"Time": models, code, and papers
Semi-supervised Semantic Segmentation of Organs at Risk on 3D Pelvic CT Images
Sep 21, 2020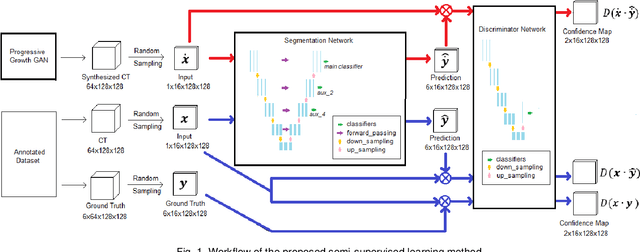

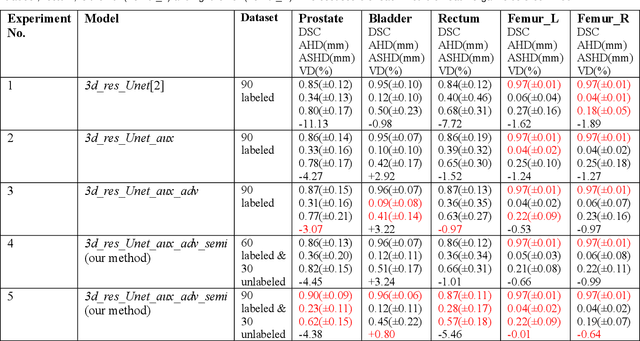
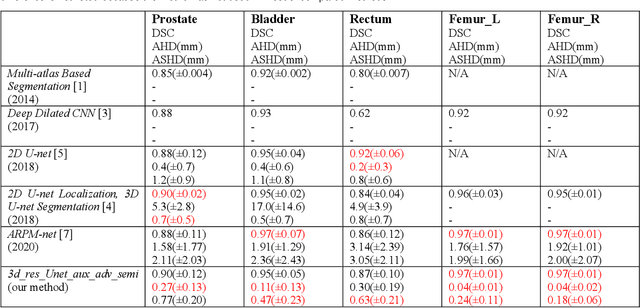
Automated segmentation of organs-at-risk in pelvic computed tomography (CT) images can assist the radiotherapy treatment planning by saving time and effort of manual contouring and reducing intra-observer and inter-observer variation. However, training high-performance deep-learning segmentation models usually requires broad labeled data, which are labor-intensive to collect. Lack of annotated data presents a significant challenge for many medical imaging-related deep learning solutions. This paper proposes a novel end-to-end convolutional neural network-based semi-supervised adversarial method that can segment multiple organs-at-risk, including prostate, bladder, rectum, left femur, and right femur. New design schemes are introduced to enhance the baseline residual U-net architecture to improve performance. Importantly, new unlabeled CT images are synthesized by a generative adversarial network (GAN) that is trained on given images to overcome the inherent problem of insufficient annotated data in practice. A semi-supervised adversarial strategy is then introduced to utilize labeled and unlabeled 3D CT images. The new method is evaluated on a dataset of 100 training cases and 20 testing cases. Experimental results, including four metrics (dice similarity coefficient, average Hausdorff distance, average surface Hausdorff distance, and relative volume difference), show that the new method outperforms several state-of-the-art segmentation approaches.
Recurrent Neural Network Control of a Hybrid Dynamic Transfemoral Prosthesis with EdgeDRNN Accelerator
Mar 05, 2020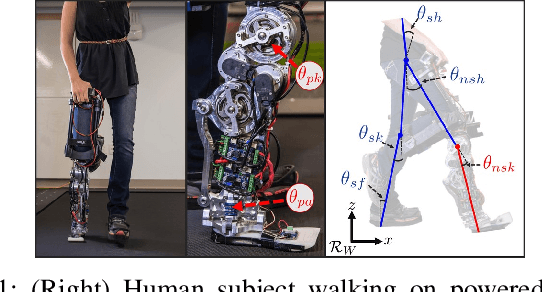
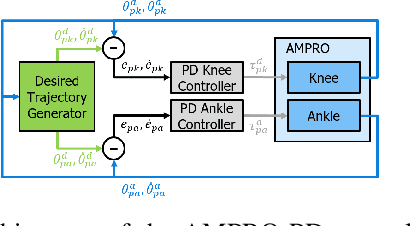
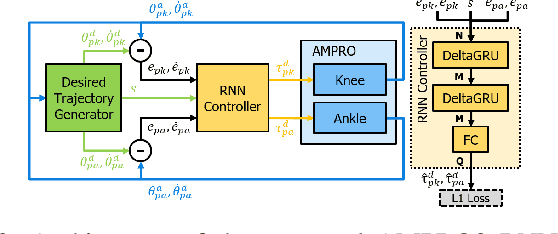
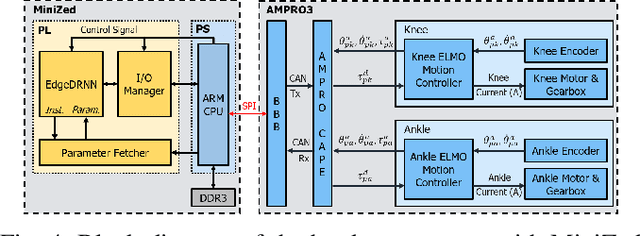
Lower leg prostheses could improve the life quality of amputees by increasing comfort and reducing energy to locomote, but currently control methods are limited in modulating behaviors based upon the human's experience. This paper describes the first steps toward learning complex controllers for dynamical robotic assistive devices. We provide the first example of behavioral cloning to control a powered transfemoral prostheses using a Gated Recurrent Unit (GRU) based recurrent neural network (RNN) running on a custom hardware accelerator that exploits temporal sparsity. The RNN is trained on data collected from the original prosthesis controller. The RNN inference is realized by a novel EdgeDRNN accelerator in real-time. Experimental results show that the RNN can replace the nominal PD controller to realize end-to-end control of the AMPRO3 prosthetic leg walking on flat ground and unforeseen slopes with comparable tracking accuracy. EdgeDRNN computes the RNN about 240 times faster than real time, opening the possibility of running larger networks for more complex tasks in the future. Implementing an RNN on this real-time dynamical system with impacts sets the ground work to incorporate other learned elements of the human-prosthesis system into prosthesis control.
Olympus: a benchmarking framework for noisy optimization and experiment planning
Oct 08, 2020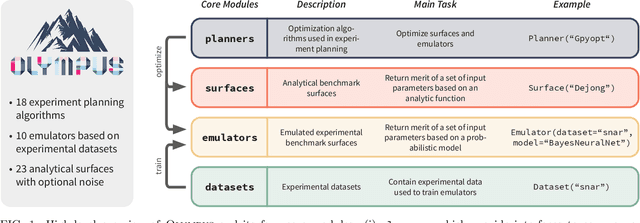
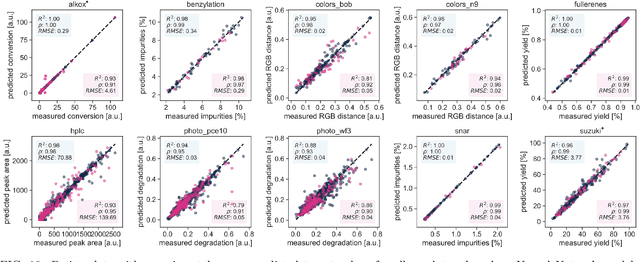
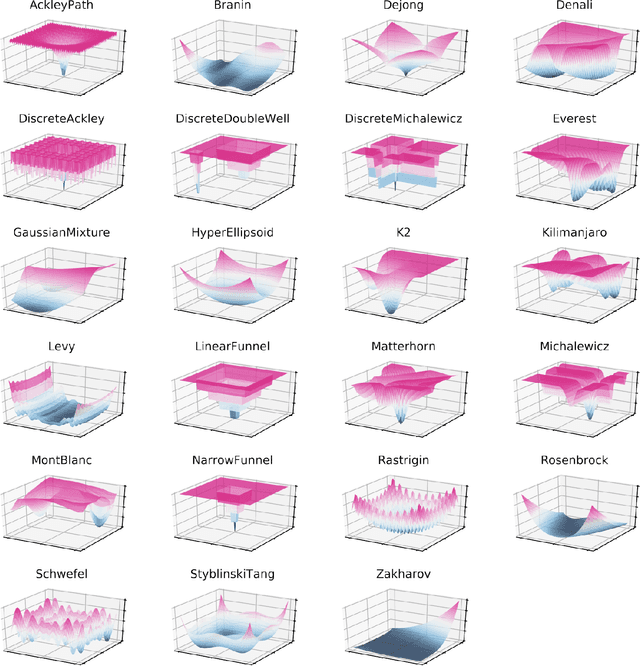
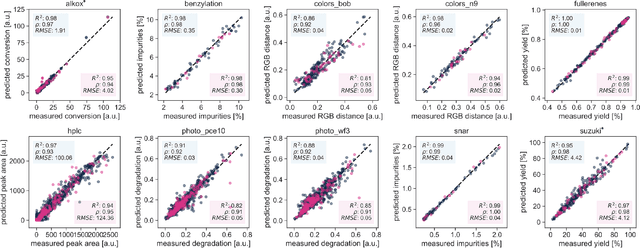
Research challenges encountered across science, engineering, and economics can frequently be formulated as optimization tasks. In chemistry and materials science, recent growth in laboratory digitization and automation has sparked interest in optimization-guided autonomous discovery and closed-loop experimentation. Experiment planning strategies based on off-the-shelf optimization algorithms can be employed in fully autonomous research platforms to achieve desired experimentation goals with the minimum number of trials. However, the experiment planning strategy that is most suitable to a scientific discovery task is a priori unknown while rigorous comparisons of different strategies are highly time and resource demanding. As optimization algorithms are typically benchmarked on low-dimensional synthetic functions, it is unclear how their performance would translate to noisy, higher-dimensional experimental tasks encountered in chemistry and materials science. We introduce Olympus, a software package that provides a consistent and easy-to-use framework for benchmarking optimization algorithms against realistic experiments emulated via probabilistic deep-learning models. Olympus includes a collection of experimentally derived benchmark sets from chemistry and materials science and a suite of experiment planning strategies that can be easily accessed via a user-friendly python interface. Furthermore, Olympus facilitates the integration, testing, and sharing of custom algorithms and user-defined datasets. In brief, Olympus mitigates the barriers associated with benchmarking optimization algorithms on realistic experimental scenarios, promoting data sharing and the creation of a standard framework for evaluating the performance of experiment planning strategies
Policy Gradient for Continuing Tasks in Non-stationary Markov Decision Processes
Oct 16, 2020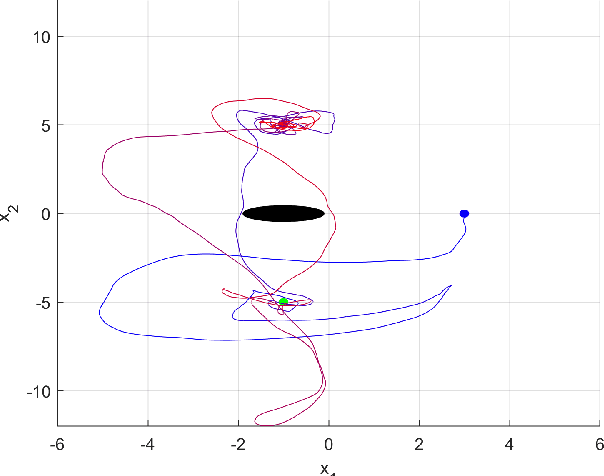
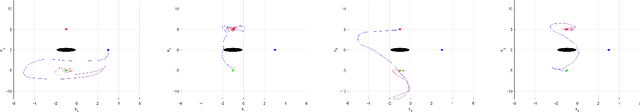
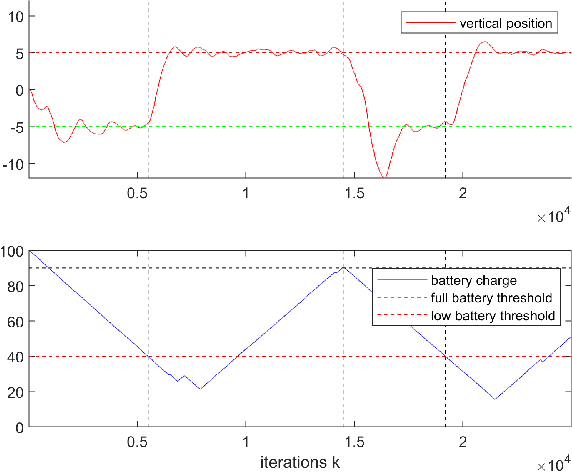
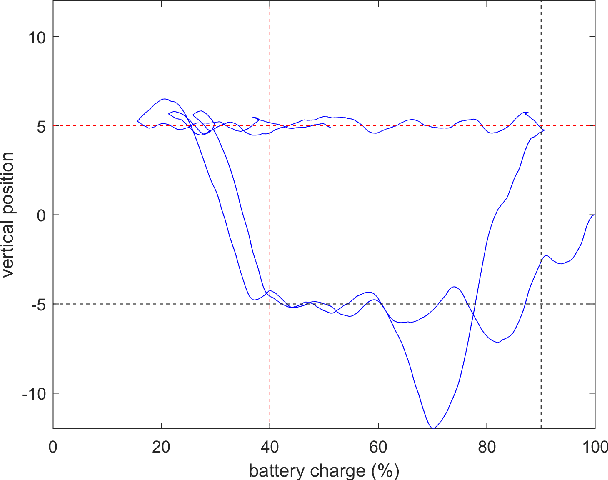
Reinforcement learning considers the problem of finding policies that maximize an expected cumulative reward in a Markov decision process with unknown transition probabilities. In this paper we consider the problem of finding optimal policies assuming that they belong to a reproducing kernel Hilbert space (RKHS). To that end we compute unbiased stochastic gradients of the value function which we use as ascent directions to update the policy. A major drawback of policy gradient-type algorithms is that they are limited to episodic tasks unless stationarity assumptions are imposed. Hence preventing these algorithms to be fully implemented online, which is a desirable property for systems that need to adapt to new tasks and/or environments in deployment. The main requirement for a policy gradient algorithm to work is that the estimate of the gradient at any point in time is an ascent direction for the initial value function. In this work we establish that indeed this is the case which enables to show the convergence of the online algorithm to the critical points of the initial value function. A numerical example shows the ability of our online algorithm to learn to solve a navigation and surveillance problem, in which an agent must loop between to goal locations. This example corroborates our theoretical findings about the ascent directions of subsequent stochastic gradients. It also shows how the agent running our online algorithm succeeds in learning to navigate, following a continuing cyclic trajectory that does not comply with the standard stationarity assumptions in the literature for non episodic training.
Waymo Public Road Safety Performance Data
Oct 30, 2020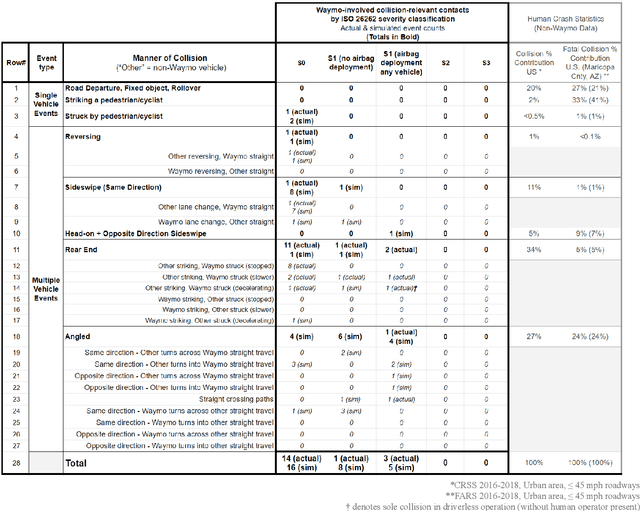

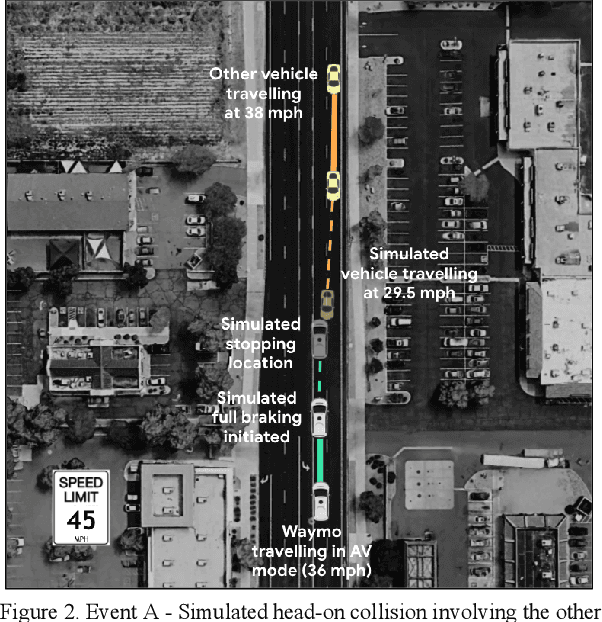
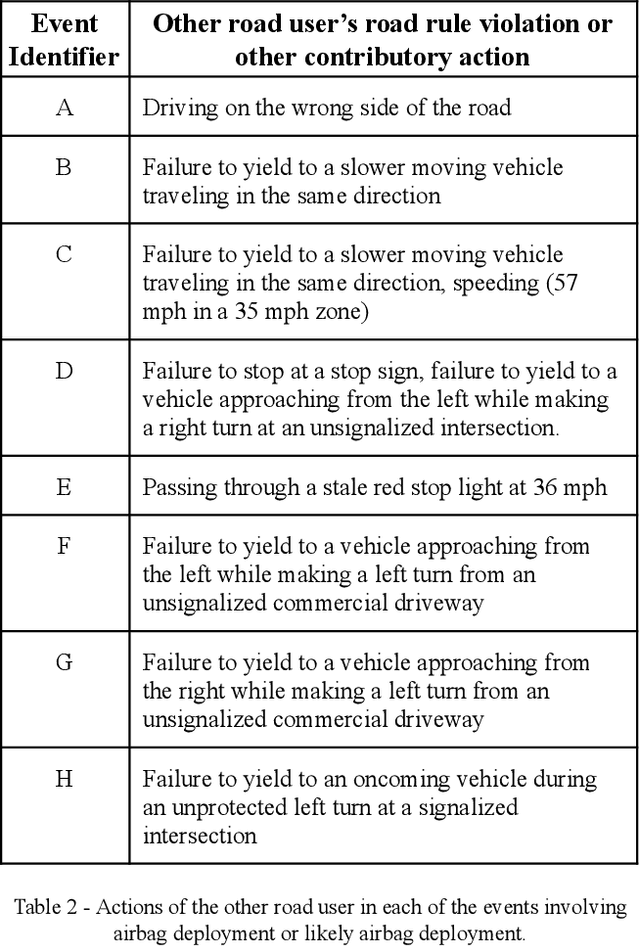
Waymo's mission to reduce traffic injuries and fatalities and improve mobility for all has led us to expand deployment of automated vehicles on public roads without a human driver behind the wheel. As part of this process, Waymo is committed to providing the public with informative and relevant data regarding the demonstrated safety of Waymo's automated driving system, which we call the Waymo Driver. The data presented in this paper represents more than 6.1 million miles of automated driving in the Phoenix, Arizona metropolitan area, including operations with a trained operator behind the steering wheel from calendar year 2019 and 65,000 miles of driverless operation without a human behind the steering wheel from 2019 and the first nine months of 2020. The paper includes every collision and minor contact experienced during these operations as well as every predicted contact identified using Waymo's counterfactual, what if, simulation of events had the vehicle's trained operator not disengaged automated driving. There were 47 contact events that occurred over this time period, consisting of 18 actual and 29 simulated contact events, none of which would be expected to result in severe or life threatening injuries. This paper presents the collision typology and severity for each actual and simulated event, along with diagrams depicting each of the most significant events. Nearly all the events involved one or more road rule violations or other errors by a human driver or road user, including all eight of the most severe events, which we define as involving actual or expected airbag deployment in any involved vehicle. When compared to national collision statistics, the Waymo Driver completely avoided certain collision modes that human driven vehicles are frequently involved in, including road departure and collisions with fixed objects.
Training Variational Networks with Multi-Domain Simulations: Speed-of-Sound Image Reconstruction
Jun 25, 2020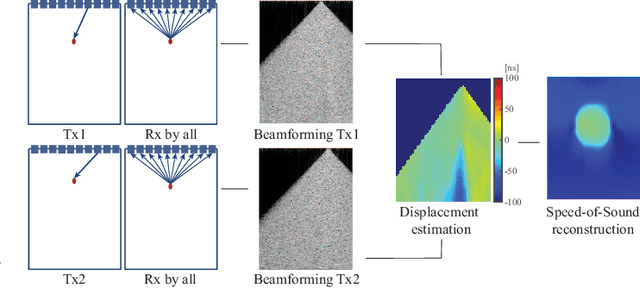
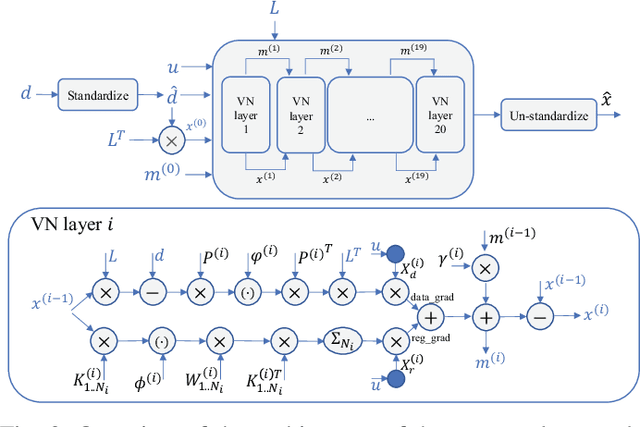
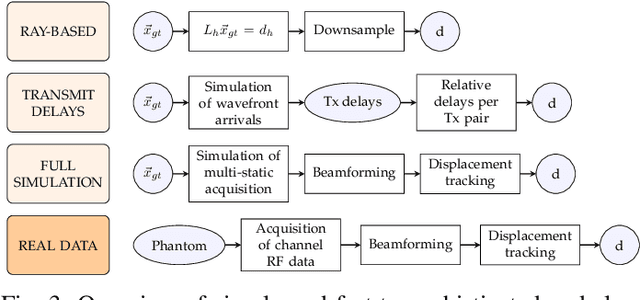
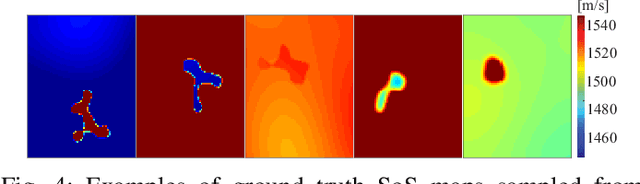
Speed-of-sound has been shown as a potential biomarker for breast cancer imaging, successfully differentiating malignant tumors from benign ones. Speed-of-sound images can be reconstructed from time-of-flight measurements from ultrasound images acquired using conventional handheld ultrasound transducers. Variational Networks (VN) have recently been shown to be a potential learning-based approach for optimizing inverse problems in image reconstruction. Despite earlier promising results, these methods however do not generalize well from simulated to acquired data, due to the domain shift. In this work, we present for the first time a VN solution for a pulse-echo SoS image reconstruction problem using diverging waves with conventional transducers and single-sided tissue access. This is made possible by incorporating simulations with varying complexity into training. We use loop unrolling of gradient descent with momentum, with an exponentially weighted loss of outputs at each unrolled iteration in order to regularize training. We learn norms as activation functions regularized to have smooth forms for robustness to input distribution variations. We evaluate reconstruction quality on ray-based and full-wave simulations as well as on tissue-mimicking phantom data, in comparison to a classical iterative (L-BFGS) optimization of this image reconstruction problem. We show that the proposed regularization techniques combined with multi-source domain training yield substantial improvements in the domain adaptation capabilities of VN, reducing median RMSE by 54% on a wave-based simulation dataset compared to the baseline VN. We also show that on data acquired from a tissue-mimicking breast phantom the proposed VN provides improved reconstruction in 12 milliseconds.
Using Overlapping Communities and Network Structure for Identifying Reduced Groups of Stress Responsive Genes
Oct 23, 2020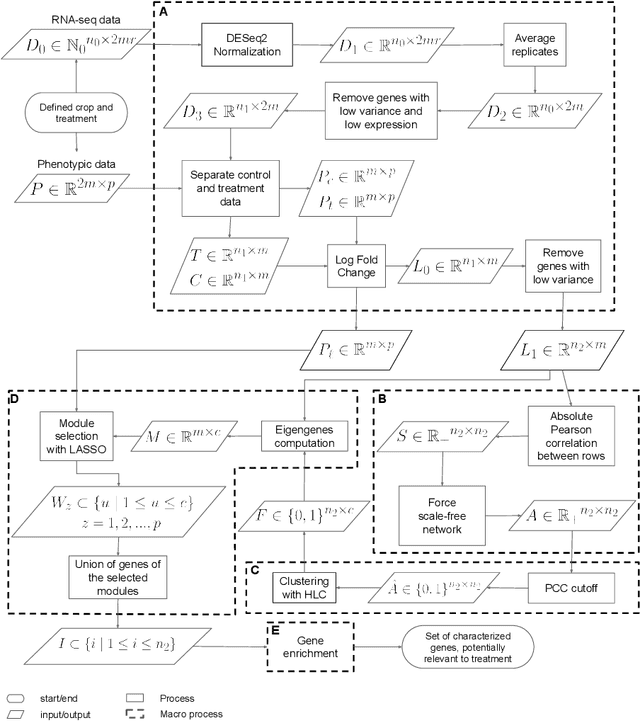
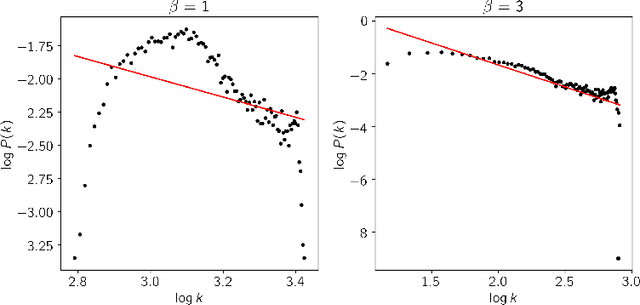
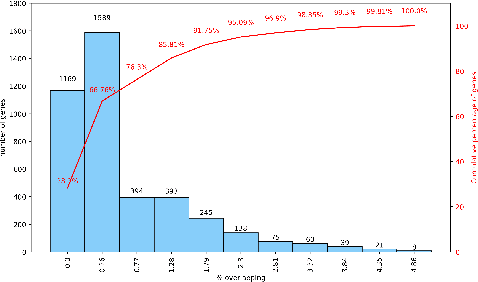
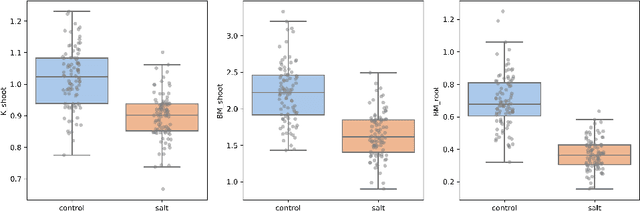
This paper proposes a workflow to identify genes responding to a specific treatment in an organism, such as abiotic stresses, a main cause of extensive agricultural production losses worldwide. On input RNA sequencing read counts (measured for genotypes under control and treatment conditions) and biological replicates, it outputs a collection of characterized genes, potentially relevant to treatment. Technically, the proposed approach is both a generalization and an extension of WGCNA; its main goal is to identify specific modules in a network of genes after a sequence of normalization and filtering steps. In this work, module detection is achieved by using Hierarchical Link Clustering, which can recognize overlapping communities and thus have more biological meaning given the overlapping regulatory domains of systems that generate co-expression. Additional steps and information are also added to the workflow, where some networks in the intermediate steps are forced to be scale-free and LASSO regression is employed to select the most significant modules of phenotypical responses to stress. Finally, the workflow is showcased with a systematic study on rice (Oryza sativa), a major food source that is known to be highly sensitive to salt stress: a total of 6 modules are detected as relevant in the response to salt stress in rice; these genes may act as potential targets for the improvement of salinity tolerance in rice cultivars. The proposed workflow has the potential to ultimately reduce the search-space for candidate genes responding to a specific treatment, which can considerably optimize the effort, time, and money invested by researchers in the experimental validation of stress responsive genes.
Deep Volumetric Universal Lesion Detection using Light-Weight Pseudo 3D Convolution and Surface Point Regression
Aug 30, 2020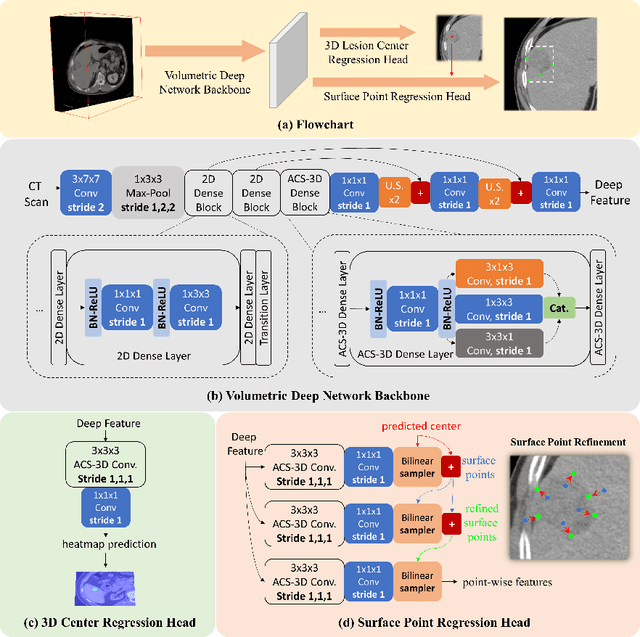
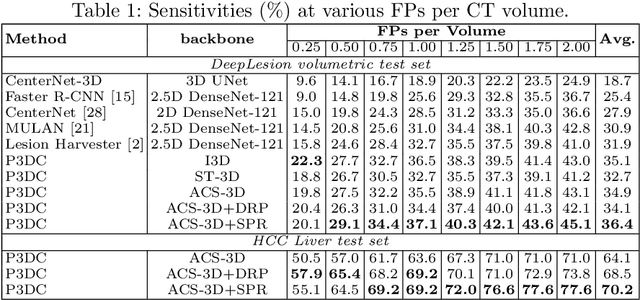
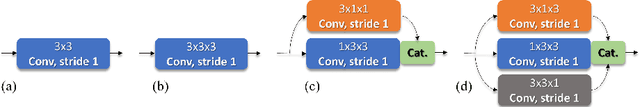
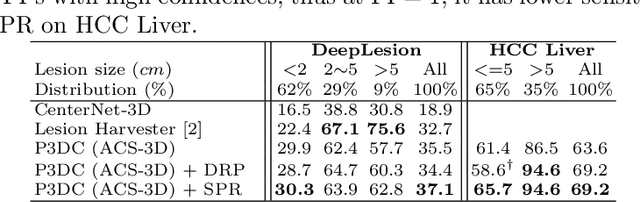
Identifying, measuring and reporting lesions accurately and comprehensively from patient CT scans are important yet time-consuming procedures for physicians. Computer-aided lesion/significant-findings detection techniques are at the core of medical imaging, which remain very challenging due to the tremendously large variability of lesion appearance, location and size distributions in 3D imaging. In this work, we propose a novel deep anchor-free one-stage VULD framework that incorporates (1) P3DC operators to recycle the architectural configurations and pre-trained weights from the off-the-shelf 2D networks, especially ones with large capacities to cope with data variance, and (2) a new SPR method to effectively regress the 3D lesion spatial extents by pinpointing their representative key points on lesion surfaces. Experimental validations are first conducted on the public large-scale NIH DeepLesion dataset where our proposed method delivers new state-of-the-art quantitative performance. We also test VULD on our in-house dataset for liver tumor detection. VULD generalizes well in both large-scale and small-sized tumor datasets in CT imaging.
Emulator-based global sensitivity analysis for flow-like landslide run-out models
Oct 08, 2020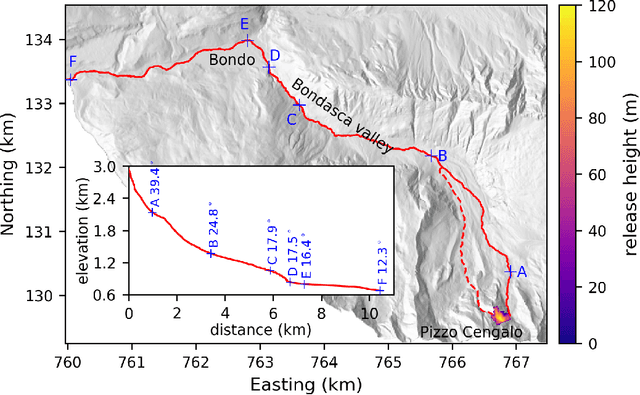
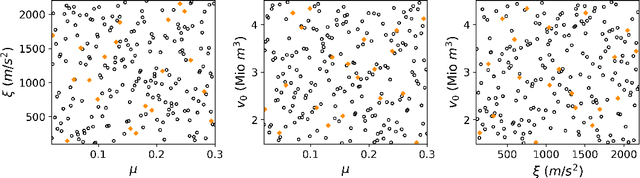
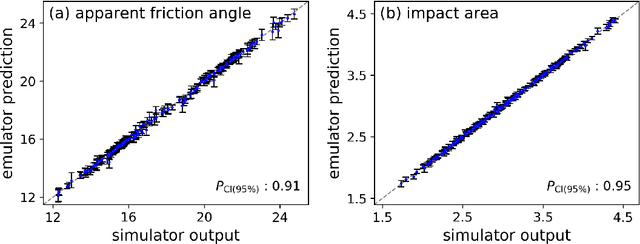
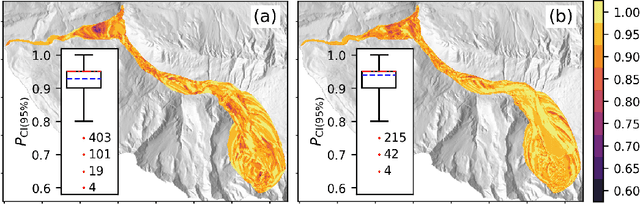
Landslide run-out modeling involves various uncertainties originating from model input data. It is therefore desirable to assess the model's sensitivity. A global sensitivity analysis that is capable of exploring the entire input space and accounts for all interactions, often remains limited due to computational challenges resulting from a large number of necessary model runs. We address this research gap by integrating Gaussian process emulation into landslide run-out modeling and apply it to the open-source simulation tool r.avaflow. The feasibility and efficiency of our approach is illustrated based on the 2017 Bondo landslide event. The sensitivity of aggregated model outputs, such as the apparent friction angle, impact area, as well as spatially resolved maximum flow height and velocity, to the dry-Coulomb friction coefficient, turbulent friction coefficient and the release volume are studied. The results of first-order effects are consistent with previous results of common one-at-a-time sensitivity analyses. In addition to that, our approach allows to rigorously investigate interactions. Strong interactions are detected on the margins of the flow path where the expectation and variation of maximum flow height and velocity are small. The interactions generally become weak with increasing variation of maximum flow height and velocity. Besides, there are stronger interactions between the two friction coefficients than between the release volume and each friction coefficient. In the future, it is promising to extend the approach for other computationally expensive tasks like uncertainty quantification, model calibration, and smart early warning.
Exemplary Natural Images Explain CNN Activations Better than Feature Visualizations
Oct 23, 2020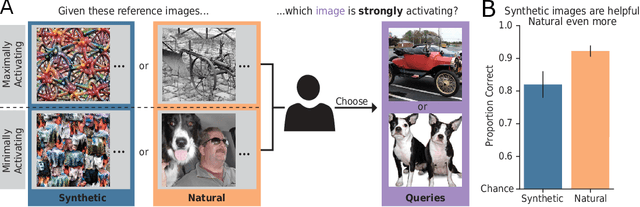
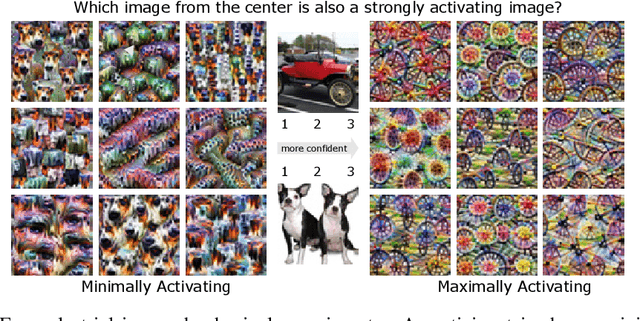
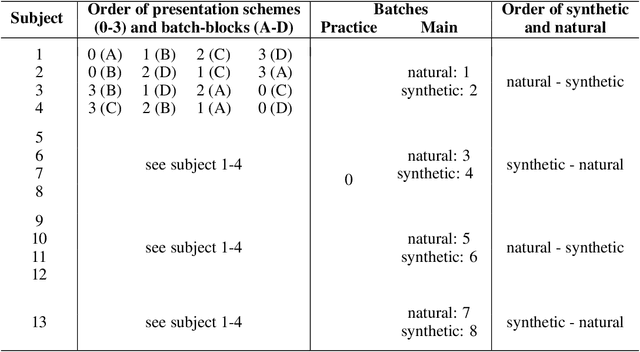

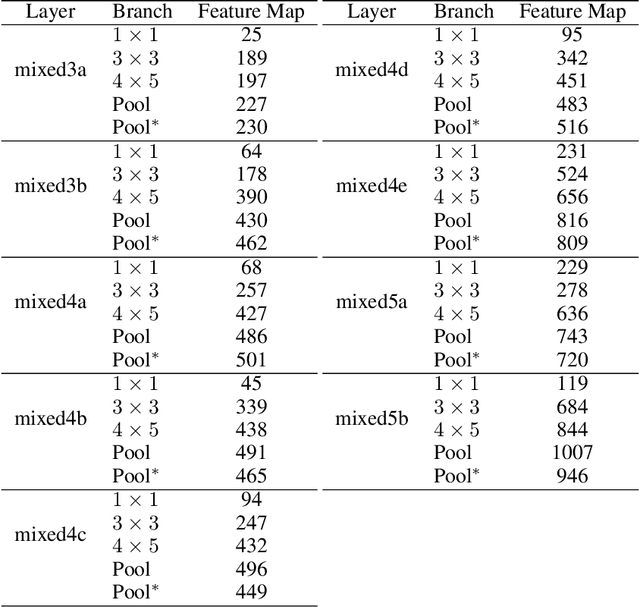
Feature visualizations such as synthetic maximally activating images are a widely used explanation method to better understand the information processing of convolutional neural networks (CNNs). At the same time, there are concerns that these visualizations might not accurately represent CNNs' inner workings. Here, we measure how much extremely activating images help humans to predict CNN activations. Using a well-controlled psychophysical paradigm, we compare the informativeness of synthetic images (Olah et al., 2017) with a simple baseline visualization, namely exemplary natural images that also strongly activate a specific feature map. Given either synthetic or natural reference images, human participants choose which of two query images leads to strong positive activation. The experiment is designed to maximize participants' performance, and is the first to probe intermediate instead of final layer representations. We find that synthetic images indeed provide helpful information about feature map activations (82% accuracy; chance would be 50%). However, natural images-originally intended to be a baseline-outperform synthetic images by a wide margin (92% accuracy). Additionally, participants are faster and more confident for natural images, whereas subjective impressions about the interpretability of feature visualization are mixed. The higher informativeness of natural images holds across most layers, for both expert and lay participants as well as for hand- and randomly-picked feature visualizations. Even if only a single reference image is given, synthetic images provide less information than natural images (65% vs. 73%). In summary, popular synthetic images from feature visualizations are significantly less informative for assessing CNN activations than natural images. We argue that future visualization methods should improve over this simple baseline.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge