"Time": models, code, and papers
Multi-Slice Fusion for Sparse-View and Limited-Angle 4D CT Reconstruction
Aug 01, 2020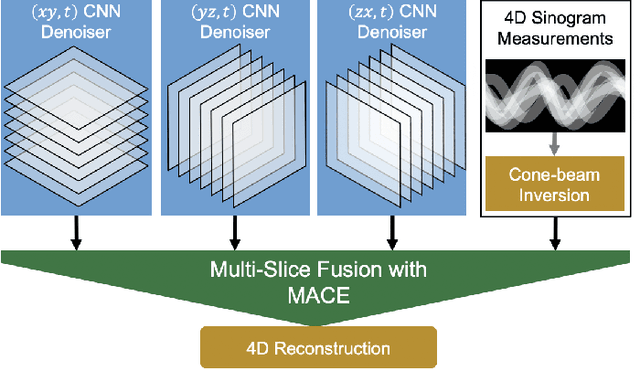
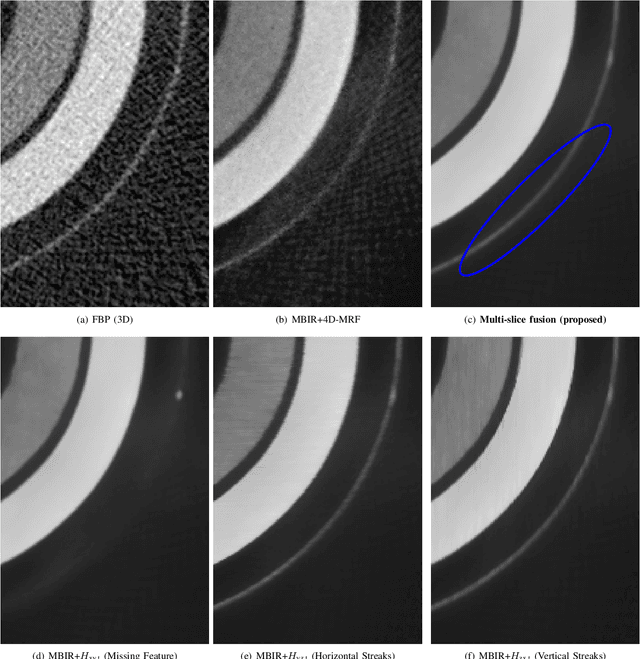

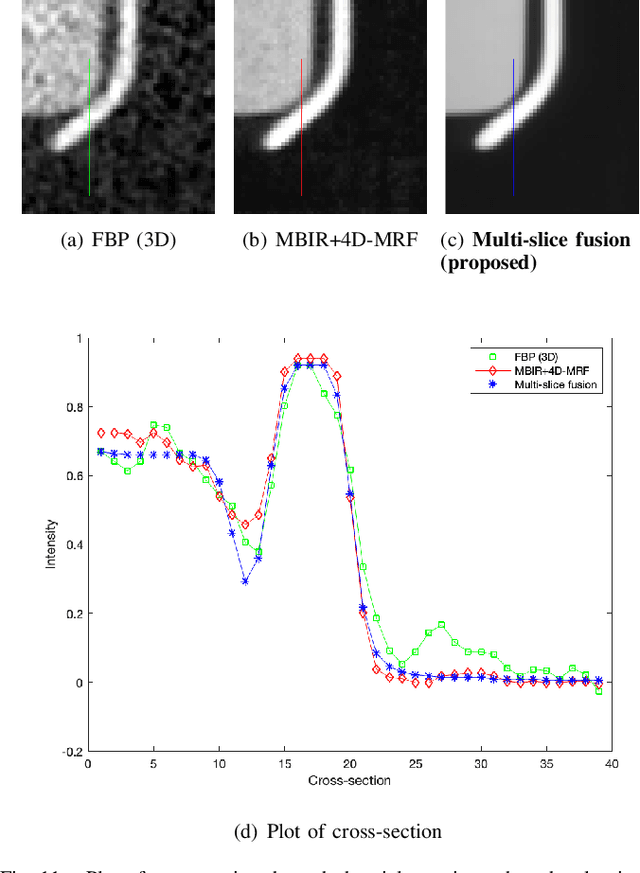
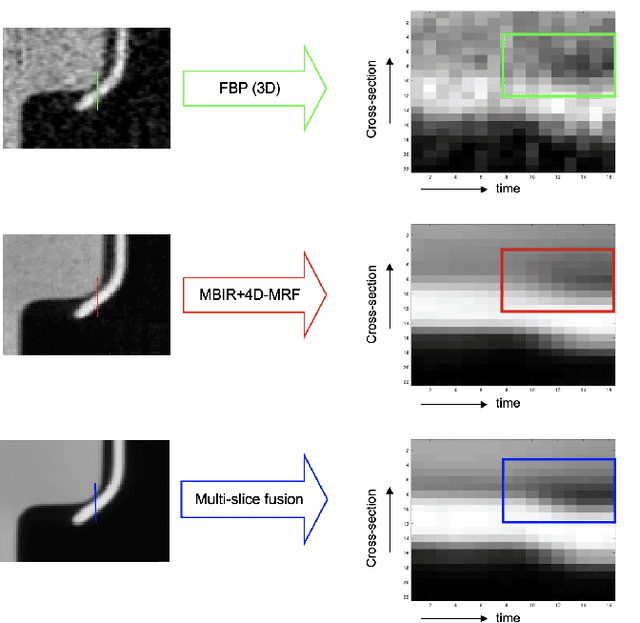
Inverse problems spanning four or more dimensions such as space, time and other independent parameters have become increasingly important. State-of-the-art 4D reconstruction methods use model based iterative reconstruction (MBIR), but depend critically on the quality of the prior modeling. Recently, plug-and-play (PnP) methods have been shown to be an effective way to incorporate advanced prior models using state-of-the-art denoising algorithms. However, state-of-the-art denoisers such as BM4D and deep convolutional neural networks (CNNs) are primarily available for 2D or 3D images and extending them to higher dimensions is difficult due to algorithmic complexity and the increased difficulty of effective training. In this paper, we present multi-slice fusion, a novel algorithm for 4D reconstruction, based on the fusion of multiple low-dimensional denoisers. Our approach uses multi-agent consensus equilibrium (MACE), an extension of plug-and-play, as a framework for integrating the multiple lower-dimensional models. We apply our method to 4D cone-beam X-ray CT reconstruction for non destructive evaluation (NDE) of samples that are dynamically moving during acquisition. We implement multi-slice fusion on distributed, heterogeneous clusters in order to reconstruct large 4D volumes in reasonable time and demonstrate the inherent parallelizable nature of the algorithm. We present simulated and real experimental results on sparse-view and limited-angle CT data to demonstrate that multi-slice fusion can substantially improve the quality of reconstructions relative to traditional methods, while also being practical to implement and train.
Compliant Conditions for Polynomial Time Approximation of Operator Counts
Jul 05, 2016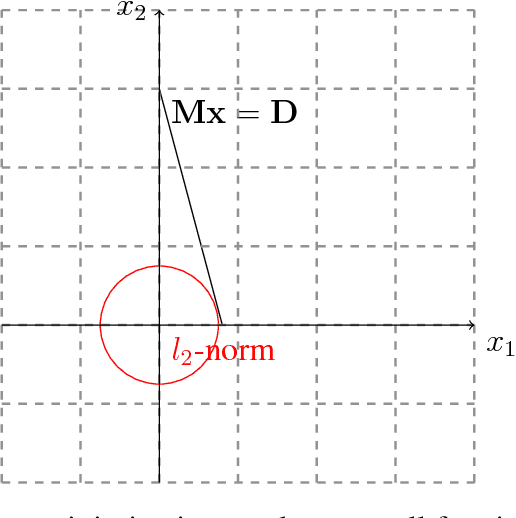
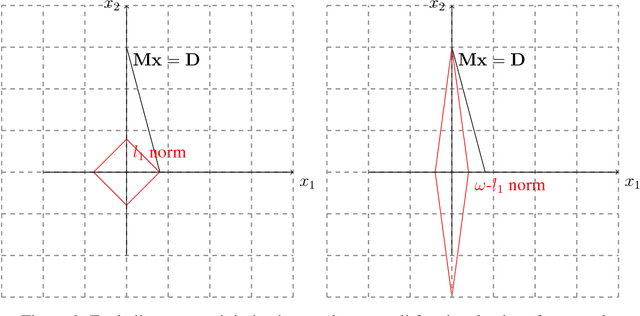
In this paper, we develop a computationally simpler version of the operator count heuristic for a particular class of domains. The contribution of this abstract is threefold, we (1) propose an efficient closed form approximation to the operator count heuristic using the Lagrangian dual; (2) leverage compressed sensing techniques to obtain an integer approximation for operator counts in polynomial time; and (3) discuss the relationship of the proposed formulation to existing heuristics and investigate properties of domains where such approaches appear to be useful.
Learning to Translate in Real-time with Neural Machine Translation
Jan 10, 2017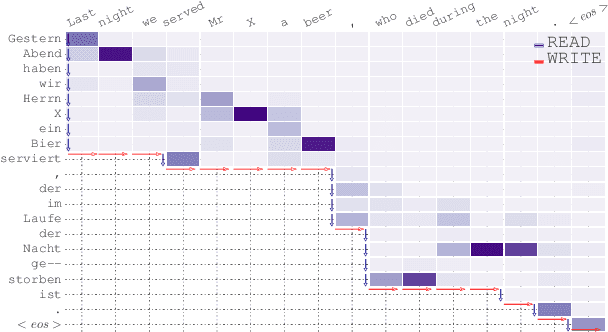
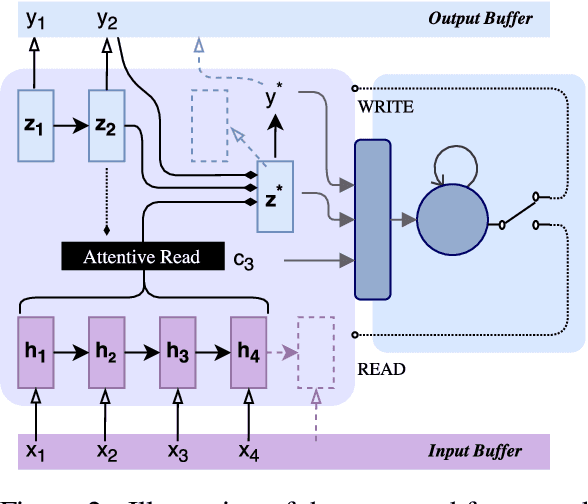
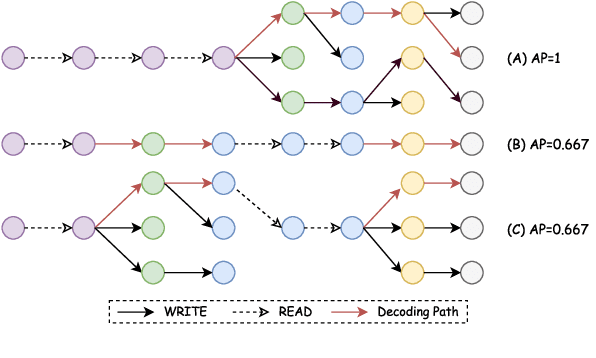
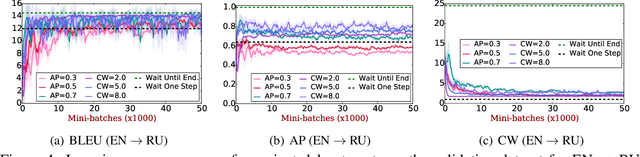
Translating in real-time, a.k.a. simultaneous translation, outputs translation words before the input sentence ends, which is a challenging problem for conventional machine translation methods. We propose a neural machine translation (NMT) framework for simultaneous translation in which an agent learns to make decisions on when to translate from the interaction with a pre-trained NMT environment. To trade off quality and delay, we extensively explore various targets for delay and design a method for beam-search applicable in the simultaneous MT setting. Experiments against state-of-the-art baselines on two language pairs demonstrate the efficacy of the proposed framework both quantitatively and qualitatively.
Time-projection control to recover inter-sample disturbances, application to bipedal walking control
Jan 07, 2018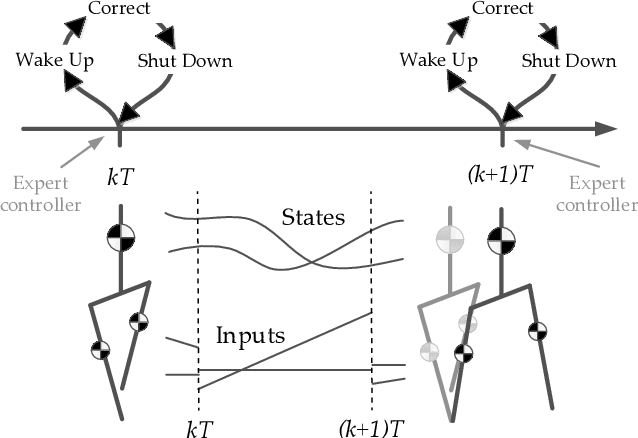
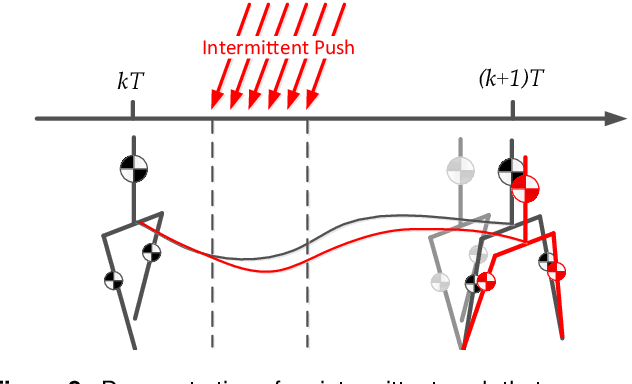
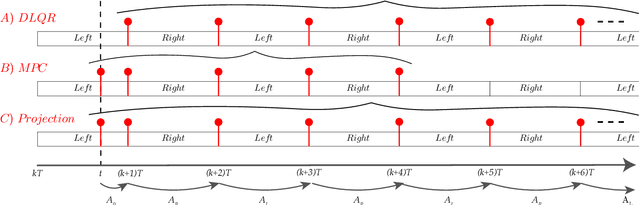
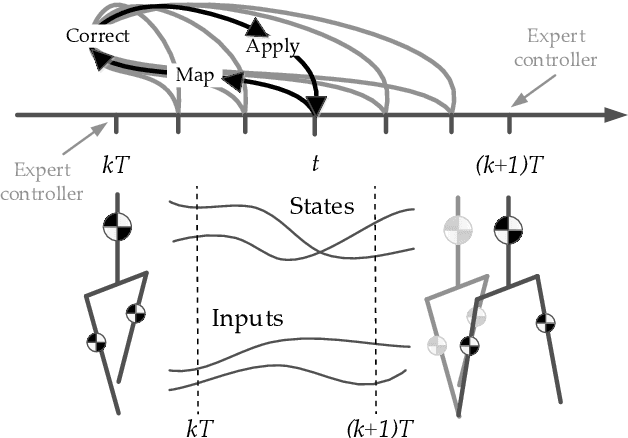
We present a new walking controller based on 3LP, a 3D model of bipedal walking that is composed of three pendulums to simulate falling, swing and torso dynamics. Taking advantage of linear equations and closed-form solutions of 3LP, the proposed controller projects intermediate states of the biped back to the beginning of the phase for which a discrete LQR controller is designed. After the projection, a proper control policy is generated by this LQR controller and used at the intermediate time. The projection controller reacts to disturbances immediately and compared to the discrete LQR controller, it provides superior performance in recovering intermittent external pushes. Further analysis of closed-loop eigenvalues and disturbance rejection strength show strong stabilization properties for this architecture. An analysis of viable regions also show that the proposed controller covers most of the maximal viable set of states. It is computationally much faster than Model Predictive Controllers (MPC) and yet optimal over an infinite horizon.
Nonseparable Symplectic Neural Networks
Oct 23, 2020
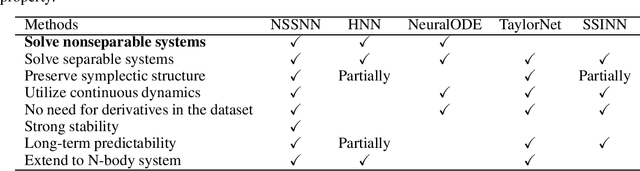
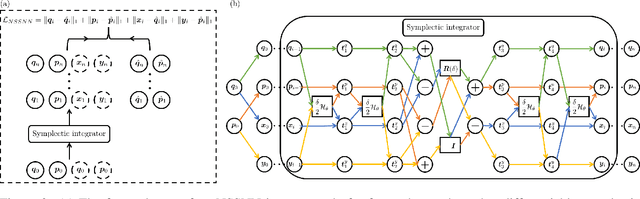
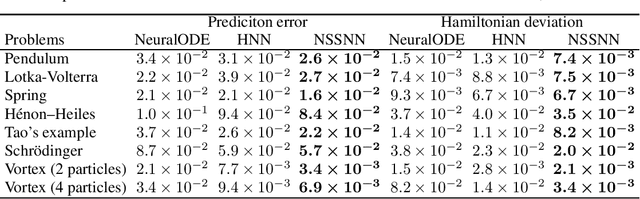
Predicting the behaviors of Hamiltonian systems has been drawing increasing attention in scientific machine learning. However, the vast majority of the literature was focused on predicting separable Hamiltonian systems with their kinematic and potential energy terms being explicitly decoupled, while building data-driven paradigms to predict nonseparable Hamiltonian systems that are ubiquitous in fluid dynamics and quantum mechanics were rarely explored. The main computational challenge lies in the effective embedding of symplectic priors to describe the inherently coupled evolution of position and momentum, which typically exhibits intricate dynamics with many degrees of freedom. To solve the problem, we propose a novel neural network architecture, Nonseparable Symplectic Neural Networks (NSSNNs), to uncover and embed the symplectic structure of a nonseparable Hamiltonian system from limited observation data. The enabling mechanics of our approach is an augmented symplectic time integrator to decouple the position and momentum energy terms and facilitate their evolution. We demonstrated the efficacy and versatility of our method by predicting a wide range of Hamiltonian systems, both separable and nonseparable, including vortical flow and quantum system. We showed the unique computational merits of our approach to yield long-term, accurate, and robust predictions for large-scale Hamiltonian systems by rigorously enforcing symplectomorphism.
Generate Novel Molecules With Target Properties Using Conditional Generative Models
Sep 15, 2020
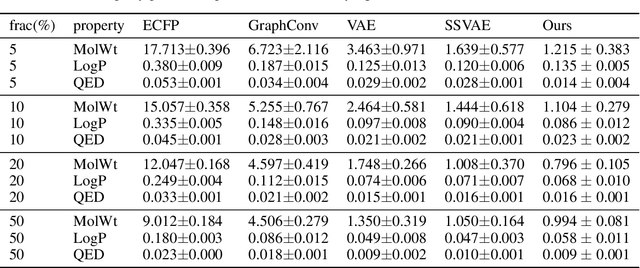
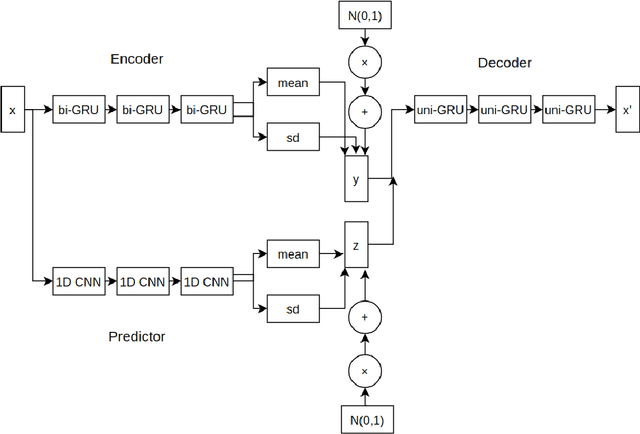
Drug discovery using deep learning has attracted a lot of attention of late as it has obvious advantages like higher efficiency, less manual guessing and faster process time. In this paper, we present a novel neural network for generating small molecules similar to the ones in the training set. Our network consists of an encoder made up of bi-GRU layers for converting the input samples to a latent space, predictor for enhancing the capability of encoder made up of 1D-CNN layers and a decoder comprised of uni-GRU layers for reconstructing the samples from the latent space representation. Condition vector in latent space is used for generating molecules with the desired properties. We present the loss functions used for training our network, experimental details and property prediction metrics. Our network outperforms previous methods using Molecular weight, LogP and Quantitative Estimation of Drug-likeness as the evaluation metrics.
Time-Varying Formation Controllers for Unmanned Aerial Vehicles Using Deep Reinforcement Learning
Jun 05, 2017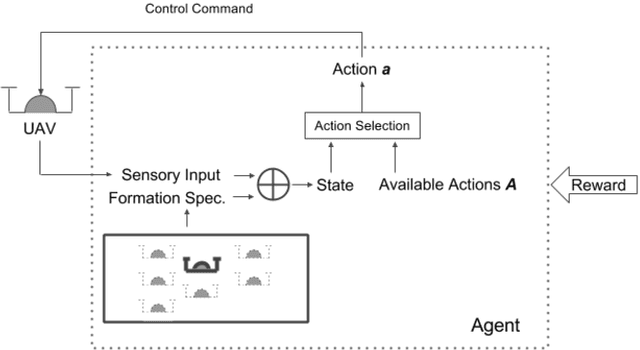
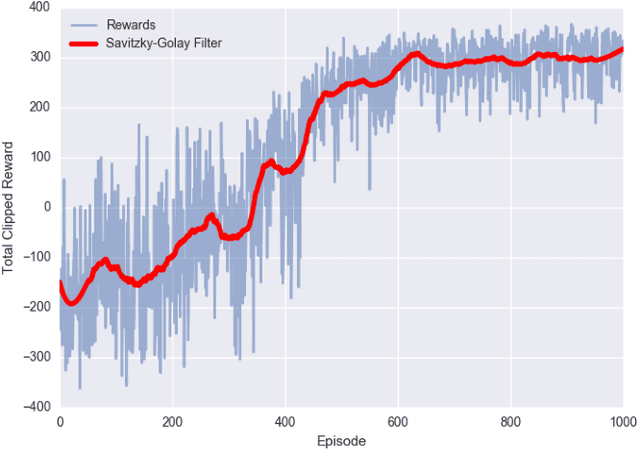
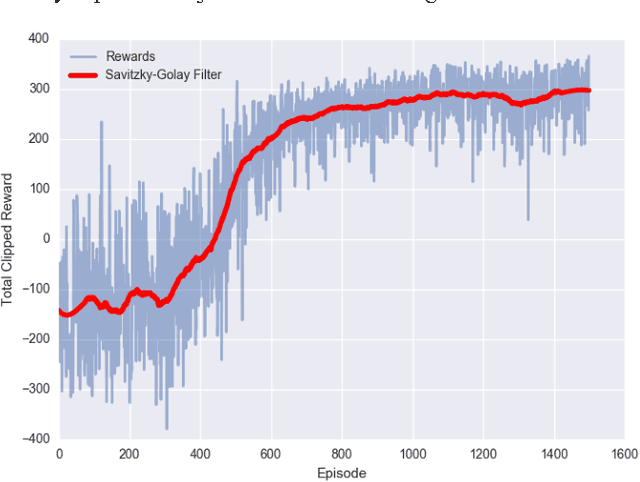
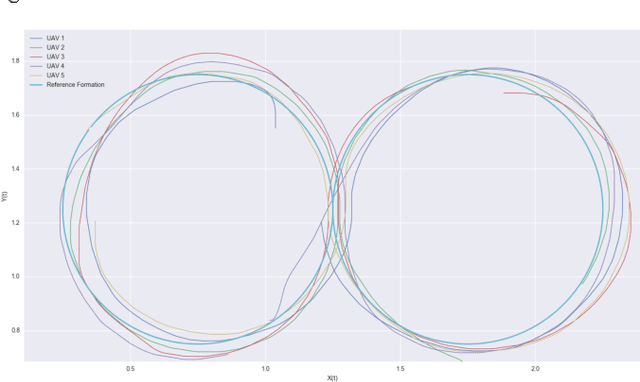
We consider the problem of designing scalable and portable controllers for unmanned aerial vehicles (UAVs) to reach time-varying formations as quickly as possible. This brief confirms that deep reinforcement learning can be used in a multi-agent fashion to drive UAVs to reach any formation while taking into account optimality and portability. We use a deep neural network to estimate how good a state is, so the agent can choose actions accordingly. The system is tested with different non-high-dimensional sensory inputs without any change in the neural network architecture, algorithm or hyperparameters, just with additional training.
Identification of state functions by physically-guided neural networks with physically-meaningful internal layers
Nov 17, 2020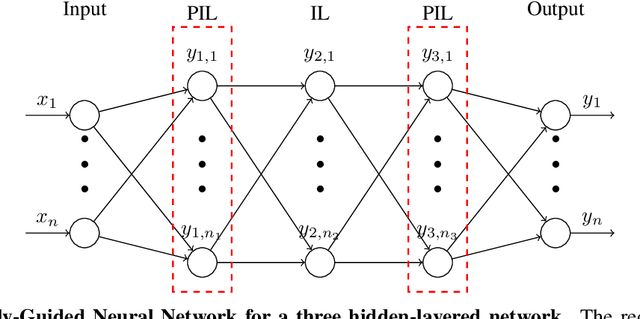

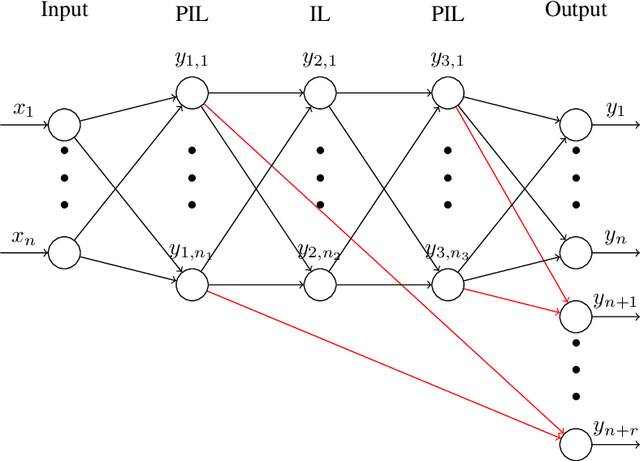

Substitution of well-grounded theoretical models by data-driven predictions is not as simple in engineering and sciences as it is in social and economic fields. Scientific problems suffer most times from paucity of data, while they may involve a large number of variables and parameters that interact in complex and non-stationary ways, obeying certain physical laws. Moreover, a physically-based model is not only useful for making predictions, but to gain knowledge by the interpretation of its structure, parameters, and mathematical properties. The solution to these shortcomings seems to be the seamless blending of the tremendous predictive power of the data-driven approach with the scientific consistency and interpretability of physically-based models. We use here the concept of physically-constrained neural networks (PCNN) to predict the input-output relation in a physical system, while, at the same time fulfilling the physical constraints. With this goal, the internal hidden state variables of the system are associated with a set of internal neuron layers, whose values are constrained by known physical relations, as well as any additional knowledge on the system. Furthermore, when having enough data, it is possible to infer knowledge about the internal structure of the system and, if parameterized, to predict the state parameters for a particular input-output relation. We show that this approach, besides getting physically-based predictions, accelerates the training process, reduces the amount of data required to get similar accuracy, filters partly the intrinsic noise in the experimental data and provides improved extrapolation capacity.
Non-Sparse PCA in High Dimensions via Cone Projected Power Iteration
May 15, 2020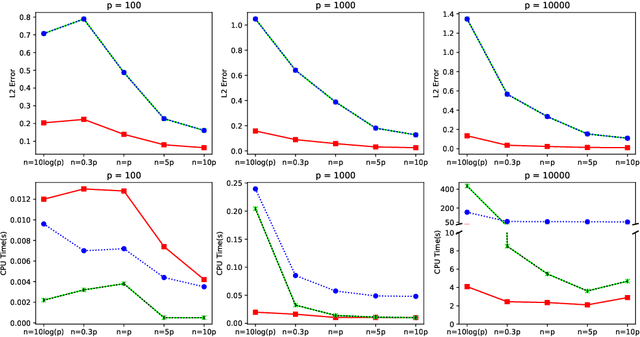
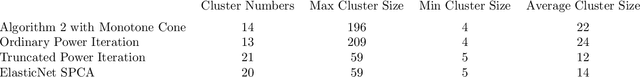
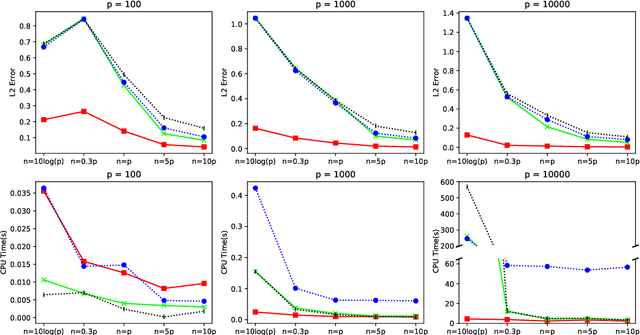
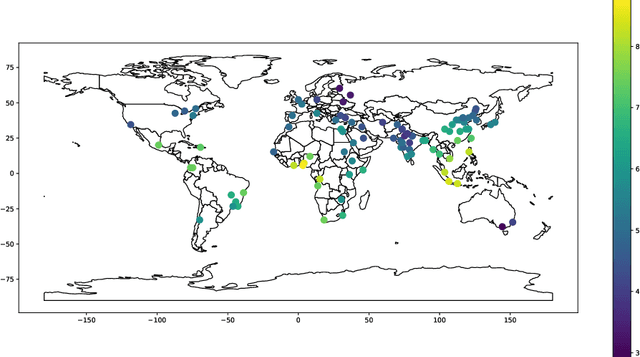
In this paper, we propose a cone projected power iteration algorithm to recover the principal eigenvector from a noisy positive semidefinite matrix. When the true principal eigenvector is assumed to belong to a convex cone, the proposed algorithm is fast and has a tractable error. Specifically, the method achieves polynomial time complexity for certain convex cones equipped with fast projection such as the monotone cone. It attains a small error when the noisy matrix has a small cone-restricted operator norm. We supplement the above results with a minimax lower bound of the error under the spiked covariance model. Our numerical experiments on simulated and real data, show that our method achieves shorter run time and smaller error in comparison to the ordinary power iteration and some sparse principal component analysis algorithms if the principal eigenvector is in a convex cone.
Fast and Accurate Forecasting of COVID-19 Deaths Using the SIkJ$α$ Model
Jul 13, 2020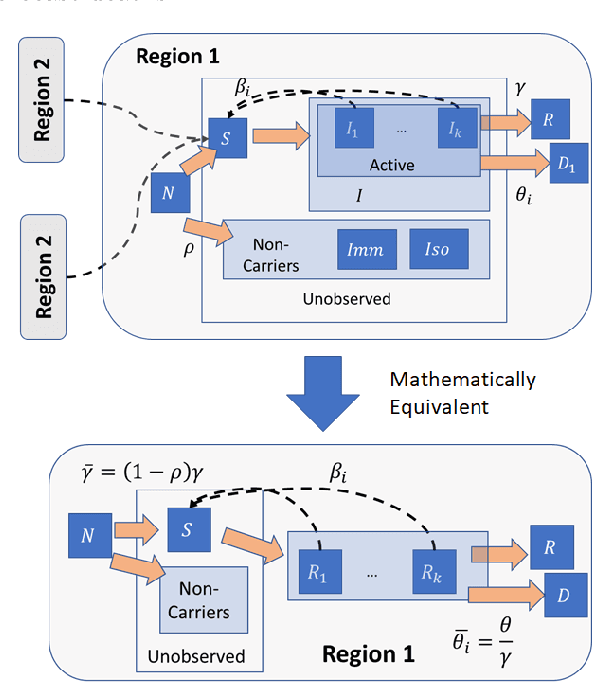
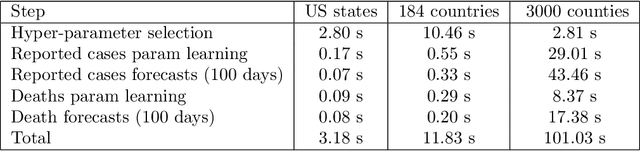
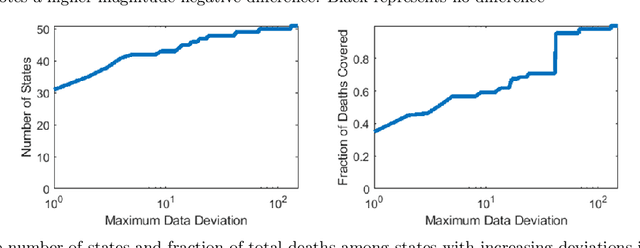
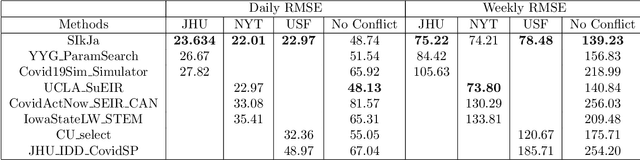
Forecasting the effect of COVID-19 is essential to design policies that may prepare us to handle the pandemic. Many methods have already been proposed, particularly, to forecast reported cases and deaths at country-level and state-level. Many of these methods are based on traditional epidemiological model which rely on simulations or Bayesian inference to simultaneously learn many parameters at a time. This makes them prone to over-fitting and slow execution. We propose an extension to our model SIkJ$\alpha$ to forecast deaths and show that it can consider the effect of many complexities of the epidemic process and yet be simplified to a few parameters that are learned using fast linear regressions. We also present an evaluation of our method against seven approaches currently being used by the CDC, based on their two weeks forecast at various times during the pandemic. We demonstrate that our method achieves better root mean squared error compared to these seven approaches during majority of the evaluation period. Further, on a 2 core desktop machine, our approach takes only 3.18s to tune hyper-parameters, learn parameters and generate 100 days of forecasts of reported cases and deaths for all the states in the US. The total execution time for 184 countries is 11.83s and for all the US counties ($>$ 3000) is 101.03s.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge