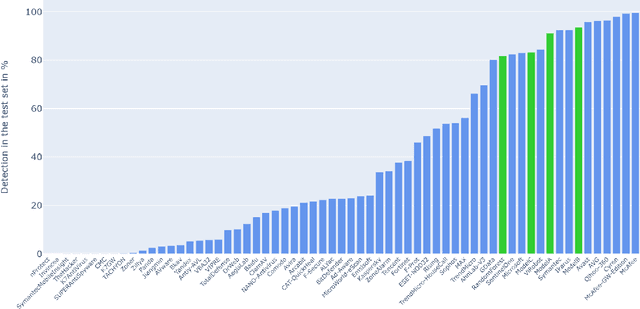
Malicious PDF files represent one of the biggest threats to computer security. To detect them, significant research has been done using handwritten signatures or machine learning based on manual feature extraction. Those approaches are both time-consuming, require significant prior knowledge and the list of features has to be updated with each newly discovered vulnerability. In this work, we propose a novel algorithm that uses an ensemble of Convolutional Neural Network (CNN) on the byte level of the file, without any handcrafted features. We show, using a data set of 90000 files downloadable online, that our approach maintains a high detection rate (94%) of PDF malware and even detects new malicious files, still undetected by most antiviruses. Using automatically generated features from our CNN network, and applying a clustering algorithm, we also obtain high similarity between the antiviruses' labels and the resulting clusters.
Comparing recurrent and convolutional neural networks for predicting wave propagation
Feb 20, 2020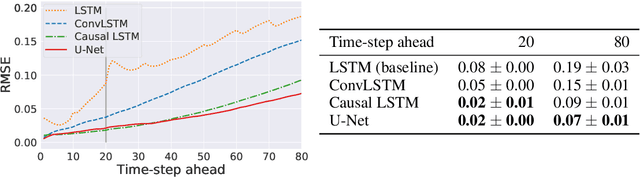
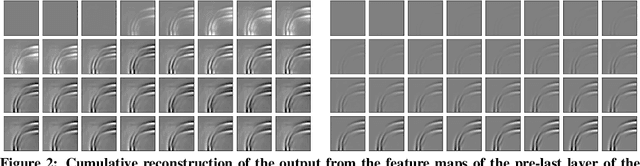

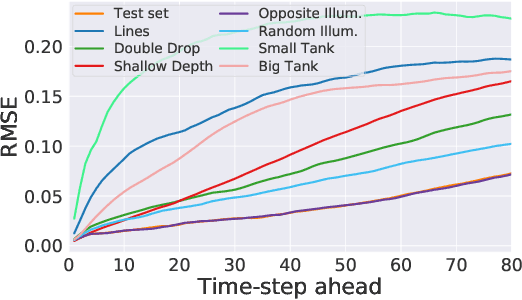
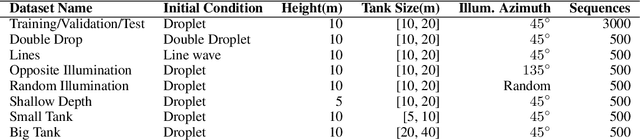
Dynamical systems can be modelled by partial differential equations and numerical computations are used everywhere in science and engineering. In this work, we investigate the performance of recurrent and convolutional deep neural network architectures to predict the surface waves. The system is governed by the Saint-Venant equations. We improve on the long-term prediction over previous methods while keeping the inference time at a fraction of numerical simulations. We also show that convolutional networks perform at least as well as recurrent networks in this task. Finally, we assess the generalisation capability of each network by extrapolating in longer time-frames and in different physical settings.
Traffic Control Gesture Recognition for Autonomous Vehicles
Jul 31, 2020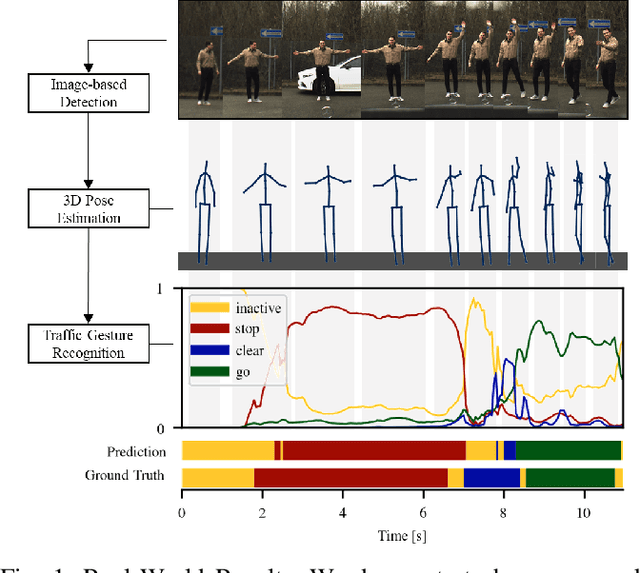
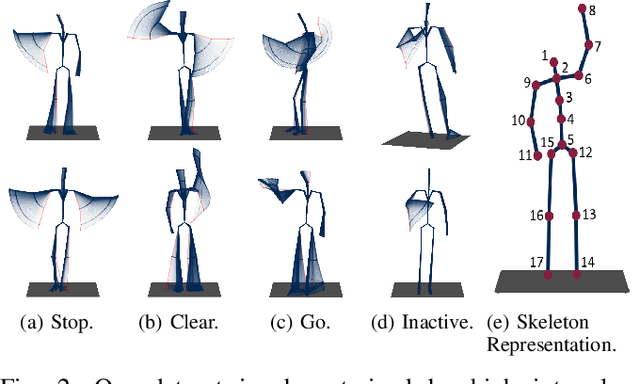
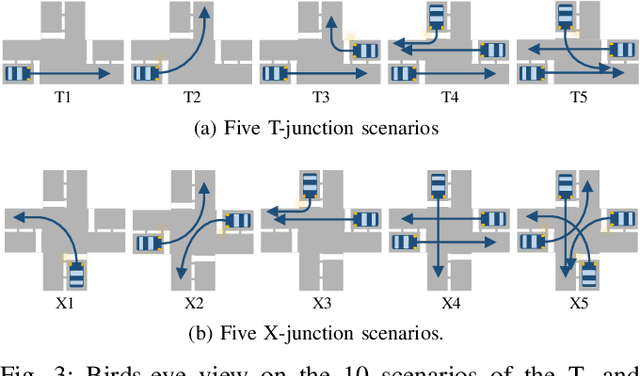
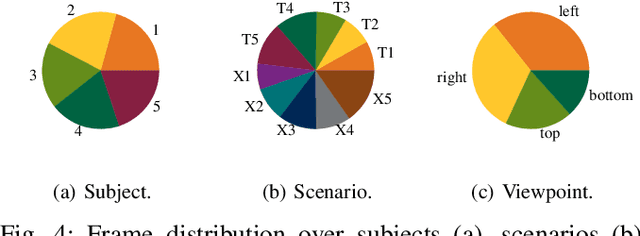
A car driver knows how to react on the gestures of the traffic officers. Clearly, this is not the case for the autonomous vehicle, unless it has road traffic control gesture recognition functionalities. In this work, we address the limitation of the existing autonomous driving datasets to provide learning data for traffic control gesture recognition. We introduce a dataset that is based on 3D body skeleton input to perform traffic control gesture classification on every time step. Our dataset consists of 250 sequences from several actors, ranging from 16 to 90 seconds per sequence. To evaluate our dataset, we propose eight sequential processing models based on deep neural networks such as recurrent networks, attention mechanism, temporal convolutional networks and graph convolutional networks. We present an extensive evaluation and analysis of all approaches for our dataset, as well as real-world quantitative evaluation. The code and dataset is publicly available.
Learning Anatomical Segmentations for Tractography from Diffusion MRI
Sep 09, 2020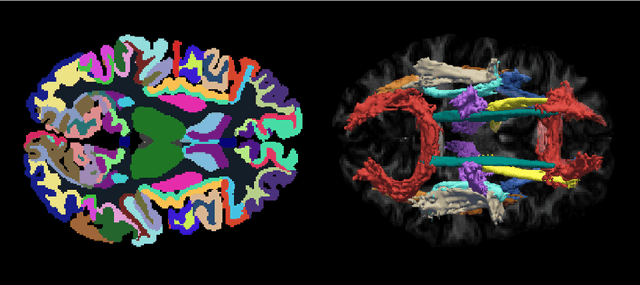
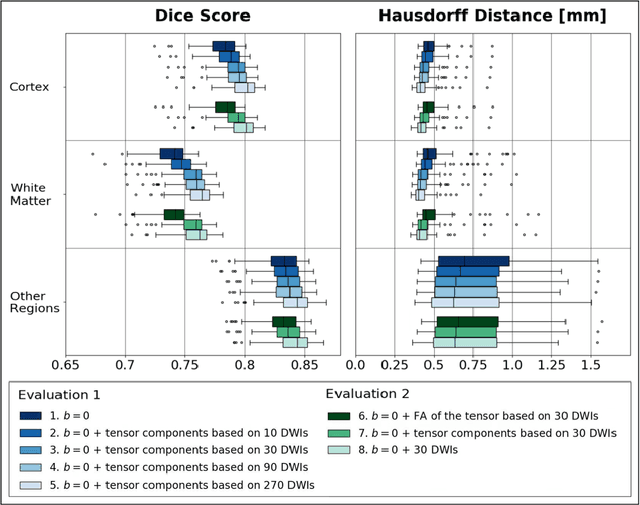
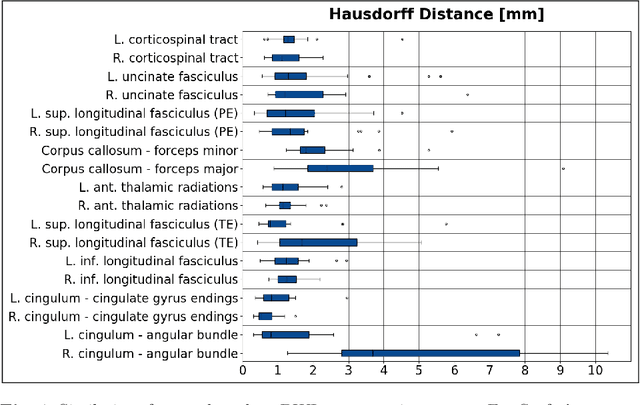
Deep learning approaches for diffusion MRI have so far focused primarily on voxel-based segmentation of lesions or white-matter fiber tracts. A drawback of representing tracts as volumetric labels, rather than sets of streamlines, is that it precludes point-wise analyses of microstructural or geometric features along a tract. Traditional tractography pipelines, which do allow such analyses, can benefit from detailed whole-brain segmentations to guide tract reconstruction. Here, we introduce fast, deep learning-based segmentation of 170 anatomical regions directly on diffusion-weighted MR images, removing the dependency of conventional segmentation methods on T 1-weighted images and slow pre-processing pipelines. Working natively in diffusion space avoids non-linear distortions and registration errors across modalities, as well as interpolation artifacts. We demonstrate consistent segmentation results between 0 .70 and 0 .87 Dice depending on the tissue type. We investigate various combinations of diffusion-derived inputs and show generalization across different numbers of gradient directions. Finally, integrating our approach to provide anatomical priors for tractography pipelines, such as TRACULA, removes hours of pre-processing time and permits processing even in the absence of high-quality T 1-weighted scans, without degrading the quality of the resulting tract estimates.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge