"Time": models, code, and papers
Stability and Generalization of the Decentralized Stochastic Gradient Descent
Feb 02, 2021
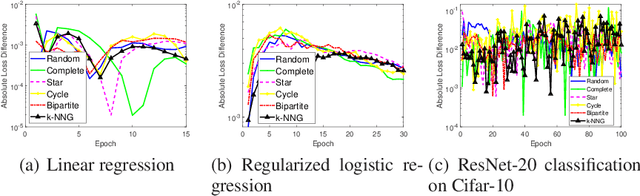
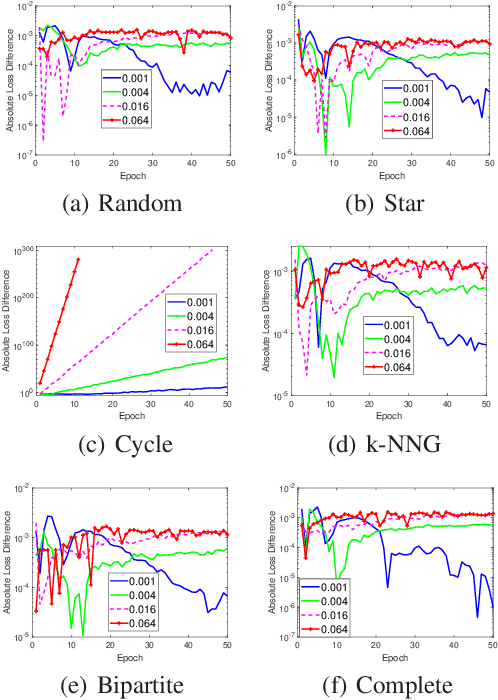
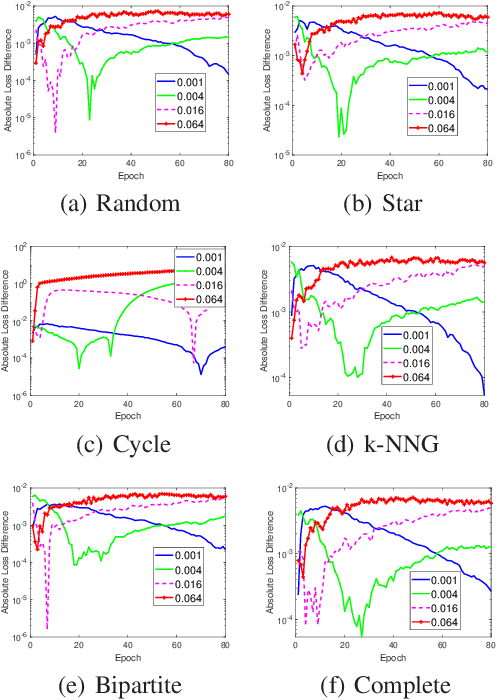
The stability and generalization of stochastic gradient-based methods provide valuable insights into understanding the algorithmic performance of machine learning models. As the main workhorse for deep learning, stochastic gradient descent has received a considerable amount of studies. Nevertheless, the community paid little attention to its decentralized variants. In this paper, we provide a novel formulation of the decentralized stochastic gradient descent. Leveraging this formulation together with (non)convex optimization theory, we establish the first stability and generalization guarantees for the decentralized stochastic gradient descent. Our theoretical results are built on top of a few common and mild assumptions and reveal that the decentralization deteriorates the stability of SGD for the first time. We verify our theoretical findings by using a variety of decentralized settings and benchmark machine learning models.
Decomposing Normal and Abnormal Features of Medical Images into Discrete Latent Codes for Content-Based Image Retrieval
Mar 23, 2021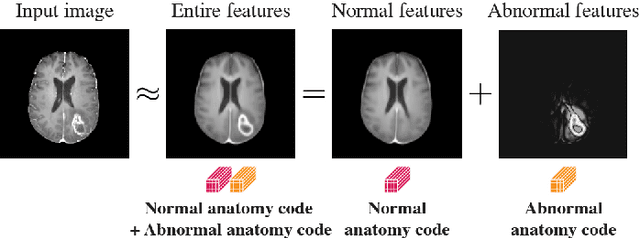
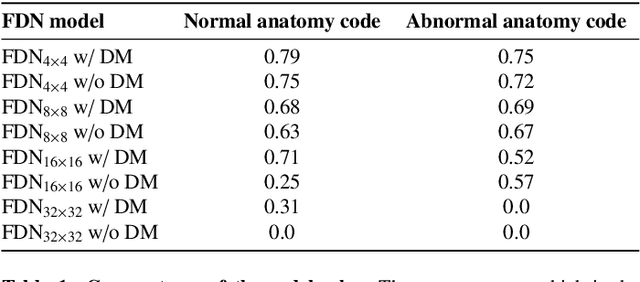
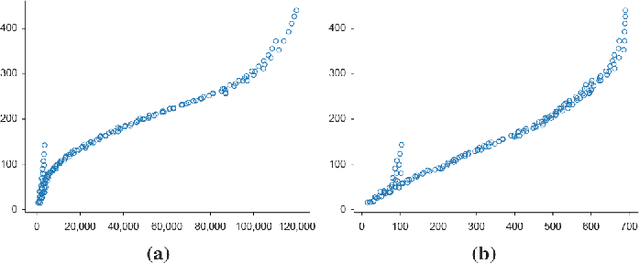
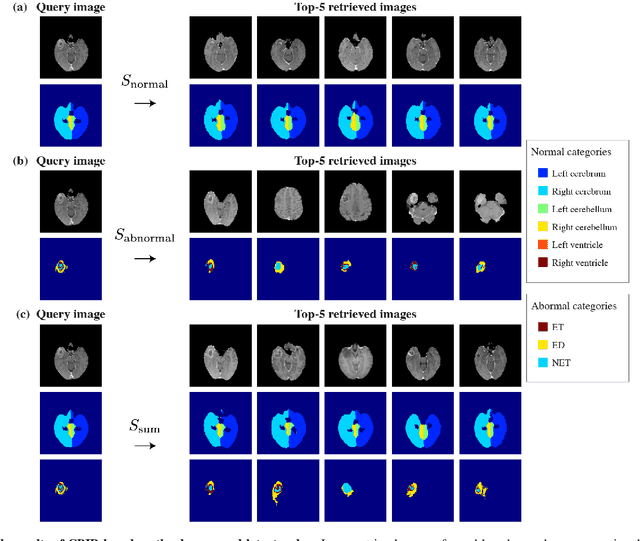
In medical imaging, the characteristics purely derived from a disease should reflect the extent to which abnormal findings deviate from the normal features. Indeed, physicians often need corresponding images without abnormal findings of interest or, conversely, images that contain similar abnormal findings regardless of normal anatomical context. This is called comparative diagnostic reading of medical images, which is essential for a correct diagnosis. To support comparative diagnostic reading, content-based image retrieval (CBIR), which can selectively utilize normal and abnormal features in medical images as two separable semantic components, will be useful. Therefore, we propose a neural network architecture to decompose the semantic components of medical images into two latent codes: normal anatomy code and abnormal anatomy code. The normal anatomy code represents normal anatomies that should have existed if the sample is healthy, whereas the abnormal anatomy code attributes to abnormal changes that reflect deviation from the normal baseline. These latent codes are discretized through vector quantization to enable binary hashing, which can reduce the computational burden at the time of similarity search. By calculating the similarity based on either normal or abnormal anatomy codes or the combination of the two codes, our algorithm can retrieve images according to the selected semantic component from a dataset consisting of brain magnetic resonance images of gliomas. Our CBIR system qualitatively and quantitatively achieves remarkable results.
On the Temporality of Priors in Entity Linking
Jan 14, 2021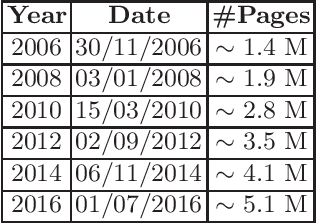
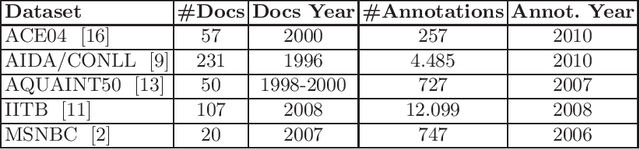
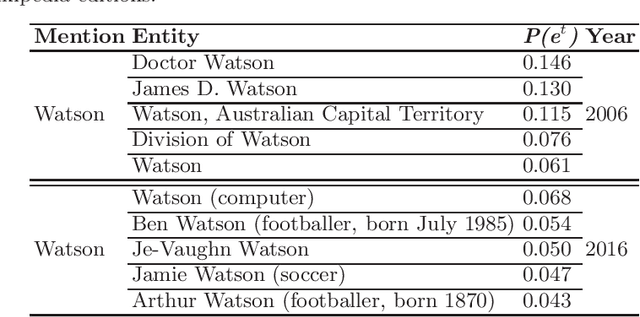
Entity linking is a fundamental task in natural language processing which deals with the lexical ambiguity in texts. An important component in entity linking approaches is the mention-to-entity prior probability. Even though there is a large number of works in entity linking, the existing approaches do not explicitly consider the time aspect, specifically the temporality of an entity's prior probability. We posit that this prior probability is temporal in nature and affects the performance of entity linking systems. In this paper we systematically study the effect of the prior on the entity linking performance over the temporal validity of both texts and KBs.
Parameter-free Locally Accelerated Conditional Gradients
Feb 12, 2021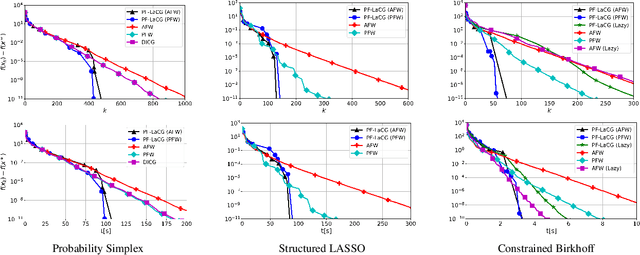
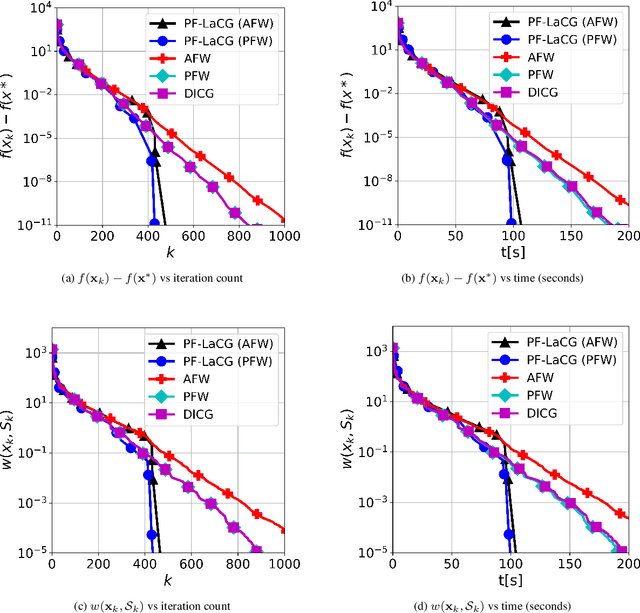
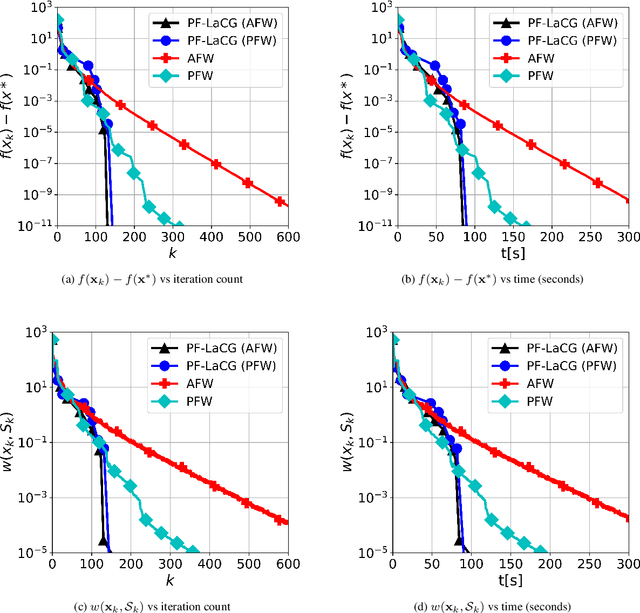
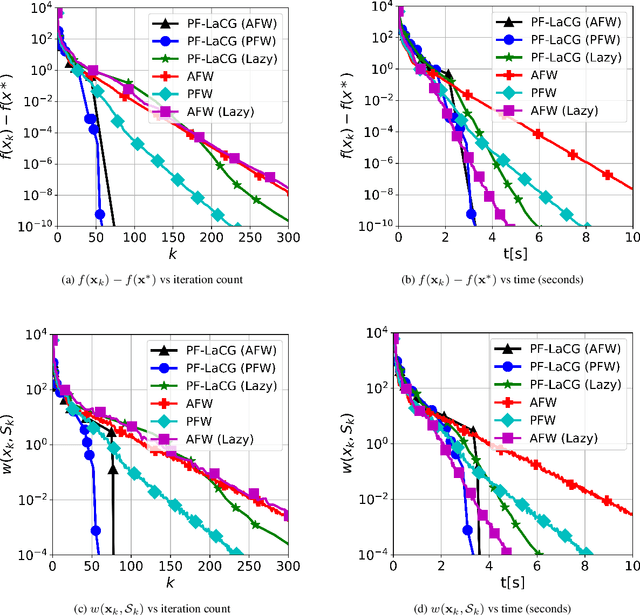
Projection-free conditional gradient (CG) methods are the algorithms of choice for constrained optimization setups in which projections are often computationally prohibitive but linear optimization over the constraint set remains computationally feasible. Unlike in projection-based methods, globally accelerated convergence rates are in general unattainable for CG. However, a very recent work on Locally accelerated CG (LaCG) has demonstrated that local acceleration for CG is possible for many settings of interest. The main downside of LaCG is that it requires knowledge of the smoothness and strong convexity parameters of the objective function. We remove this limitation by introducing a novel, Parameter-Free Locally accelerated CG (PF-LaCG) algorithm, for which we provide rigorous convergence guarantees. Our theoretical results are complemented by numerical experiments, which demonstrate local acceleration and showcase the practical improvements of PF-LaCG over non-accelerated algorithms, both in terms of iteration count and wall-clock time.
Adaptive-Control-Oriented Meta-Learning for Nonlinear Systems
Mar 07, 2021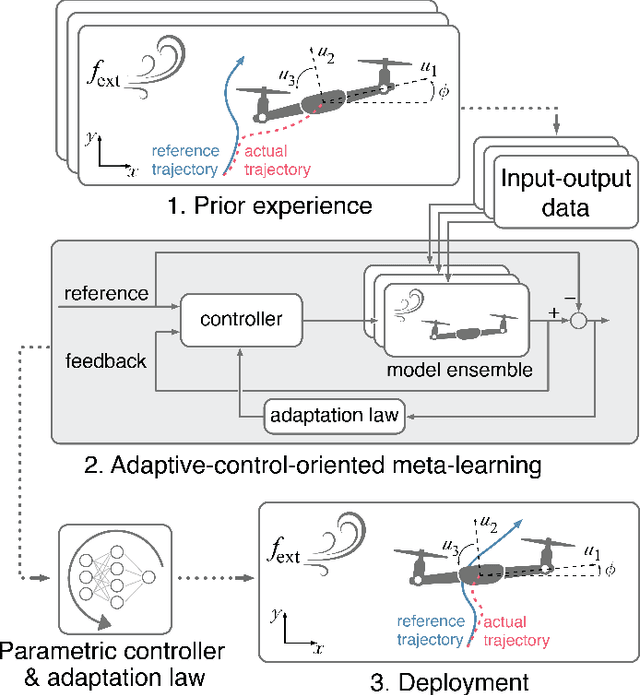
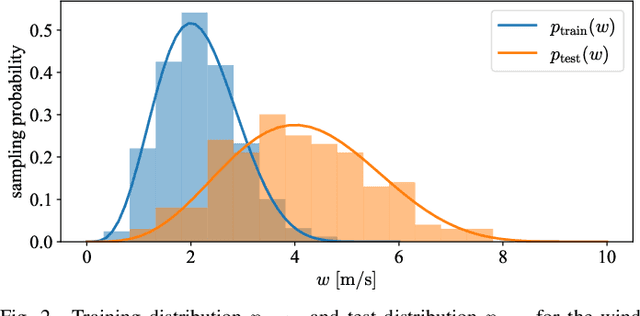
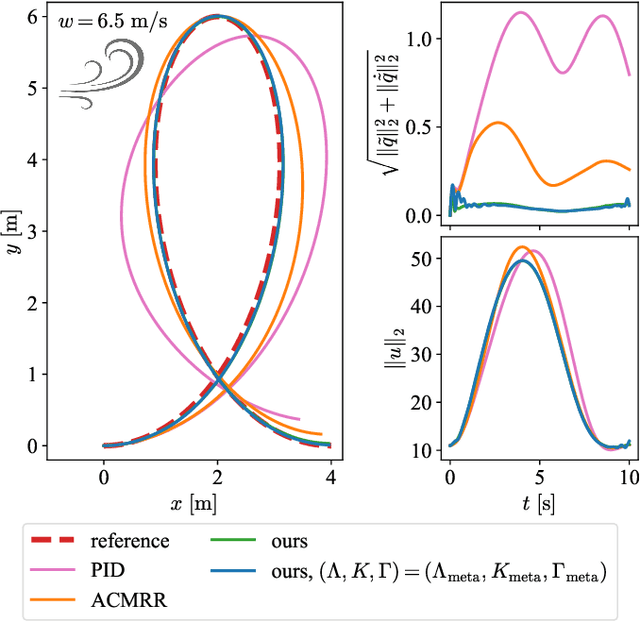
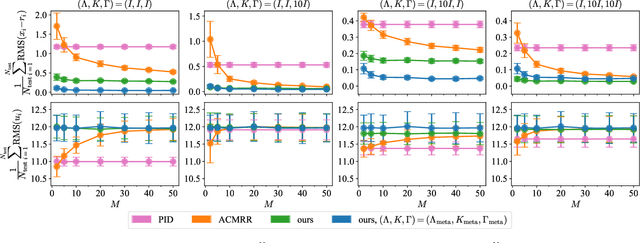
Real-time adaptation is imperative to the control of robots operating in complex, dynamic environments. Adaptive control laws can endow even nonlinear systems with good trajectory tracking performance, provided that any uncertain dynamics terms are linearly parameterizable with known nonlinear features. However, it is often difficult to specify such features a priori, such as for aerodynamic disturbances on rotorcraft or interaction forces between a manipulator arm and various objects. In this paper, we turn to data-driven modeling with neural networks to learn, offline from past data, an adaptive controller with an internal parametric model of these nonlinear features. Our key insight is that we can better prepare the controller for deployment with control-oriented meta-learning of features in closed-loop simulation, rather than regression-oriented meta-learning of features to fit input-output data. Specifically, we meta-learn the adaptive controller with closed-loop tracking simulation as the base-learner and the average tracking error as the meta-objective. With a nonlinear planar rotorcraft subject to wind, we demonstrate that our adaptive controller outperforms other controllers trained with regression-oriented meta-learning when deployed in closed-loop for trajectory tracking control.
Recurrent Neural Networks for Multivariate Time Series with Missing Values
Nov 07, 2016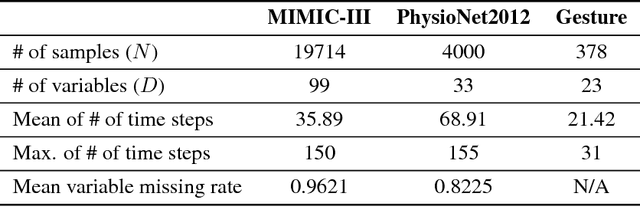
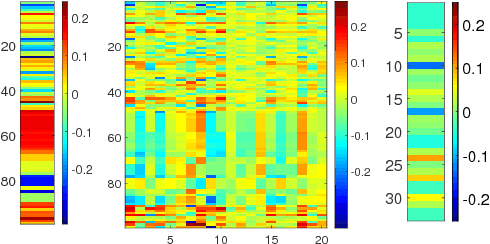
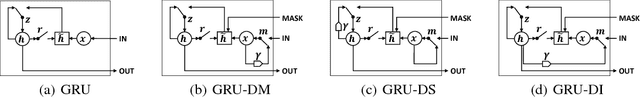
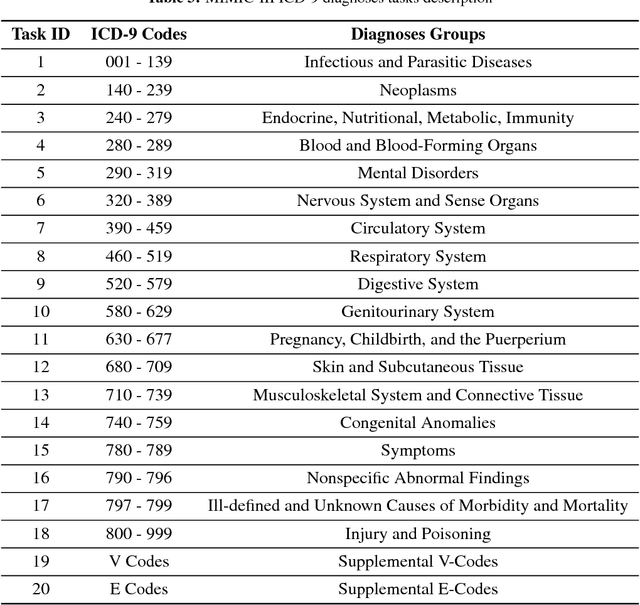
Multivariate time series data in practical applications, such as health care, geoscience, and biology, are characterized by a variety of missing values. In time series prediction and other related tasks, it has been noted that missing values and their missing patterns are often correlated with the target labels, a.k.a., informative missingness. There is very limited work on exploiting the missing patterns for effective imputation and improving prediction performance. In this paper, we develop novel deep learning models, namely GRU-D, as one of the early attempts. GRU-D is based on Gated Recurrent Unit (GRU), a state-of-the-art recurrent neural network. It takes two representations of missing patterns, i.e., masking and time interval, and effectively incorporates them into a deep model architecture so that it not only captures the long-term temporal dependencies in time series, but also utilizes the missing patterns to achieve better prediction results. Experiments of time series classification tasks on real-world clinical datasets (MIMIC-III, PhysioNet) and synthetic datasets demonstrate that our models achieve state-of-the-art performance and provides useful insights for better understanding and utilization of missing values in time series analysis.
Evaluating phase synchronization methods in fMRI: a comparison study and new approaches
Sep 21, 2020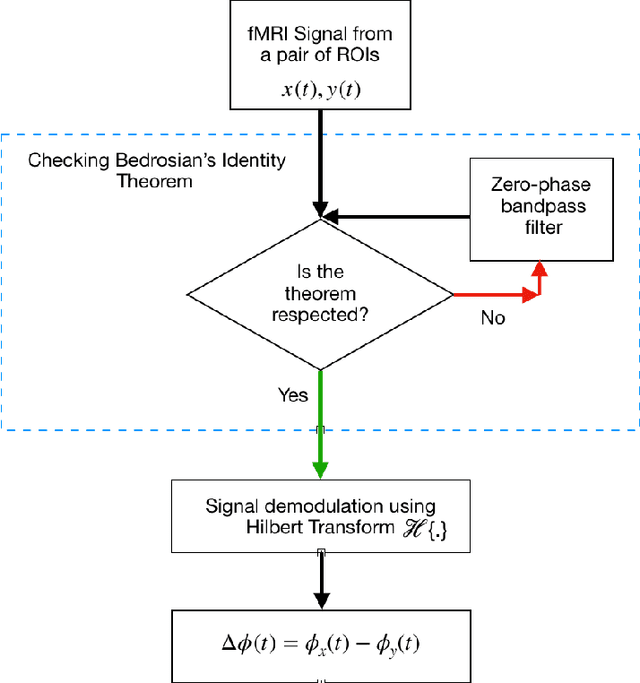
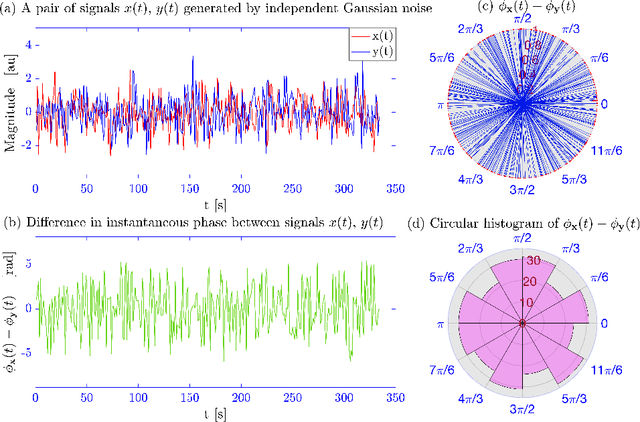
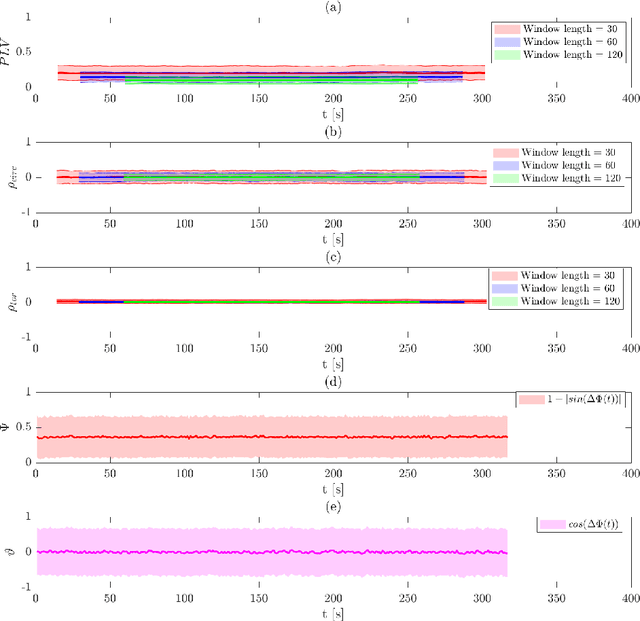
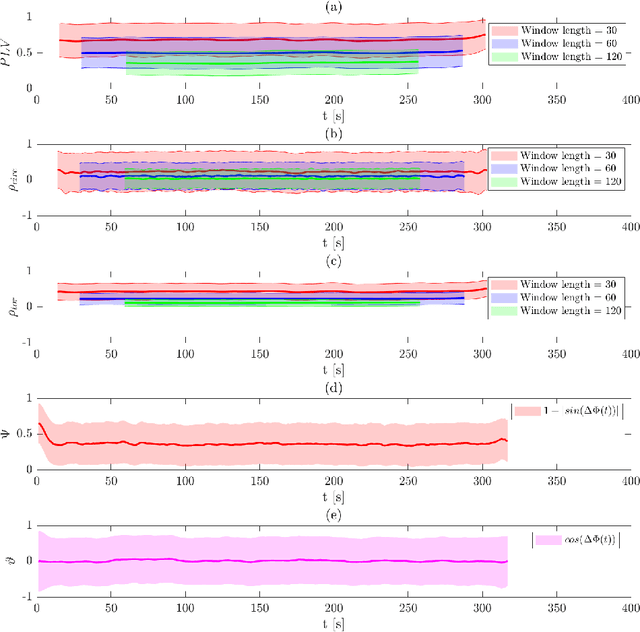
In recent years there has been growing interest in measuring time-varying functional connectivity between different brain regions using resting-state functional magnetic resonance imaging (rs-fMRI) data. One way to assess the relationship between signals from different brain regions is to measure their phase synchronization (PS) across time. There are several ways to perform such analyses, and here we compare methods that utilize a PS metric together with a sliding window, referred to here as windowed phase synchronization (WPS), with those that directly measure the instantaneous phase synchronization (IPS). In particular, IPS has recently gained popularity as it offers single time-point resolution of time-resolved fMRI connectivity. In this paper, we discuss the underlying assumptions required for performing PS analyses and emphasize the necessity of band-pass filtering the data to obtain valid results. We review various methods for evaluating PS and introduce a new approach within the IPS framework denoted the cosine of the relative phase (CRP). We contrast methods through a series of simulations and application to rs-fMRI data. Our results indicate that CRP outperforms other tested methods and overcomes issues related to undetected temporal transitions from positive to negative associations common in IPS analysis. Further, in contrast to phase coherence, CRP unfolds the distribution of PS measures, which benefits subsequent clustering of PS matrices into recurring brain states.
Physical model simulator-trained neural network for computational 3D phase imaging of multiple-scattering samples
Mar 29, 2021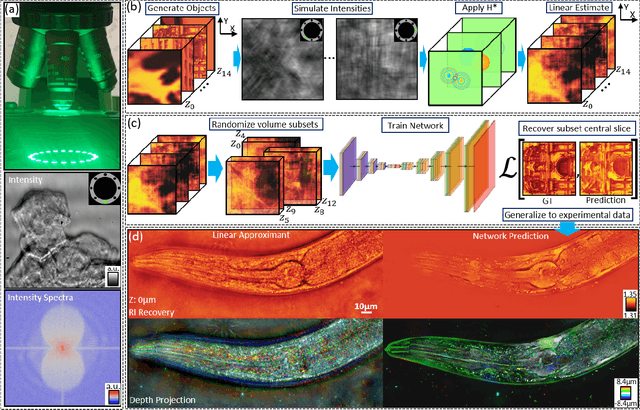
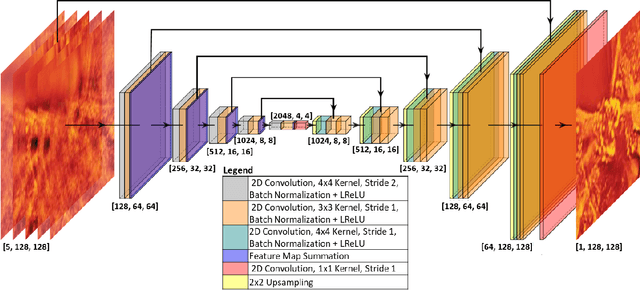
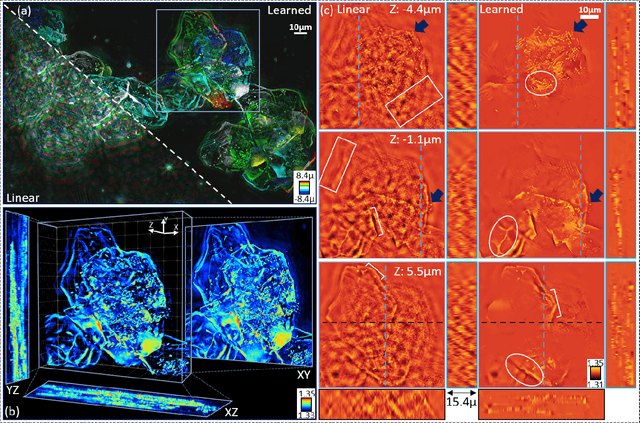
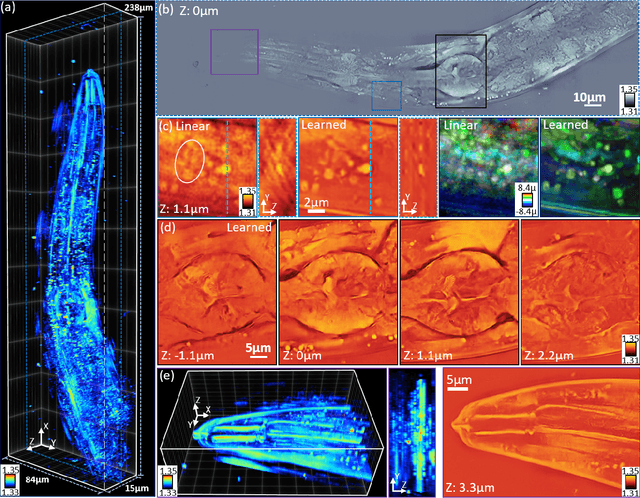
Recovering 3D phase features of complex, multiple-scattering biological samples traditionally sacrifices computational efficiency and processing time for physical model accuracy and reconstruction quality. This trade-off hinders the rapid analysis of living, dynamic biological samples that are often of greatest interest to biological research. Here, we overcome this bottleneck by combining annular intensity diffraction tomography (aIDT) with an approximant-guided deep learning framework. Using a novel physics model simulator-based learning strategy trained entirely on natural image datasets, we show our network can robustly reconstruct complex 3D biological samples of arbitrary size and structure. This approach highlights that large-scale multiple-scattering models can be leveraged in place of acquiring experimental datasets for achieving highly generalizable deep learning models. We devise a new model-based data normalization pre-processing procedure for homogenizing the sample contrast and achieving uniform prediction quality regardless of scattering strength. To achieve highly efficient training and prediction, we implement a lightweight 2D network structure that utilizes a multi-channel input for encoding the axial information. We demonstrate this framework's capabilities on experimental measurements of epithelial buccal cells and Caenorhabditis elegans worms. We highlight the robustness of this approach by evaluating dynamic samples on a living worm video, and we emphasize our approach's generalizability by recovering algae samples evaluated with different experimental setups. To assess the prediction quality, we develop a novel quantitative evaluation metric and show that our predictions are consistent with our experimental measurements and multiple-scattering physics.
A Real-time and Registration-free Framework for Dynamic Shape Instantiation
Dec 30, 2017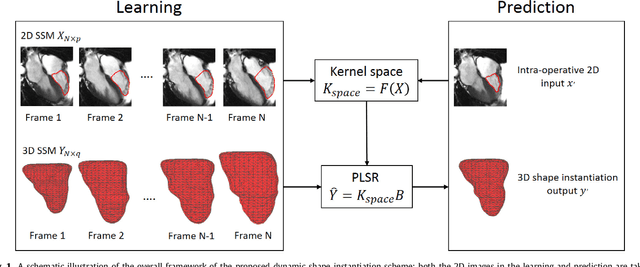
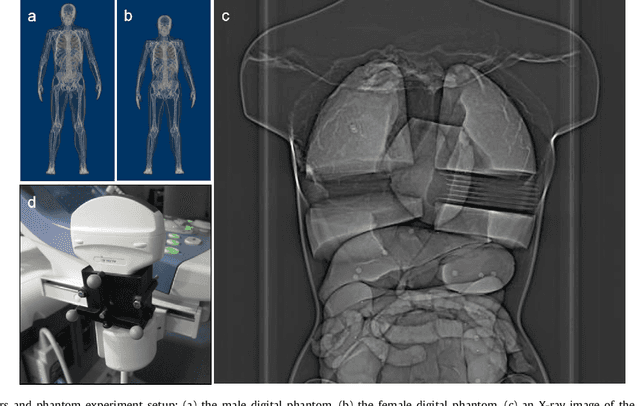
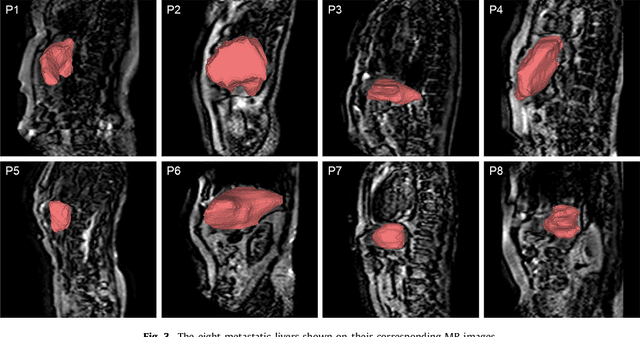
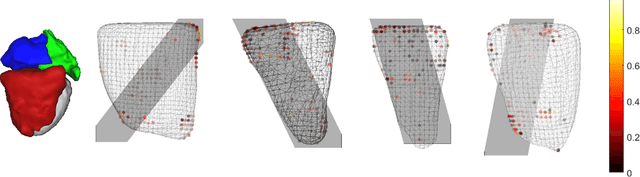
Real-time 3D navigation during minimally invasive procedures is an essential yet challenging task, especially when considerable tissue motion is involved. To balance image acquisition speed and resolution, only 2D images or low-resolution 3D volumes can be used clinically. In this paper, a real-time and registration-free framework for dynamic shape instantiation, generalizable to multiple anatomical applications, is proposed to instantiate high-resolution 3D shapes of an organ from a single 2D image intra-operatively. Firstly, an approximate optimal scan plane was determined by analyzing the pre-operative 3D statistical shape model (SSM) of the anatomy with sparse principal component analysis (SPCA) and considering practical constraints . Secondly, kernel partial least squares regression (KPLSR) was used to learn the relationship between the pre-operative 3D SSM and a synchronized 2D SSM constructed from 2D images obtained at the approximate optimal scan plane. Finally, the derived relationship was applied to the new intra-operative 2D image obtained at the same scan plane to predict the high-resolution 3D shape intra-operatively. A major feature of the proposed framework is that no extra registration between the pre-operative 3D SSM and the synchronized 2D SSM is required. Detailed validation was performed on studies including the liver and right ventricle (RV) of the heart. The derived results (mean accuracy of 2.19mm on patients and computation speed of 1ms) demonstrate its potential clinical value for real-time, high-resolution, dynamic and 3D interventional guidance.
* 35 Pages, 11 figures
EPNE: Evolutionary Pattern Preserving Network Embedding
Sep 24, 2020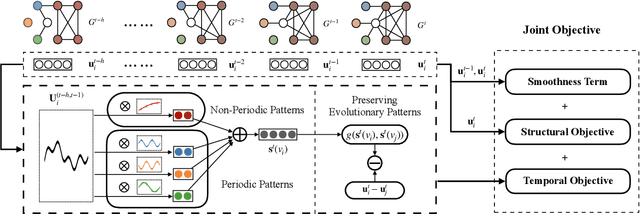
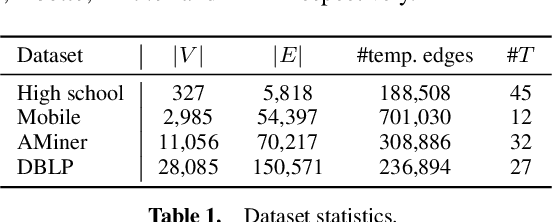
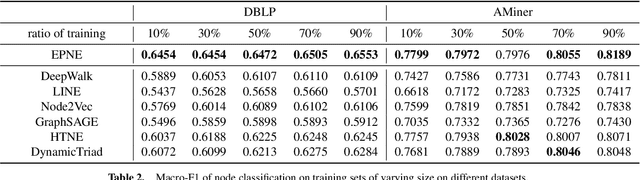
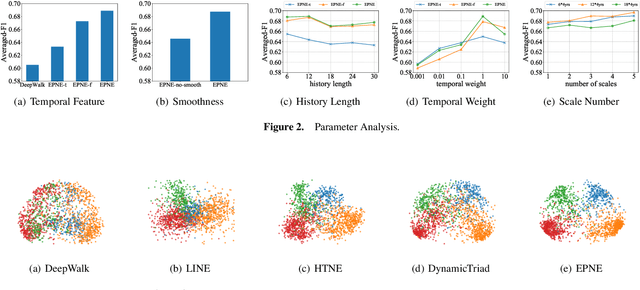
Information networks are ubiquitous and are ideal for modeling relational data. Networks being sparse and irregular, network embedding algorithms have caught the attention of many researchers, who came up with numerous embeddings algorithms in static networks. Yet in real life, networks constantly evolve over time. Hence, evolutionary patterns, namely how nodes develop itself over time, would serve as a powerful complement to static structures in embedding networks, on which relatively few works focus. In this paper, we propose EPNE, a temporal network embedding model preserving evolutionary patterns of the local structure of nodes. In particular, we analyze evolutionary patterns with and without periodicity and design strategies correspondingly to model such patterns in time-frequency domains based on causal convolutions. In addition, we propose a temporal objective function which is optimized simultaneously with proximity ones such that both temporal and structural information are preserved. With the adequate modeling of temporal information, our model is able to outperform other competitive methods in various prediction tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge