"Time": models, code, and papers
TorchEsegeta: Framework for Interpretability and Explainability of Image-based Deep Learning Models
Oct 16, 2021
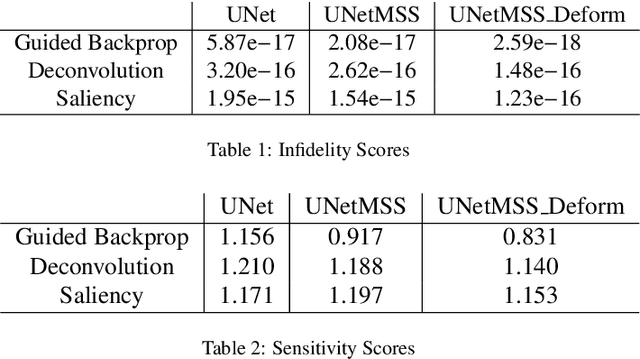
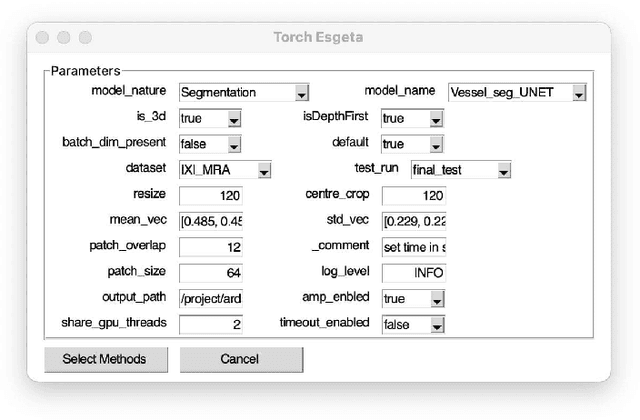
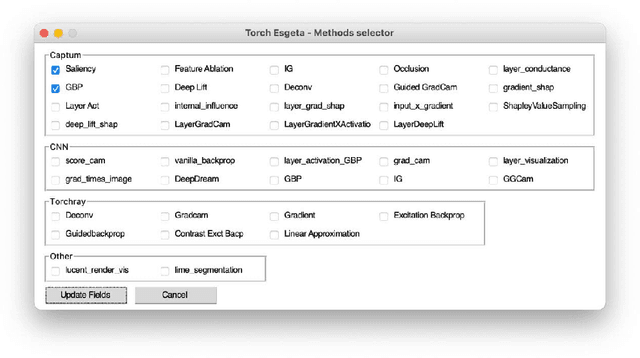
Clinicians are often very sceptical about applying automatic image processing approaches, especially deep learning based methods, in practice. One main reason for this is the black-box nature of these approaches and the inherent problem of missing insights of the automatically derived decisions. In order to increase trust in these methods, this paper presents approaches that help to interpret and explain the results of deep learning algorithms by depicting the anatomical areas which influence the decision of the algorithm most. Moreover, this research presents a unified framework, TorchEsegeta, for applying various interpretability and explainability techniques for deep learning models and generate visual interpretations and explanations for clinicians to corroborate their clinical findings. In addition, this will aid in gaining confidence in such methods. The framework builds on existing interpretability and explainability techniques that are currently focusing on classification models, extending them to segmentation tasks. In addition, these methods have been adapted to 3D models for volumetric analysis. The proposed framework provides methods to quantitatively compare visual explanations using infidelity and sensitivity metrics. This framework can be used by data scientists to perform post-hoc interpretations and explanations of their models, develop more explainable tools and present the findings to clinicians to increase their faith in such models. The proposed framework was evaluated based on a use case scenario of vessel segmentation models trained on Time-of-fight (TOF) Magnetic Resonance Angiogram (MRA) images of the human brain. Quantitative and qualitative results of a comparative study of different models and interpretability methods are presented. Furthermore, this paper provides an extensive overview of several existing interpretability and explainability methods.
You should evaluate your language model on marginal likelihood over tokenisations
Sep 21, 2021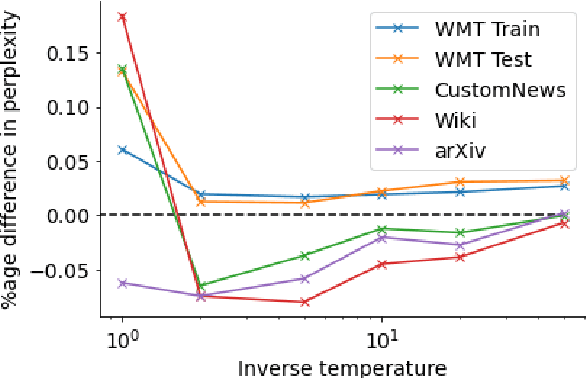
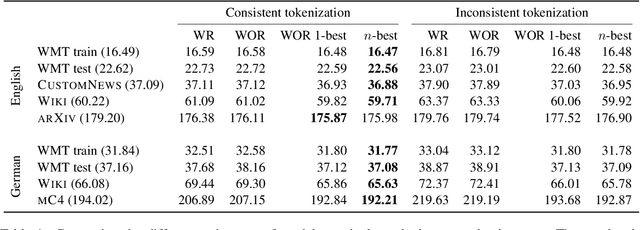
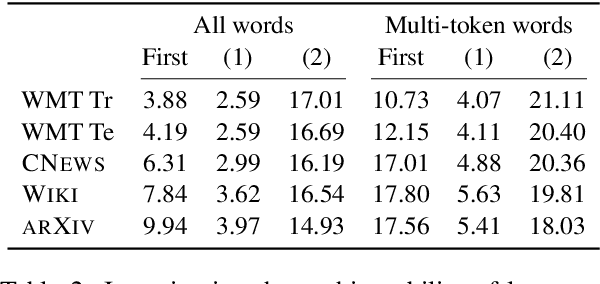
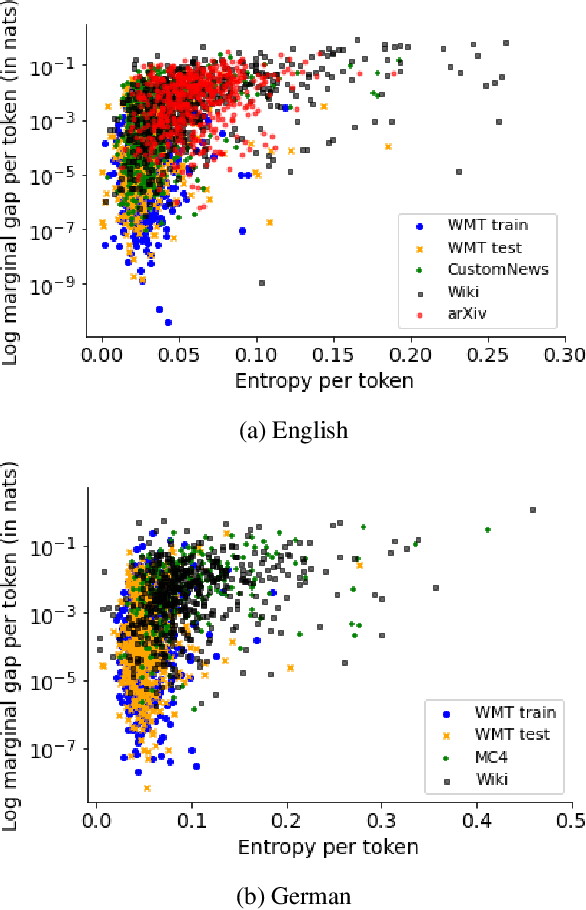
Neural language models typically tokenise input text into sub-word units to achieve an open vocabulary. The standard approach is to use a single canonical tokenisation at both train and test time. We suggest that this approach is unsatisfactory and may bottleneck our evaluation of language model performance. Using only the one-best tokenisation ignores tokeniser uncertainty over alternative tokenisations, which may hurt model out-of-domain performance. In this paper, we argue that instead, language models should be evaluated on their marginal likelihood over tokenisations. We compare different estimators for the marginal likelihood based on sampling, and show that it is feasible to estimate the marginal likelihood with a manageable number of samples. We then evaluate pretrained English and German language models on both the one-best-tokenisation and marginal perplexities, and show that the marginal perplexity can be significantly better than the one best, especially on out-of-domain data. We link this difference in perplexity to the tokeniser uncertainty as measured by tokeniser entropy. We discuss some implications of our results for language model training and evaluation, particularly with regard to tokenisation robustness.
Heterogeneous Treatment Effect Estimation using machine learning for Healthcare application: tutorial and benchmark
Sep 27, 2021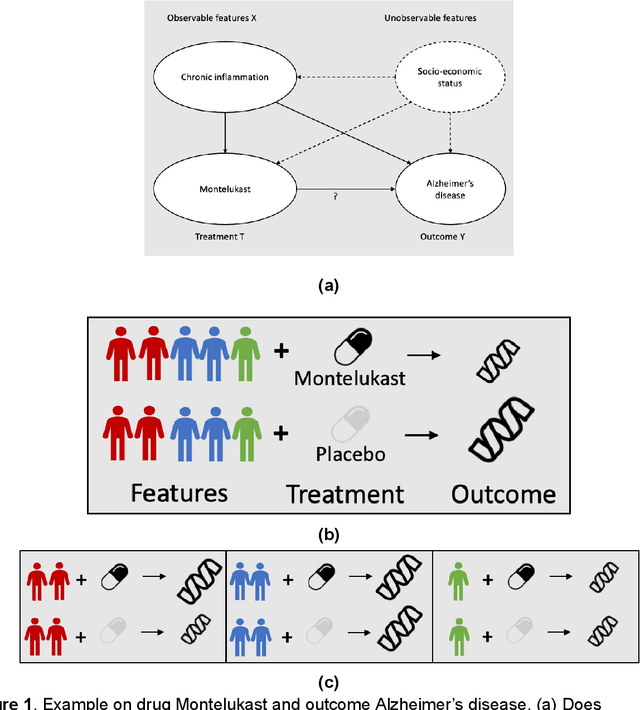
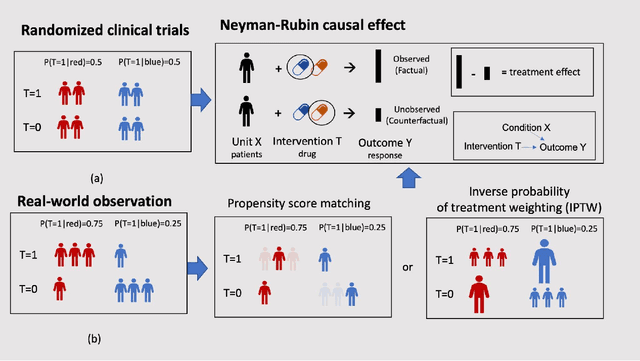
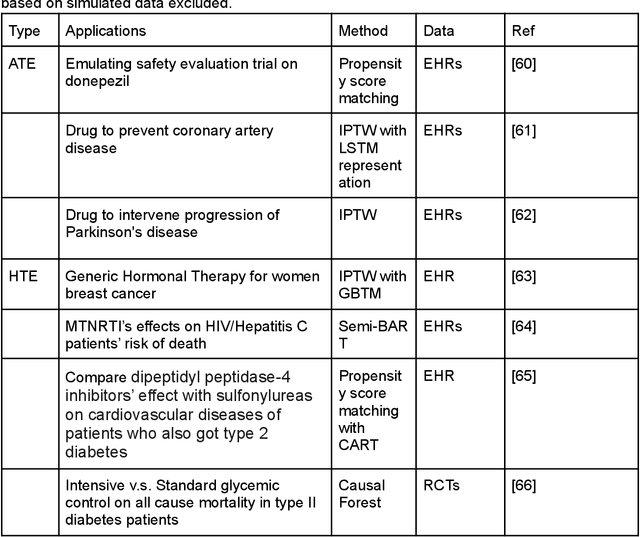
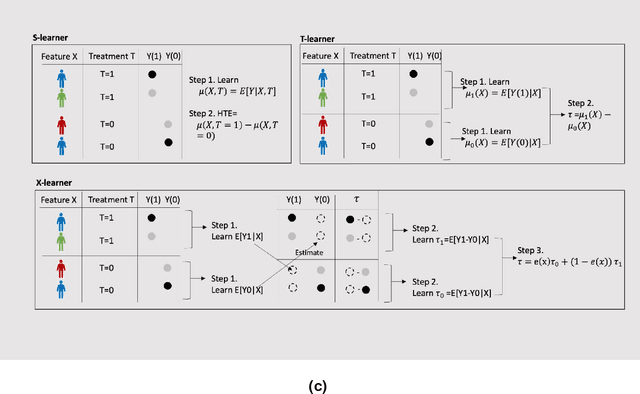
Developing new drugs for target diseases is a time-consuming and expensive task, drug repurposing has become a popular topic in the drug development field. As much health claim data become available, many studies have been conducted on the data. The real-world data is noisy, sparse, and has many confounding factors. In addition, many studies have shown that drugs effects are heterogeneous among the population. Lots of advanced machine learning models about estimating heterogeneous treatment effects (HTE) have emerged in recent years, and have been applied to in econometrics and machine learning communities. These studies acknowledge medicine and drug development as the main application area, but there has been limited translational research from the HTE methodology to drug development. We aim to introduce the HTE methodology to the healthcare area and provide feasibility consideration when translating the methodology with benchmark experiments on healthcare administrative claim data. Also, we want to use benchmark experiments to show how to interpret and evaluate the model when it is applied to healthcare research. By introducing the recent HTE techniques to a broad readership in biomedical informatics communities, we expect to promote the wide adoption of causal inference using machine learning. We also expect to provide the feasibility of HTE for personalized drug effectiveness.
FaceCook: Face Generation Based on Linear Scaling Factors
Sep 08, 2021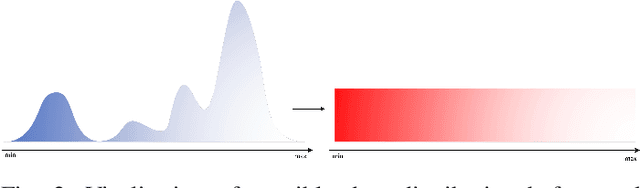
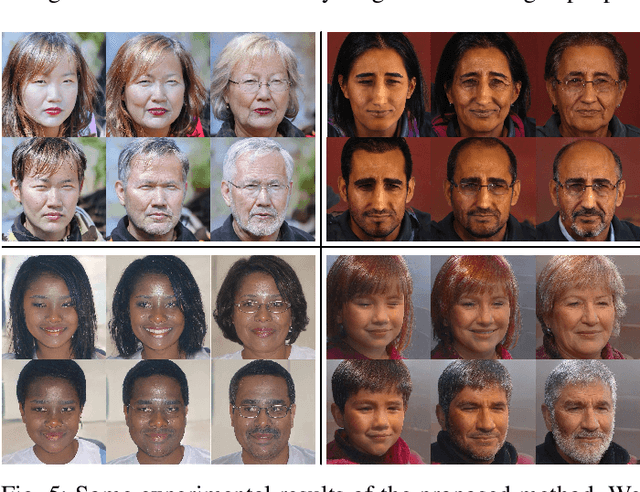
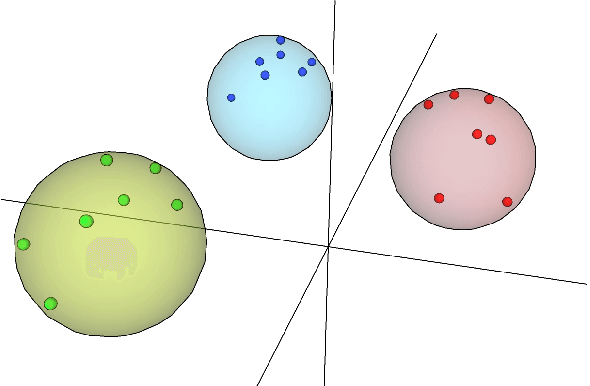
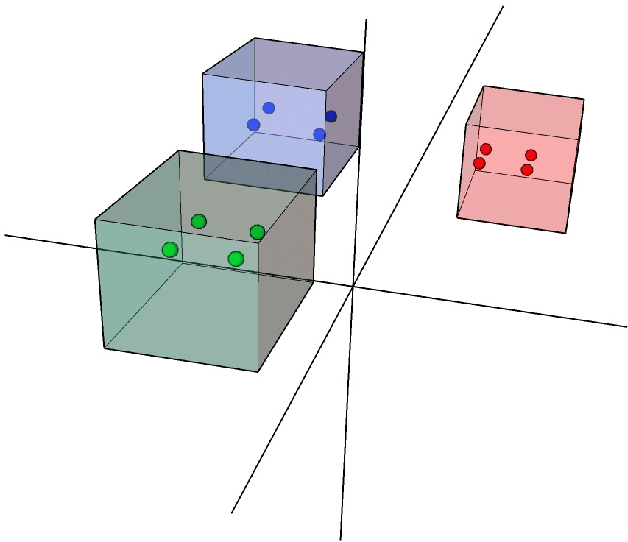

With the excellent disentanglement properties of state-of-the-art generative models, image editing has been the dominant approach to control the attributes of synthesised face images. However, these edited results often suffer from artifacts or incorrect feature rendering, especially when there is a large discrepancy between the image to be edited and the desired feature set. Therefore, we propose a new approach to mapping the latent vectors of the generative model to the scaling factors through solving a set of multivariate linear equations. The coefficients of the equations are the eigenvectors of the weight parameters of the pre-trained model, which form the basis of a hyper coordinate system. The qualitative and quantitative results both show that the proposed method outperforms the baseline in terms of image diversity. In addition, the method is much more time-efficient because you can obtain synthesised images with desirable features directly from the latent vectors, rather than the former process of editing randomly generated images requiring many processing steps.
The importance of space and time in neuromorphic cognitive agents
Feb 26, 2019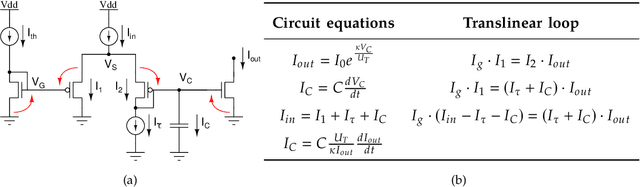
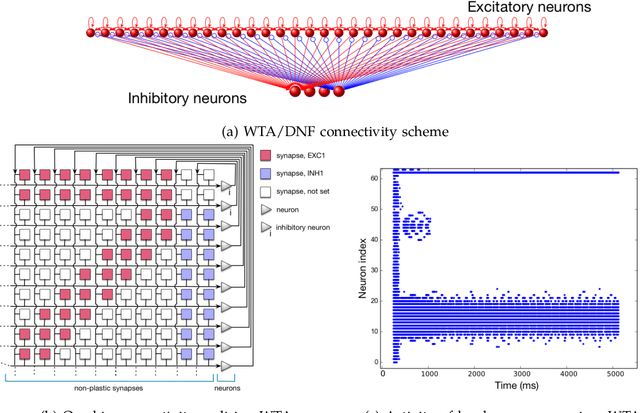
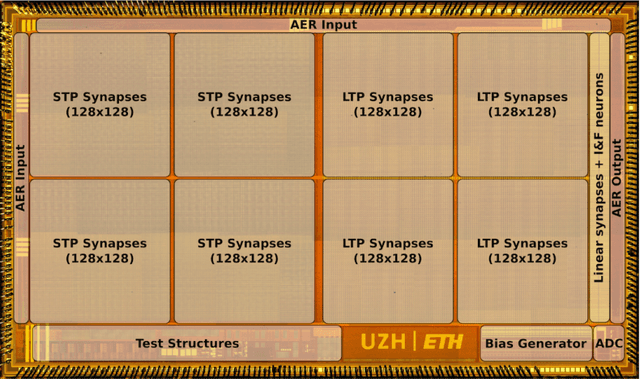
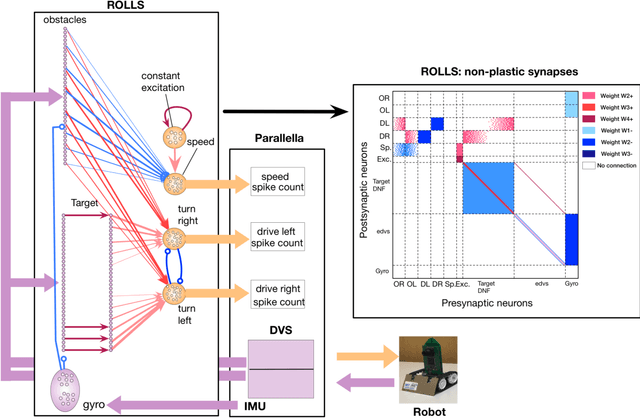
Artificial neural networks and computational neuroscience models have made tremendous progress, allowing computers to achieve impressive results in artificial intelligence (AI) applications, such as image recognition, natural language processing, or autonomous driving. Despite this remarkable progress, biological neural systems consume orders of magnitude less energy than today's artificial neural networks and are much more agile and adaptive. This efficiency and adaptivity gap is partially explained by the computing substrate of biological neural processing systems that is fundamentally different from the way today's computers are built. Biological systems use in-memory computing elements operating in a massively parallel way rather than time-multiplexed computing units that are reused in a sequential fashion. Moreover, activity of biological neurons follows continuous-time dynamics in real, physical time, instead of operating on discrete temporal cycles abstracted away from real-time. Here, we present neuromorphic processing devices that emulate the biological style of processing by using parallel instances of mixed-signal analog/digital circuits that operate in real time. We argue that this approach brings significant advantages in efficiency of computation. We show examples of embodied neuromorphic agents that use such devices to interact with the environment and exhibit autonomous learning.
HUNTER: AI based Holistic Resource Management for Sustainable Cloud Computing
Oct 11, 2021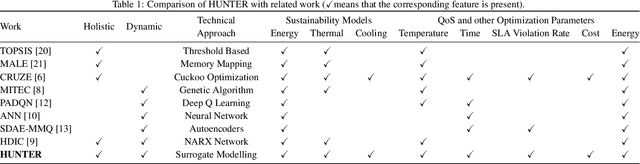
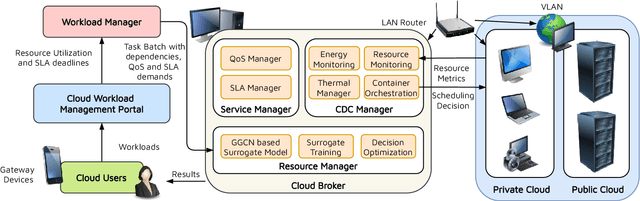
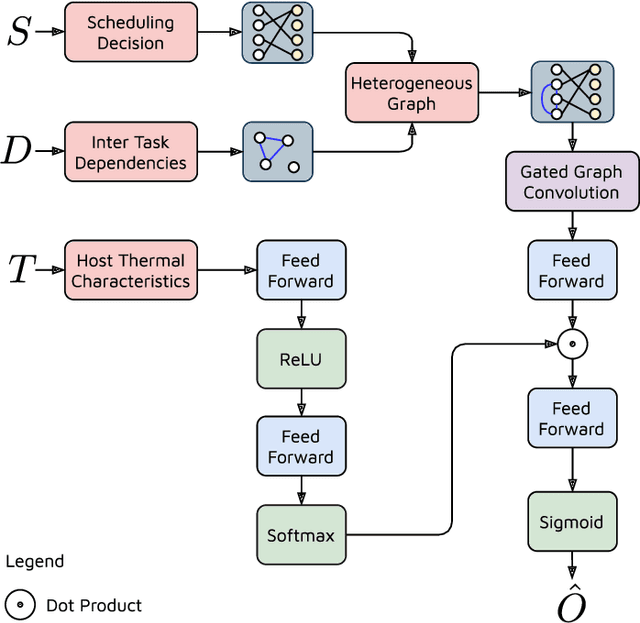
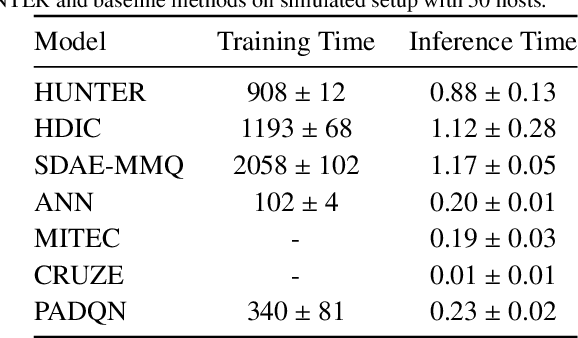
The worldwide adoption of cloud data centers (CDCs) has given rise to the ubiquitous demand for hosting application services on the cloud. Further, contemporary data-intensive industries have seen a sharp upsurge in the resource requirements of modern applications. This has led to the provisioning of an increased number of cloud servers, giving rise to higher energy consumption and, consequently, sustainability concerns. Traditional heuristics and reinforcement learning based algorithms for energy-efficient cloud resource management address the scalability and adaptability related challenges to a limited extent. Existing work often fails to capture dependencies across thermal characteristics of hosts, resource consumption of tasks and the corresponding scheduling decisions. This leads to poor scalability and an increase in the compute resource requirements, particularly in environments with non-stationary resource demands. To address these limitations, we propose an artificial intelligence (AI) based holistic resource management technique for sustainable cloud computing called HUNTER. The proposed model formulates the goal of optimizing energy efficiency in data centers as a multi-objective scheduling problem, considering three important models: energy, thermal and cooling. HUNTER utilizes a Gated Graph Convolution Network as a surrogate model for approximating the Quality of Service (QoS) for a system state and generating optimal scheduling decisions. Experiments on simulated and physical cloud environments using the CloudSim toolkit and the COSCO framework show that HUNTER outperforms state-of-the-art baselines in terms of energy consumption, SLA violation, scheduling time, cost and temperature by up to 12, 35, 43, 54 and 3 percent respectively.
Feature Identification and Matching for Hand Hygiene Pose
Aug 14, 2021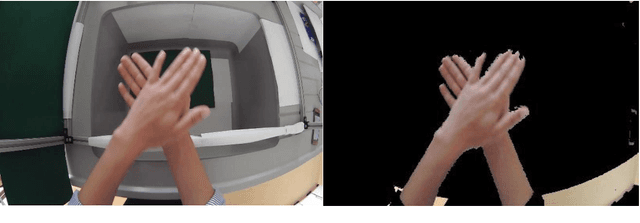
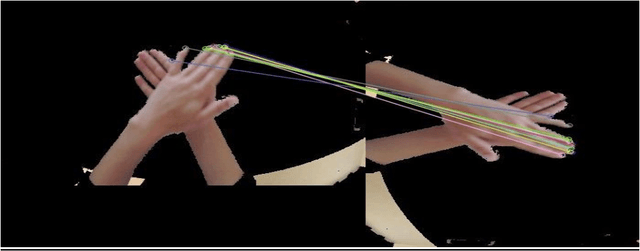
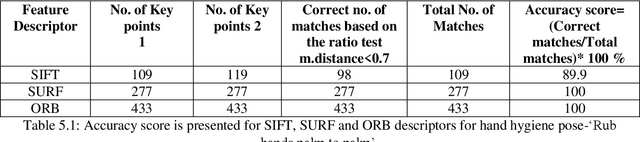


Three popular feature descriptors of computer vision such as SIFT, SURF, and ORB compared and evaluated. The number of correct features extracted and matched for the original hand hygiene pose-Rub hands palm to palm image and rotated image. An accuracy score calculated based on the total number of matches and the correct number of matches produced. The experiment demonstrated that ORB algorithm outperforms by giving the high number of correct matches in less amount of time. ORB feature detection technique applied over handwashing video recordings for feature extraction and hand hygiene pose classification as a future work. OpenCV utilized to apply the algorithms within python scripts.
Real-Time Semantic Segmentation via Auto Depth, Downsampling Joint Decision and Feature Aggregation
Mar 31, 2020
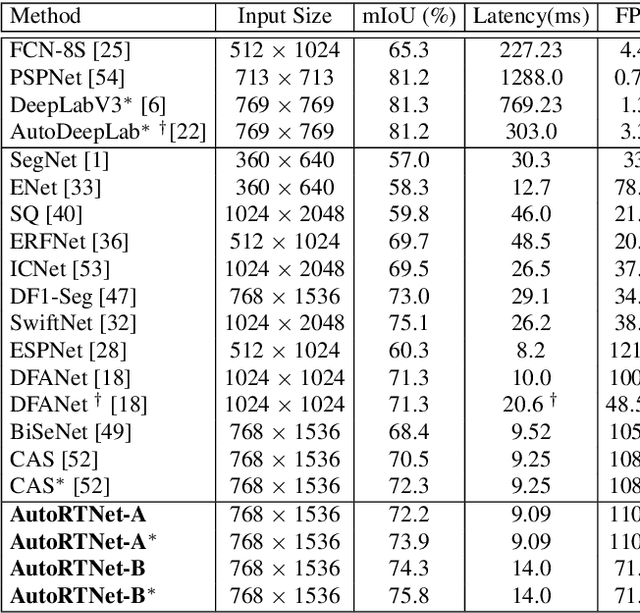

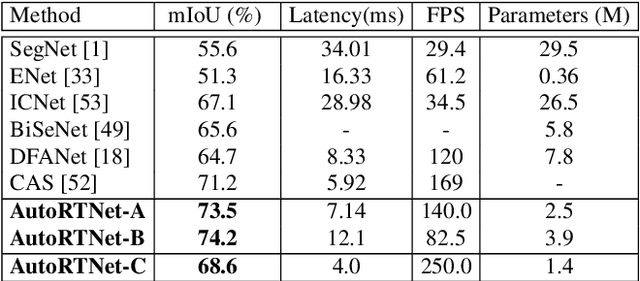
To satisfy the stringent requirements on computational resources in the field of real-time semantic segmentation, most approaches focus on the hand-crafted design of light-weight segmentation networks. Recently, Neural Architecture Search (NAS) has been used to search for the optimal building blocks of networks automatically, but the network depth, downsampling strategy, and feature aggregation way are still set in advance by trial and error. In this paper, we propose a joint search framework, called AutoRTNet, to automate the design of these strategies. Specifically, we propose hyper-cells to jointly decide the network depth and downsampling strategy, and an aggregation cell to achieve automatic multi-scale feature aggregation. Experimental results show that AutoRTNet achieves 73.9% mIoU on the Cityscapes test set and 110.0 FPS on an NVIDIA TitanXP GPU card with 768x1536 input images.
Simultaneous Monitoring of Multiple People's Vital Sign Leveraging a Single Phased-MIMO Radar
Oct 15, 2021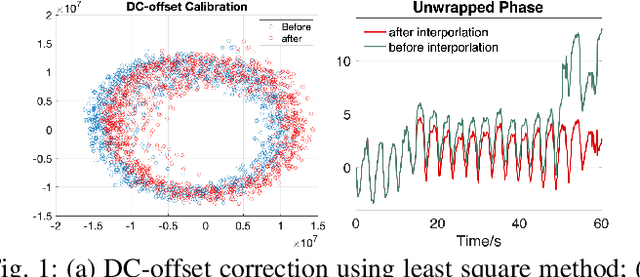
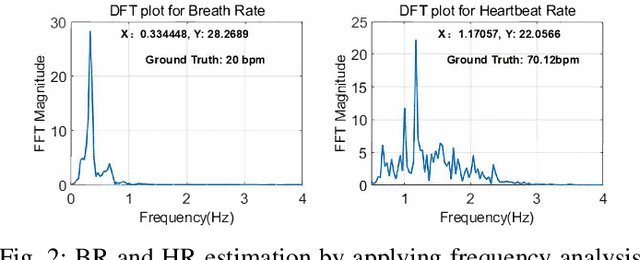
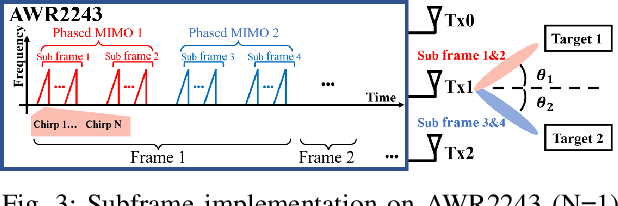
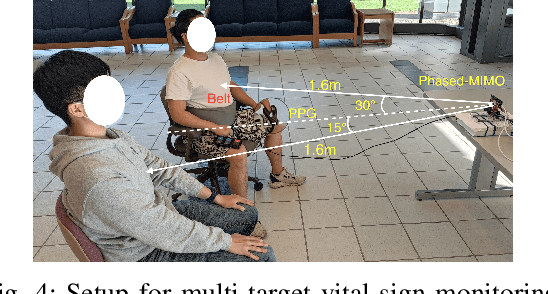
Vital sign monitoring plays a critical role in tracking the physiological state of people and enabling various health-related applications (e.g., recommending a change of lifestyle, examining the risk of diseases). Traditional approaches rely on hospitalization or body-attached instruments, which are costly and intrusive. Therefore, researchers have been exploring contact-less vital sign monitoring with radio frequency signals in recent years. Early studies with continuous wave radars/WiFi devices work on detecting vital signs of a single individual, but it still remains challenging to simultaneously monitor vital signs of multiple subjects, especially those who locate in proximity. In this paper, we design and implement a time-division multiplexing (TDM) phased-MIMO radar sensing scheme for high-precision vital sign monitoring of multiple people. Our phased-MIMO radar can steer the mmWave beam towards different directions with a micro-second delay, which enables capturing the vital signs of multiple individuals at the same radial distance to the radar. Furthermore, we develop a TDM-MIMO technique to fully utilize all transmitting antenna (TX)-receiving antenna (RX) pairs, thereby significantly boosting the signal-to-noise ratio. Based on the designed TDM phased-MIMO radar, we develop a system to automatically localize multiple human subjects and estimate their vital signs. Extensive evaluations show that under two-subject scenarios, our system can achieve an error of less than 1 beat per minute (BPM) and 3 BPM for breathing rate (BR) and heartbeat rate (HR) estimations, respectively, at a subject-to-radar distance of $1.6~m$. The minimal subject-to-subject angle separation is $40{\deg}$, corresponding to a close distance of $0.5~m$ between two subjects, which outperforms the state-of-the-art.
MLMOD Package: Machine Learning Methods for Data-Driven Modeling in LAMMPS
Jul 29, 2021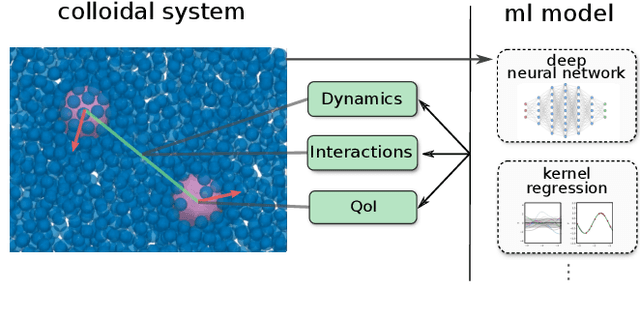
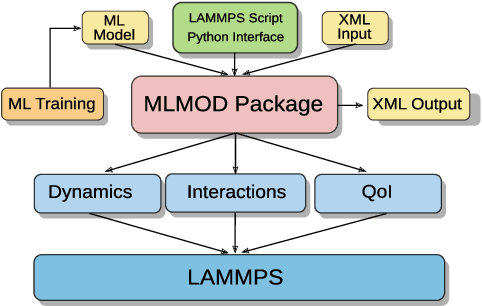
We discuss a software package for incorporating into simulations data-driven models trained using machine learning methods. These can be used for (i) modeling dynamics and time-step integration, (ii) modeling interactions between system components, and (iii) computing quantities of interest characterizing system state. The package allows for use of machine learning methods with general model classes including Neural Networks, Gaussian Process Regression, Kernel Models, and other approaches. We discuss in this whitepaper our prototype C++ package, aims, and example usage.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge