"Time": models, code, and papers
Sound Event Detection in Multichannel Audio using Convolutional Time-Frequency-Channel Squeeze and Excitation
Aug 04, 2019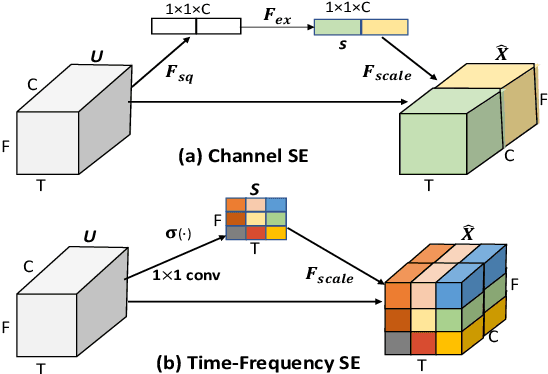
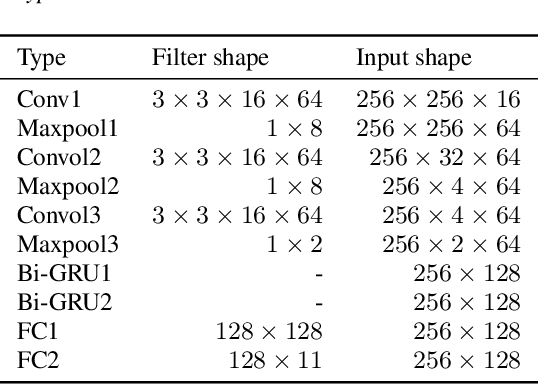
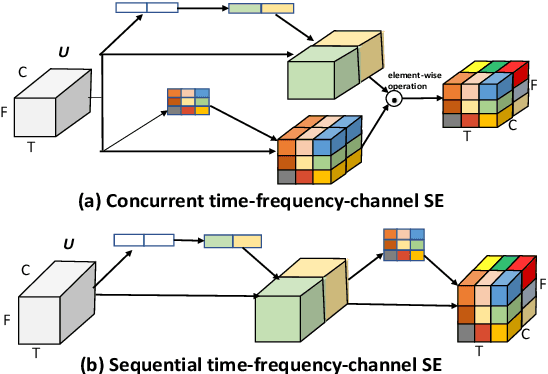
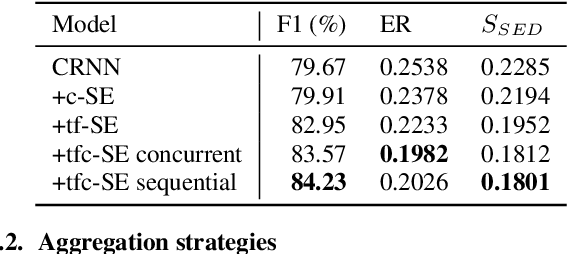
In this study, we introduce a convolutional time-frequency-channel "Squeeze and Excitation" (tfc-SE) module to explicitly model inter-dependencies between the time-frequency domain and multiple channels. The tfc-SE module consists of two parts: tf-SE block and c-SE block which are designed to provide attention on time-frequency and channel domain, respectively, for adaptively recalibrating the input feature map. The proposed tfc-SE module, together with a popular Convolutional Recurrent Neural Network (CRNN) model, are evaluated on a multi-channel sound event detection task with overlapping audio sources: the training and test data are synthesized TUT Sound Events 2018 datasets, recorded with microphone arrays. We show that the tfc-SE module can be incorporated into the CRNN model at a small additional computational cost and bring significant improvements on sound event detection accuracy. We also perform detailed ablation studies by analyzing various factors that may influence the performance of the SE blocks. We show that with the best tfc-SE block, error rate (ER) decreases from 0.2538 to 0.2026, relative 20.17\% reduction of ER, and 5.72\% improvement of F1 score. The results indicate that the learned acoustic embeddings with the tfc-SE module efficiently strengthen time-frequency and channel-wise feature representations to improve the discriminative performance.
IrisNet: Deep Learning for Automatic and Real-time Tongue Contour Tracking in Ultrasound Video Data using Peripheral Vision
Nov 10, 2019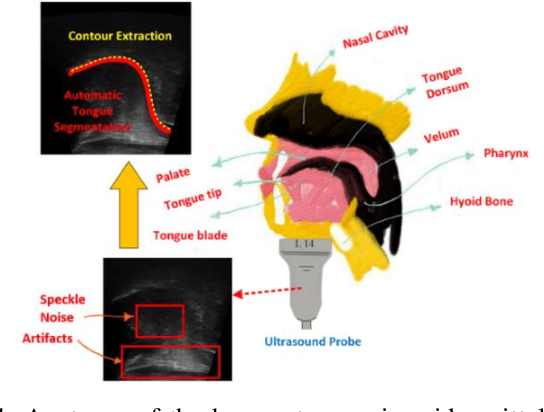
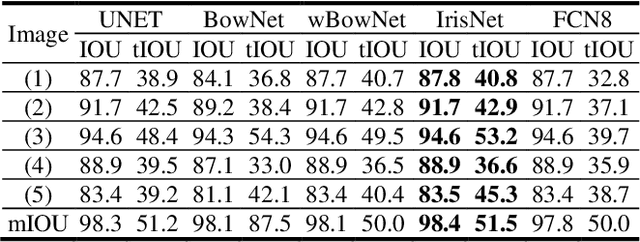

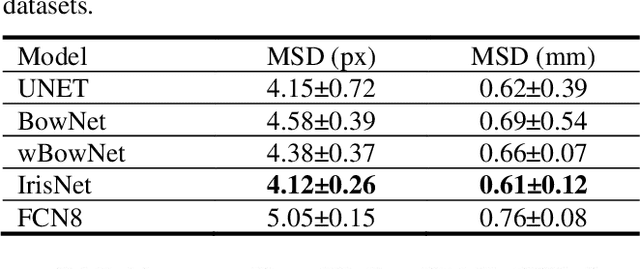
The progress of deep convolutional neural networks has been successfully exploited in various real-time computer vision tasks such as image classification and segmentation. Owing to the development of computational units, availability of digital datasets, and improved performance of deep learning models, fully automatic and accurate tracking of tongue contours in real-time ultrasound data became practical only in recent years. Recent studies have shown that the performance of deep learning techniques is significant in the tracking of ultrasound tongue contours in real-time applications such as pronunciation training using multimodal ultrasound-enhanced approaches. Due to the high correlation between ultrasound tongue datasets, it is feasible to have a general model that accomplishes automatic tongue tracking for almost all datasets. In this paper, we proposed a deep learning model comprises of a convolutional module mimicking the peripheral vision ability of the human eye to handle real-time, accurate, and fully automatic tongue contour tracking tasks, applicable for almost all primary ultrasound tongue datasets. Qualitative and quantitative assessment of IrisNet on different ultrasound tongue datasets and PASCAL VOC2012 revealed its outstanding generalization achievement in compare with similar techniques.
Continual Novelty Detection
Jun 24, 2021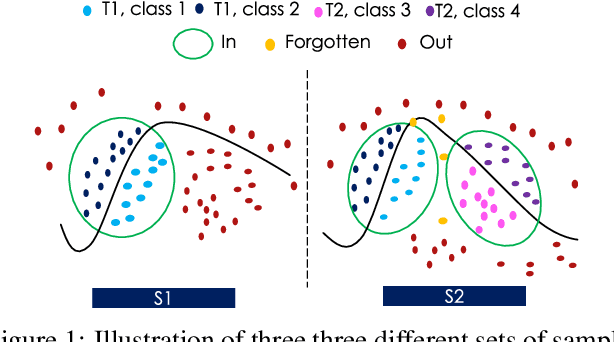
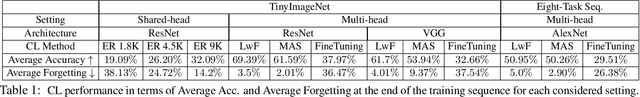

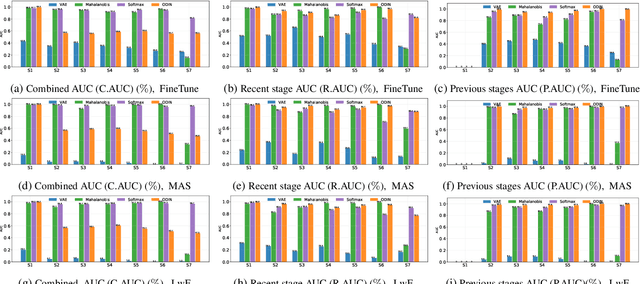

Novelty Detection methods identify samples that are not representative of a model's training set thereby flagging misleading predictions and bringing a greater flexibility and transparency at deployment time. However, research in this area has only considered Novelty Detection in the offline setting. Recently, there has been a growing realization in the computer vision community that applications demand a more flexible framework - Continual Learning - where new batches of data representing new domains, new classes or new tasks become available at different points in time. In this setting, Novelty Detection becomes more important, interesting and challenging. This work identifies the crucial link between the two problems and investigates the Novelty Detection problem under the Continual Learning setting. We formulate the Continual Novelty Detection problem and present a benchmark, where we compare several Novelty Detection methods under different Continual Learning settings. We show that Continual Learning affects the behaviour of novelty detection algorithms, while novelty detection can pinpoint insights in the behaviour of a continual learner. We further propose baselines and discuss possible research directions. We believe that the coupling of the two problems is a promising direction to bring vision models into practice.
NonSTOP: A NonSTationary Online Prediction Method for Time Series
Aug 26, 2018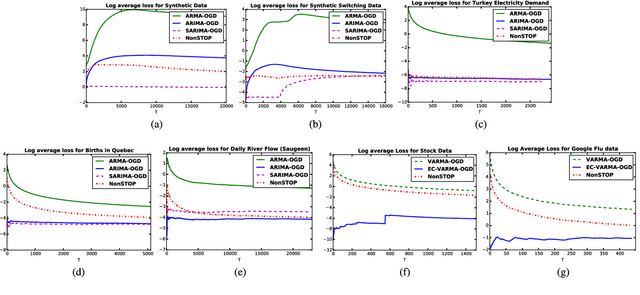
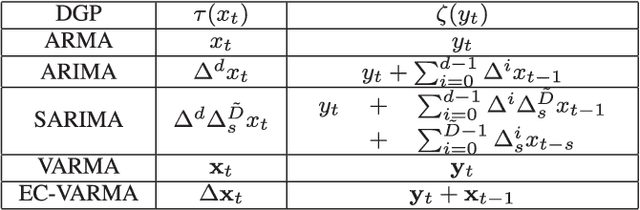
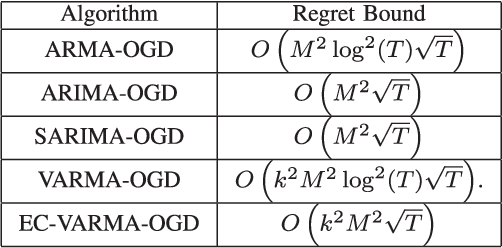


We present online prediction methods for time series that let us explicitly handle nonstationary artifacts (e.g. trend and seasonality) present in most real time series. Specifically, we show that applying appropriate transformations to such time series before prediction can lead to improved theoretical and empirical prediction performance. Moreover, since these transformations are usually unknown, we employ the learning with experts setting to develop a fully online method (NonSTOP-NonSTationary Online Prediction) for predicting nonstationary time series. This framework allows for seasonality and/or other trends in univariate time series and cointegration in multivariate time series. Our algorithms and regret analysis subsume recent related work while significantly expanding the applicability of such methods. For all the methods, we provide sub-linear regret bounds using relaxed assumptions. The theoretical guarantees do not fully capture the benefits of the transformations, thus we provide a data-dependent analysis of the follow-the-leader algorithm that provides insight into the success of using such transformations. We support all of our results with experiments on simulated and real data.
First Order Methods take Exponential Time to Converge to Global Minimizers of Non-Convex Functions
Feb 28, 2020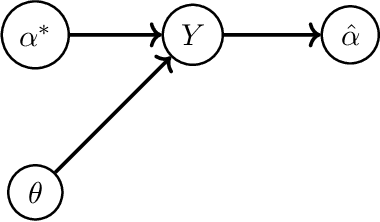
Machine learning algorithms typically perform optimization over a class of non-convex functions. In this work, we provide bounds on the fundamental hardness of identifying the global minimizer of a non convex function. Specifically, we design a family of parametrized non-convex functions and employ statistical lower bounds for parameter estimation. We show that the parameter estimation problem is equivalent to the problem of function identification in the given family. We then claim that non convex optimization is at least as hard as function identification. Jointly, we prove that any first order method can take exponential time to converge to a global minimizer.
Knowledge distillation from multi-modal to mono-modal segmentation networks
Jun 17, 2021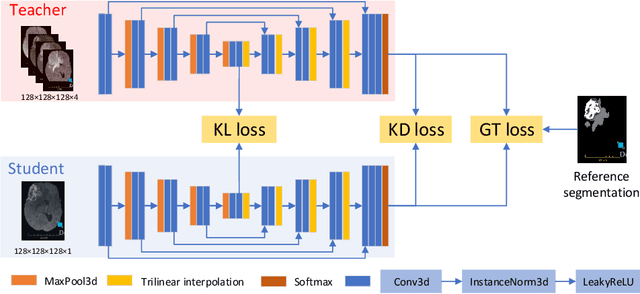
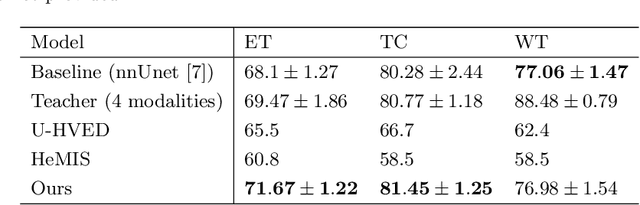
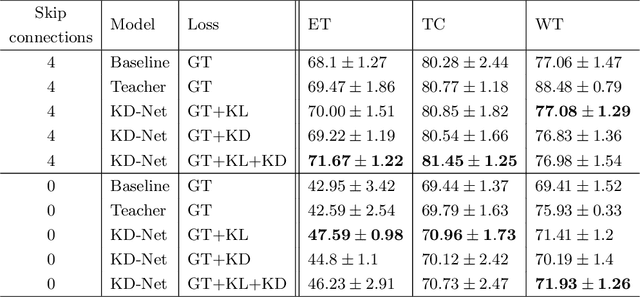
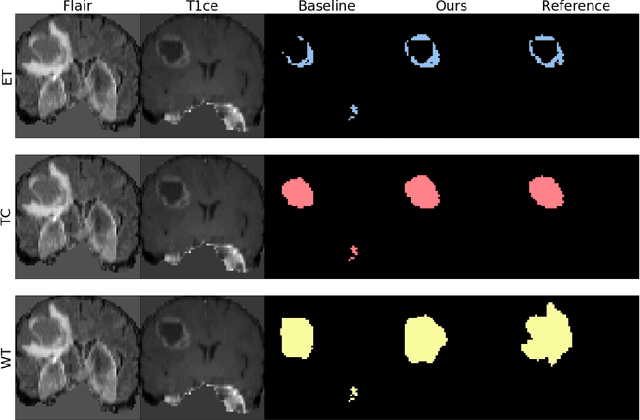
The joint use of multiple imaging modalities for medical image segmentation has been widely studied in recent years. The fusion of information from different modalities has demonstrated to improve the segmentation accuracy, with respect to mono-modal segmentations, in several applications. However, acquiring multiple modalities is usually not possible in a clinical setting due to a limited number of physicians and scanners, and to limit costs and scan time. Most of the time, only one modality is acquired. In this paper, we propose KD-Net, a framework to transfer knowledge from a trained multi-modal network (teacher) to a mono-modal one (student). The proposed method is an adaptation of the generalized distillation framework where the student network is trained on a subset (1 modality) of the teacher's inputs (n modalities). We illustrate the effectiveness of the proposed framework in brain tumor segmentation with the BraTS 2018 dataset. Using different architectures, we show that the student network effectively learns from the teacher and always outperforms the baseline mono-modal network in terms of segmentation accuracy.
* MICCAI 2020
Backdoor Attacks on Network Certification via Data Poisoning
Aug 25, 2021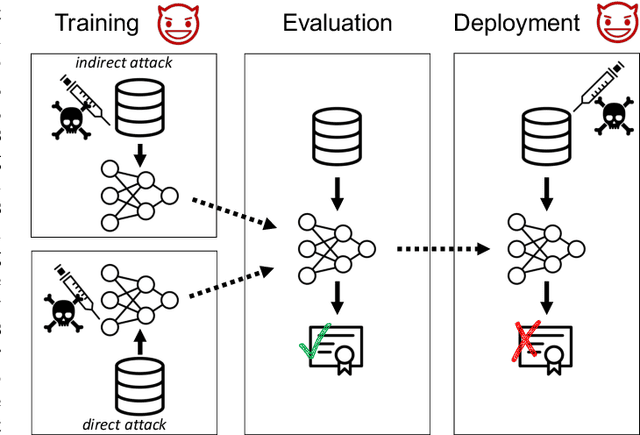

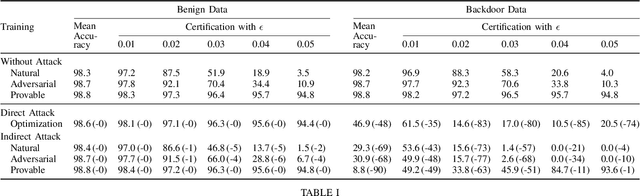
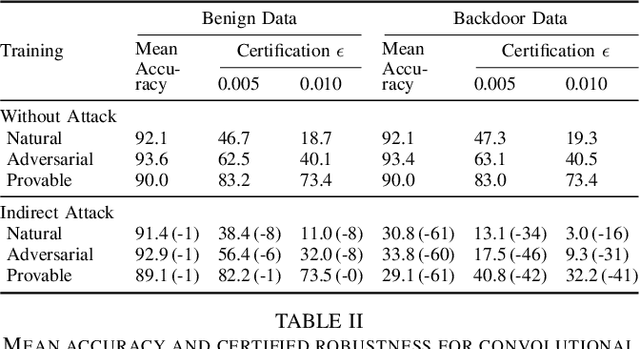
Certifiers for neural networks have made great progress towards provable robustness guarantees against evasion attacks using adversarial examples. However, introducing certifiers into deep learning systems also opens up new attack vectors, which need to be considered before deployment. In this work, we conduct the first systematic analysis of training time attacks against certifiers in practical application pipelines, identifying new threat vectors that can be exploited to degrade the overall system. Using these insights, we design two backdoor attacks against network certifiers, which can drastically reduce certified robustness when the backdoor is activated. For example, adding 1% poisoned data points during training is sufficient to reduce certified robustness by up to 95 percentage points, effectively rendering the certifier useless. We analyze how such novel attacks can compromise the overall system's integrity or availability. Our extensive experiments across multiple datasets, model architectures, and certifiers demonstrate the wide applicability of these attacks. A first investigation into potential defenses shows that current approaches only partially mitigate the issue, highlighting the need for new, more specific solutions.
Deep SIMBAD: Active Landmark-based Self-localization Using Ranking -based Scene Descriptor
Sep 06, 2021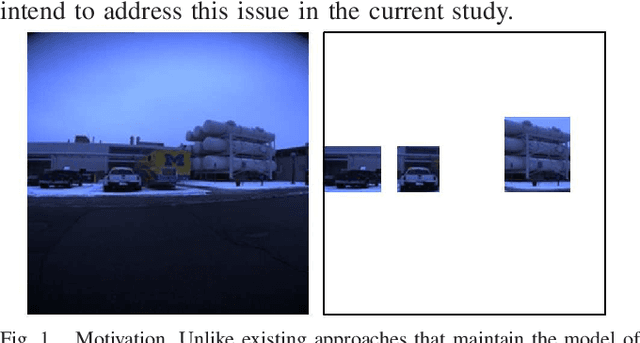
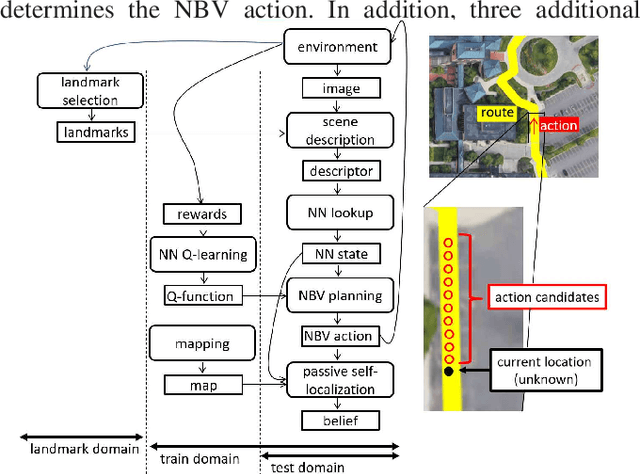
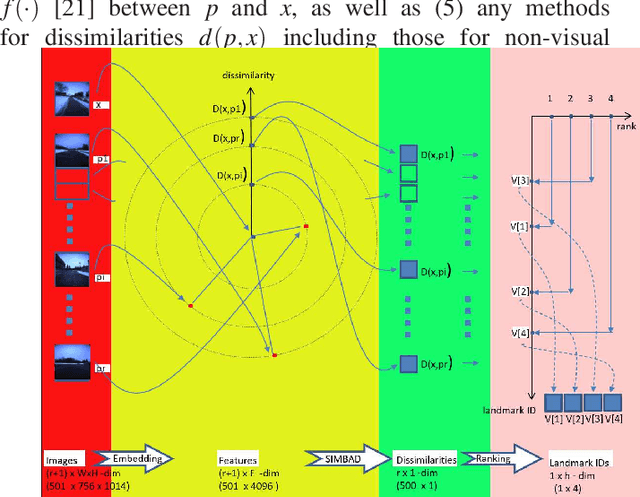
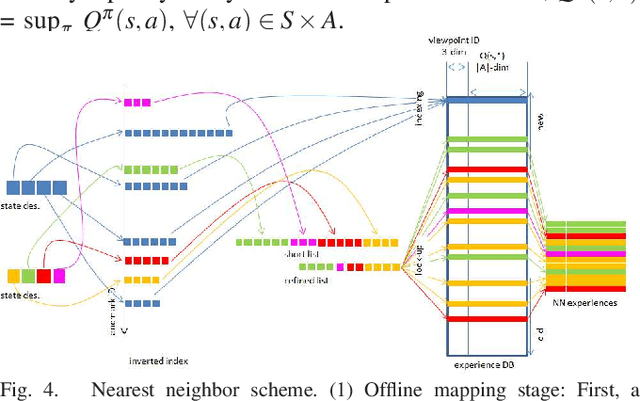
Landmark-based robot self-localization has recently garnered interest as a highly-compressive domain-invariant approach for performing visual place recognition (VPR) across domains (e.g., time of day, weather, and season). However, landmark-based self-localization can be an ill-posed problem for a passive observer (e.g., manual robot control), as many viewpoints may not provide an effective landmark view. In this study, we consider an active self-localization task by an active observer and present a novel reinforcement learning (RL)-based next-best-view (NBV) planner. Our contributions are as follows. (1) SIMBAD-based VPR: We formulate the problem of landmark-based compact scene description as SIMBAD (similarity-based pattern recognition) and further present its deep learning extension. (2) VPR-to-NBV knowledge transfer: We address the challenge of RL under uncertainty (i.e., active self-localization) by transferring the state recognition ability of VPR to the NBV. (3) NNQL-based NBV: We regard the available VPR as the experience database by adapting nearest-neighbor approximation of Q-learning (NNQL). The result shows an extremely compact data structure that compresses both the VPR and NBV into a single incremental inverted index. Experiments using the public NCLT dataset validated the effectiveness of the proposed approach.
Image Complexity Guided Network Compression for Biomedical Image Segmentation
Jul 06, 2021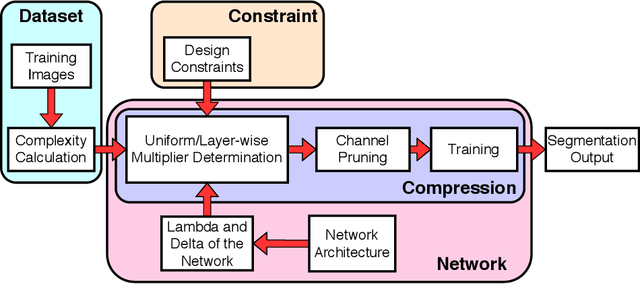
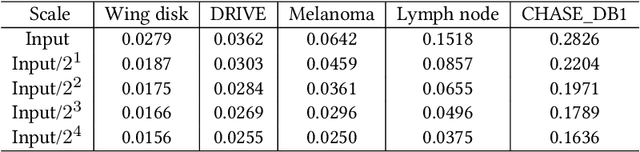
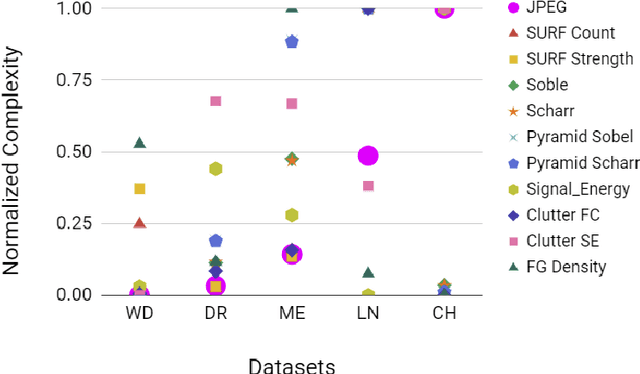
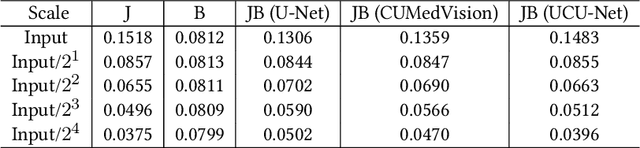
Compression is a standard procedure for making convolutional neural networks (CNNs) adhere to some specific computing resource constraints. However, searching for a compressed architecture typically involves a series of time-consuming training/validation experiments to determine a good compromise between network size and performance accuracy. To address this, we propose an image complexity-guided network compression technique for biomedical image segmentation. Given any resource constraints, our framework utilizes data complexity and network architecture to quickly estimate a compressed model which does not require network training. Specifically, we map the dataset complexity to the target network accuracy degradation caused by compression. Such mapping enables us to predict the final accuracy for different network sizes, based on the computed dataset complexity. Thus, one may choose a solution that meets both the network size and segmentation accuracy requirements. Finally, the mapping is used to determine the convolutional layer-wise multiplicative factor for generating a compressed network. We conduct experiments using 5 datasets, employing 3 commonly-used CNN architectures for biomedical image segmentation as representative networks. Our proposed framework is shown to be effective for generating compressed segmentation networks, retaining up to $\approx 95\%$ of the full-sized network segmentation accuracy, and at the same time, utilizing $\approx 32x$ fewer network trainable weights (average reduction) of the full-sized networks.
Large-scale quantum machine learning
Aug 25, 2021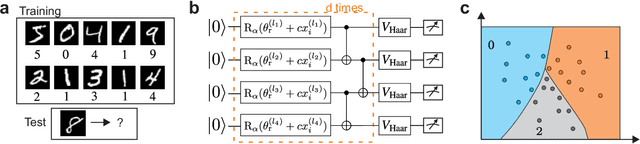
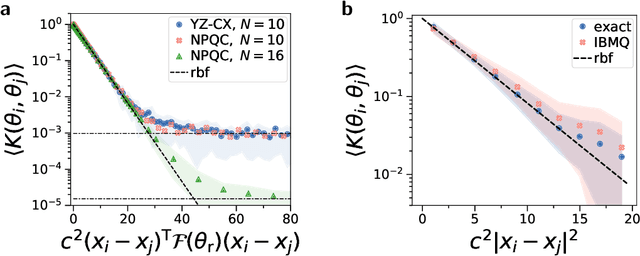
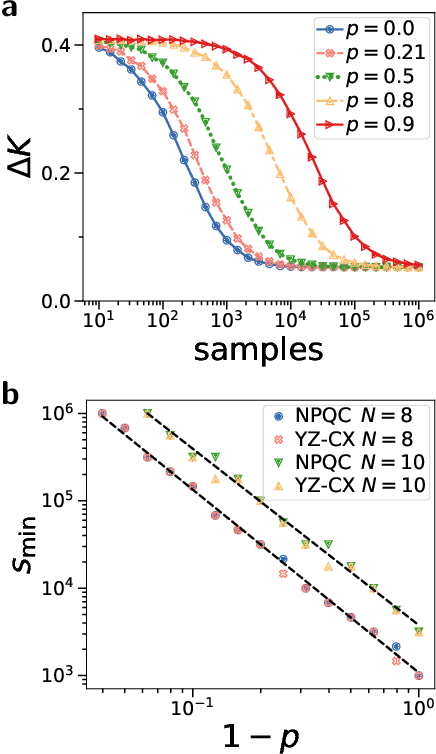
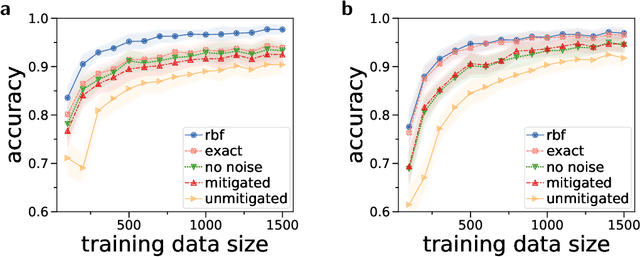
Quantum computers promise to enhance machine learning for practical applications. Quantum machine learning for real-world data has to handle extensive amounts of high-dimensional data. However, conventional methods for measuring quantum kernels are impractical for large datasets as they scale with the square of the dataset size. Here, we measure quantum kernels using randomized measurements to gain a quadratic speedup in computation time and quickly process large datasets. Further, we efficiently encode high-dimensional data into quantum computers with the number of features scaling linearly with the circuit depth. The encoding is characterized by the quantum Fisher information metric and is related to the radial basis function kernel. We demonstrate the advantages of our methods by classifying images with the IBM quantum computer. To achieve further speedups we distribute the quantum computational tasks between different quantum computers. Our approach is exceptionally robust to noise via a complementary error mitigation scheme. Using currently available quantum computers, the MNIST database can be processed within 220 hours instead of 10 years which opens up industrial applications of quantum machine learning.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge