"Time": models, code, and papers
Choosing the Right Algorithm With Hints From Complexity Theory
Sep 14, 2021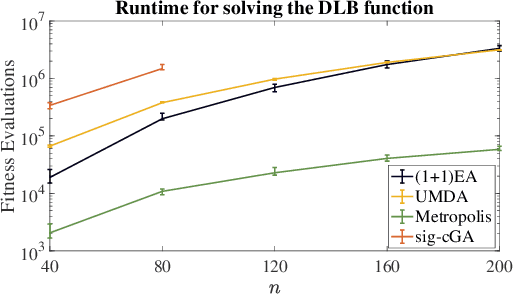
Choosing a suitable algorithm from the myriads of different search heuristics is difficult when faced with a novel optimization problem. In this work, we argue that the purely academic question of what could be the best possible algorithm in a certain broad class of black-box optimizers can give fruitful indications in which direction to search for good established optimization heuristics. We demonstrate this approach on the recently proposed DLB benchmark, for which the only known results are $O(n^3)$ runtimes for several classic evolutionary algorithms and an $O(n^2 \log n)$ runtime for an estimation-of-distribution algorithm. Our finding that the unary unbiased black-box complexity is only $O(n^2)$ suggests the Metropolis algorithm as an interesting candidate and we prove that it solves the DLB problem in quadratic time. Since we also prove that better runtimes cannot be obtained in the class of unary unbiased algorithms, we shift our attention to algorithms that use the information of more parents to generate new solutions. An artificial algorithm of this type having an $O(n \log n)$ runtime leads to the result that the significance-based compact genetic algorithm (sig-cGA) can solve the DLB problem also in time $O(n \log n)$. Our experiments show a remarkably good performance of the Metropolis algorithm, clearly the best of all algorithms regarded for reasonable problem sizes.
Breaking the curse of dimensionality with Isolation Kernel
Sep 29, 2021
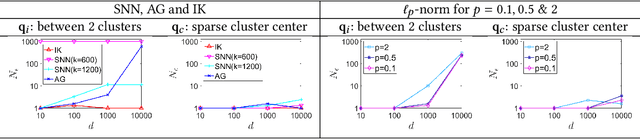
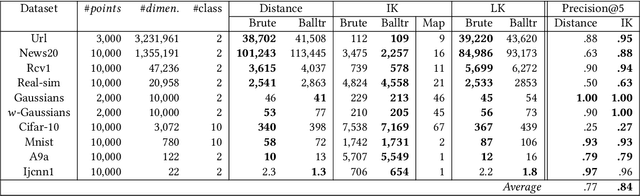
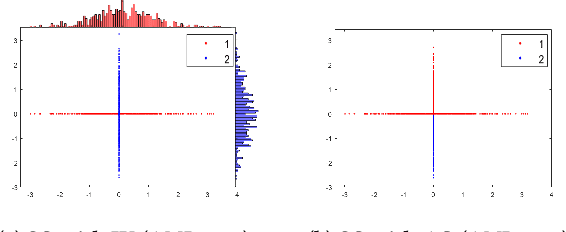
The curse of dimensionality has been studied in different aspects. However, breaking the curse has been elusive. We show for the first time that it is possible to break the curse using the recently introduced Isolation Kernel. We show that only Isolation Kernel performs consistently well in indexed search, spectral & density peaks clustering, SVM classification and t-SNE visualization in both low and high dimensions, compared with distance, Gaussian and linear kernels. This is also supported by our theoretical analyses that Isolation Kernel is the only kernel that has the provable ability to break the curse, compared with existing metric-based Lipschitz continuous kernels.
Medical Instrument Segmentation in 3D US by Hybrid Constrained Semi-Supervised Learning
Jul 30, 2021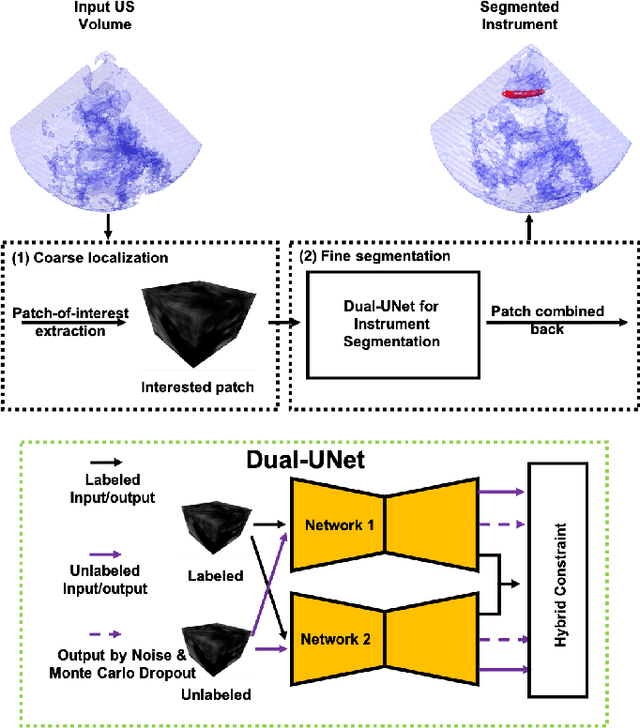
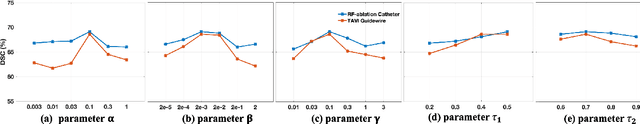
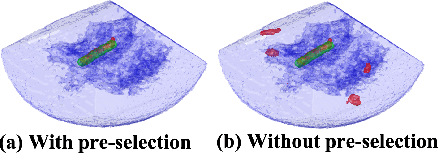
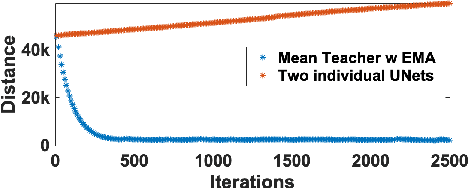
Medical instrument segmentation in 3D ultrasound is essential for image-guided intervention. However, to train a successful deep neural network for instrument segmentation, a large number of labeled images are required, which is expensive and time-consuming to obtain. In this article, we propose a semi-supervised learning (SSL) framework for instrument segmentation in 3D US, which requires much less annotation effort than the existing methods. To achieve the SSL learning, a Dual-UNet is proposed to segment the instrument. The Dual-UNet leverages unlabeled data using a novel hybrid loss function, consisting of uncertainty and contextual constraints. Specifically, the uncertainty constraints leverage the uncertainty estimation of the predictions of the UNet, and therefore improve the unlabeled information for SSL training. In addition, contextual constraints exploit the contextual information of the training images, which are used as the complementary information for voxel-wise uncertainty estimation. Extensive experiments on multiple ex-vivo and in-vivo datasets show that our proposed method achieves Dice score of about 68.6%-69.1% and the inference time of about 1 sec. per volume. These results are better than the state-of-the-art SSL methods and the inference time is comparable to the supervised approaches.
Distributed Learning over a Wireless Network with FSK-Based Majority Vote
Nov 02, 2021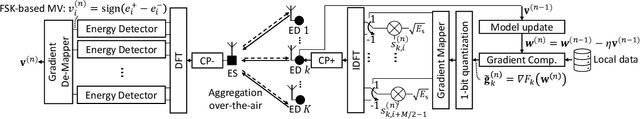
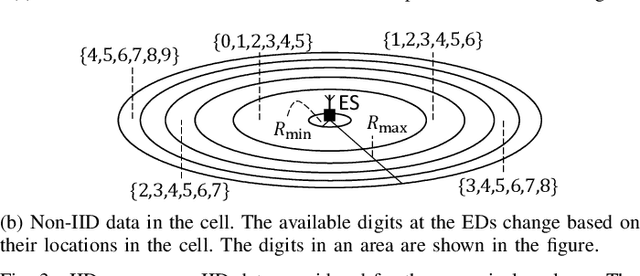
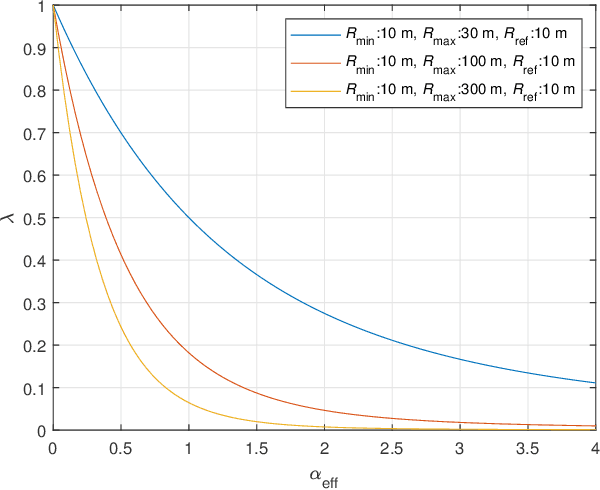

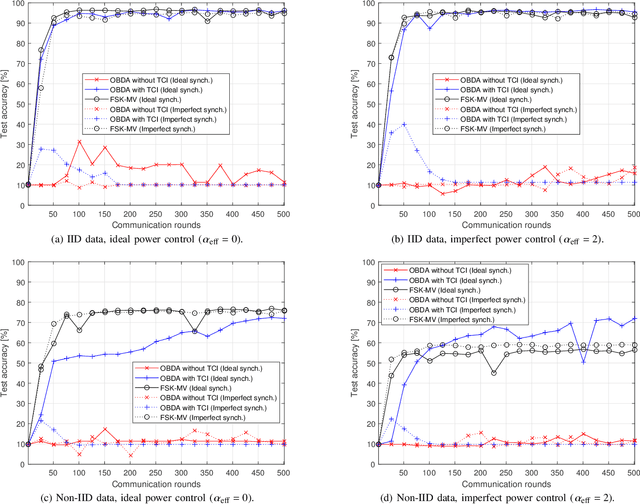
In this study, we propose an over-the-air computation (AirComp) scheme for federated edge learning (FEEL). The proposed scheme relies on the concept of distributed learning by majority vote (MV) with sign stochastic gradient descend (signSGD). As compared to the state-of-the-art solutions, with the proposed method, edge devices (EDs) transmit the signs of local stochastic gradients by activating one of two orthogonal resources, i.e., orthogonal frequency division multiplexing (OFDM) subcarriers, and the MVs at the edge server (ES) are obtained with non-coherent detectors by exploiting the energy accumulations on the subcarriers. Hence, the proposed scheme eliminates the need for channel state information (CSI) at the EDs and ES. By taking path loss, power control, cell size, and the probabilistic nature of the detected MVs in fading channel into account, we prove the convergence of the distributed learning for a non-convex function. Through simulations, we show that the proposed scheme can provide a high test accuracy in fading channels even when the time-synchronization and the power alignment at the ES are not ideal. We also provide insight into distributed learning for location-dependent data distribution for the MV-based schemes.
Computer Model Calibration with Time Series Data using Deep Learning and Quantile Regression
Aug 29, 2020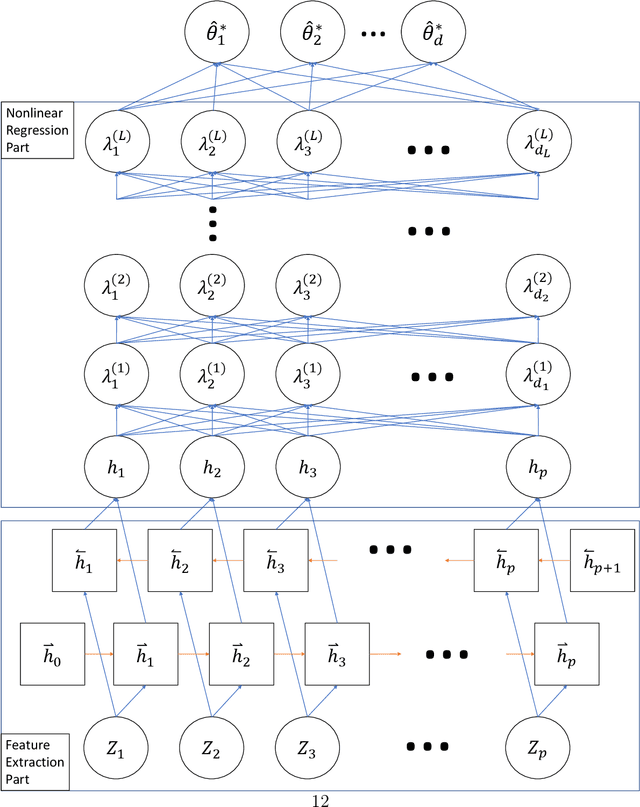
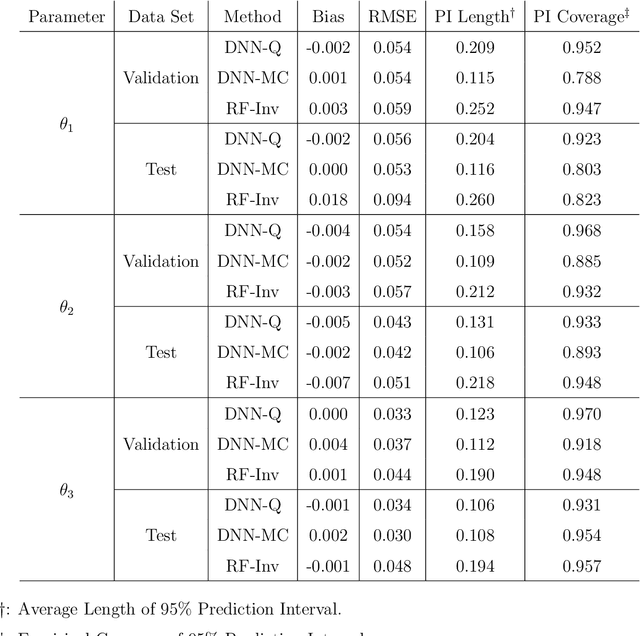
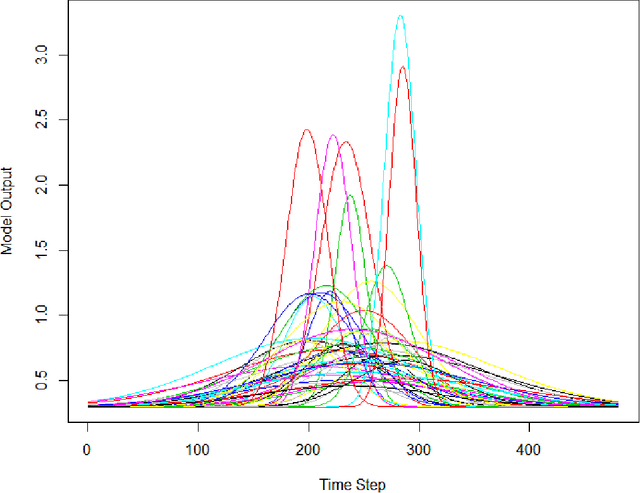
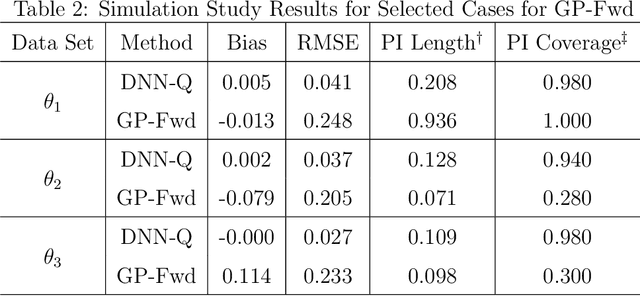
Computer models play a key role in many scientific and engineering problems. One major source of uncertainty in computer model experiment is input parameter uncertainty. Computer model calibration is a formal statistical procedure to infer input parameters by combining information from model runs and observational data. The existing standard calibration framework suffers from inferential issues when the model output and observational data are high-dimensional dependent data such as large time series due to the difficulty in building an emulator and the non-identifiability between effects from input parameters and data-model discrepancy. To overcome these challenges we propose a new calibration framework based on a deep neural network (DNN) with long-short term memory layers that directly emulates the inverse relationship between the model output and input parameters. Adopting the 'learning with noise' idea we train our DNN model to filter out the effects from data model discrepancy on input parameter inference. We also formulate a new way to construct interval predictions for DNN using quantile regression to quantify the uncertainty in input parameter estimates. Through a simulation study and real data application with WRF-hydro model we show that our approach can yield accurate point estimates and well calibrated interval estimates for input parameters.
WORD: Revisiting Organs Segmentation in the Whole Abdominal Region
Nov 17, 2021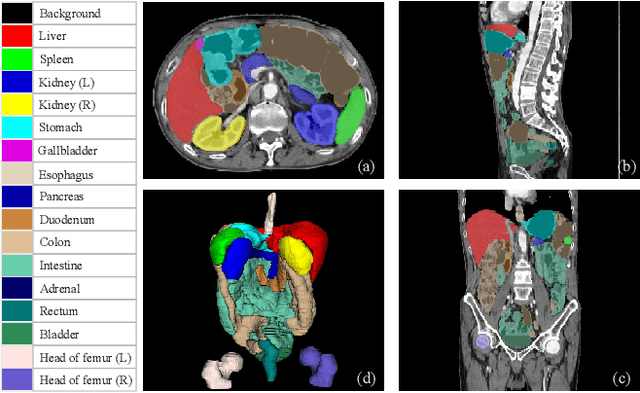
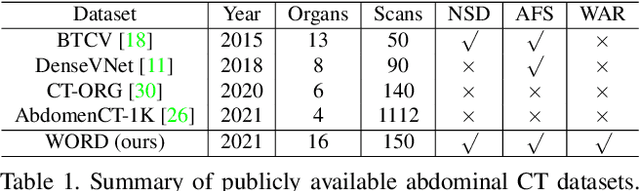
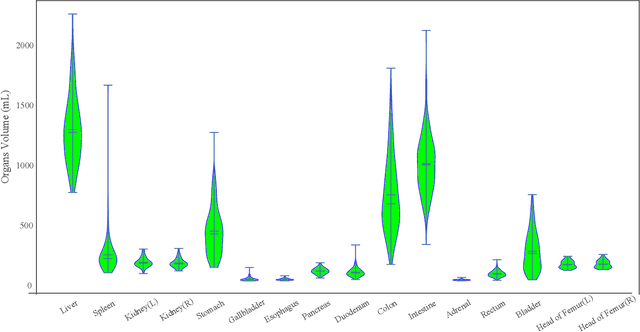
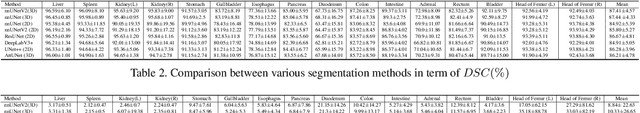
Whole abdominal organs segmentation plays an important role in abdomen lesion diagnosis, radiotherapy planning, and follow-up. However, delineating all abdominal organs by oncologists manually is time-consuming and very expensive. Recently, deep learning-based medical image segmentation has shown the potential to reduce manual delineation efforts, but it still requires a large-scale fine annotated dataset for training. Although many efforts in this task, there are still few large image datasets covering the whole abdomen region with accurate and detailed annotations for the whole abdominal organ segmentation. In this work, we establish a large-scale \textit{W}hole abdominal \textit{OR}gans \textit{D}ataset (\textit{WORD}) for algorithms research and clinical applications development. This dataset contains 150 abdominal CT volumes (30495 slices) and each volume has 16 organs with fine pixel-level annotations and scribble-based sparse annotation, which may be the largest dataset with whole abdominal organs annotation. Several state-of-the-art segmentation methods are evaluated on this dataset. And, we also invited clinical oncologists to revise the model predictions to measure the gap between the deep learning method and real oncologists. We further introduce and evaluate a new scribble-based weakly supervised segmentation on this dataset. The work provided a new benchmark for the abdominal multi-organ segmentation task and these experiments can serve as the baseline for future research and clinical application development. The codebase and dataset will be released at: https://github.com/HiLab-git/WORD
Mapping illegal waste dumping sites with neural-network classification of satellite imagery
Oct 28, 2021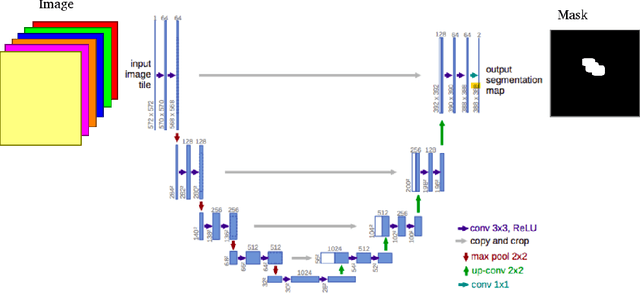

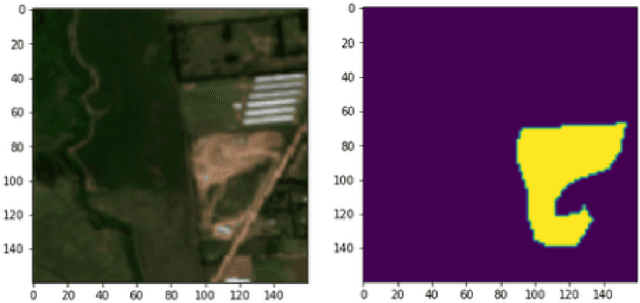
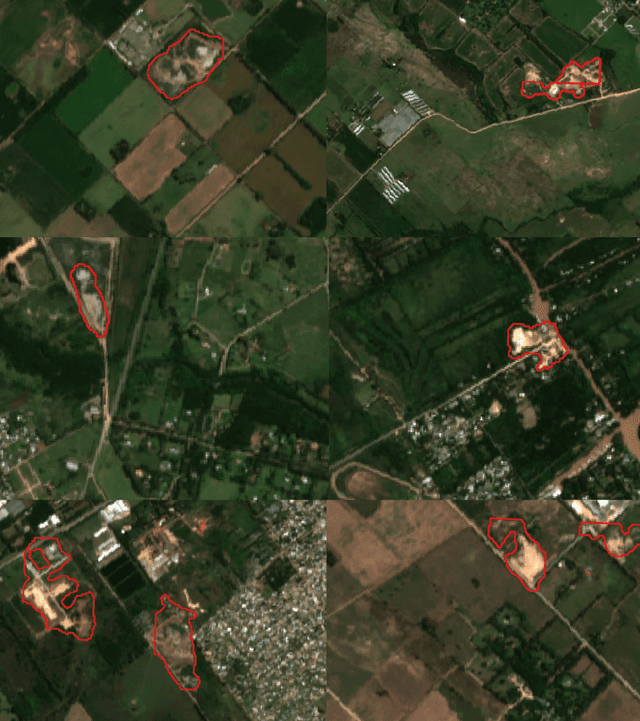
Public health and habitat quality are crucial goals of urban planning. In recent years, the severe social and environmental impact of illegal waste dumping sites has made them one of the most serious problems faced by cities in the Global South, in a context of scarce information available for decision making. To help identify the location of dumping sites and track their evolution over time we adopt a data-driven model from the machine learning domain, analyzing satellite images. This allows us to take advantage of the increasing availability of geo-spatial open-data, high-resolution satellite imagery, and open source tools to train machine learning algorithms with a small set of known waste dumping sites in Buenos Aires, and then predict the location of other sites over vast areas at high speed and low cost. This case study shows the results of a collaboration between Dymaxion Labs and Fundaci\'on Bunge y Born to harness this technique in order to create a comprehensive map of potential locations of illegal waste dumping sites in the region.
Generalizing electrocardiogram delineation: training convolutional neural networks with synthetic data augmentation
Nov 25, 2021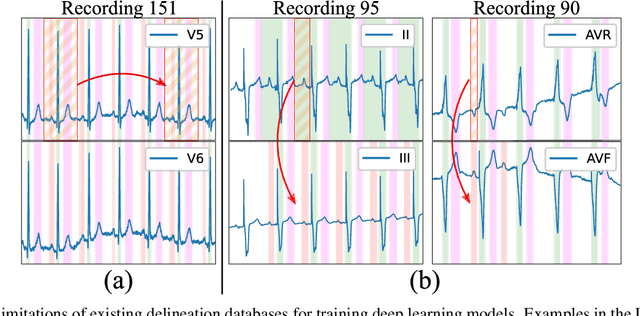
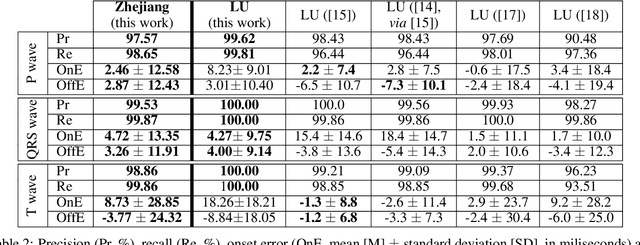
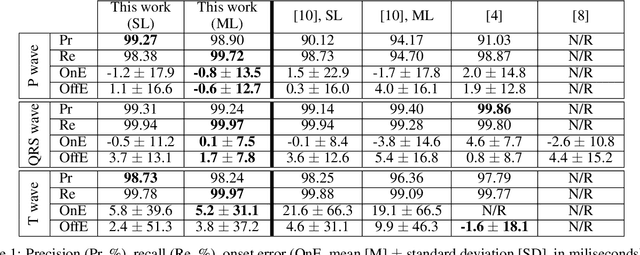
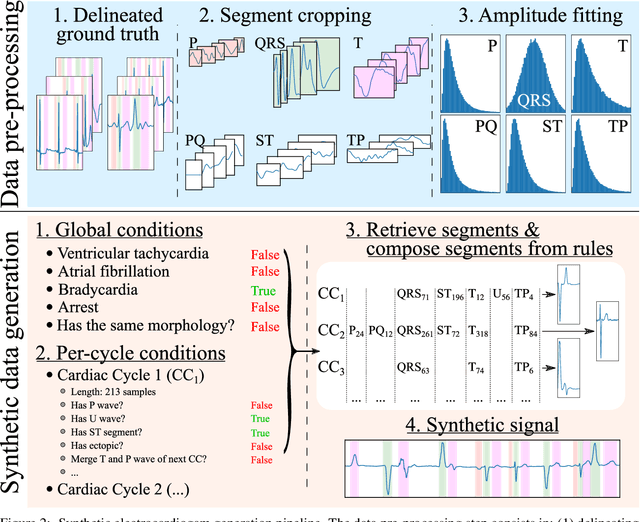

Obtaining per-beat information is a key task in the analysis of cardiac electrocardiograms (ECG), as many downstream diagnosis tasks are dependent on ECG-based measurements. Those measurements, however, are costly to produce, especially in recordings that change throughout long periods of time. However, existing annotated databases for ECG delineation are small, being insufficient in size and in the array of pathological conditions they represent. This article delves has two main contributions. First, a pseudo-synthetic data generation algorithm was developed, based in probabilistically composing ECG traces given "pools" of fundamental segments, as cropped from the original databases, and a set of rules for their arrangement into coherent synthetic traces. The generation of conditions is controlled by imposing expert knowledge on the generated trace, which increases the input variability for training the model. Second, two novel segmentation-based loss functions have been developed, which attempt at enforcing the prediction of an exact number of independent structures and at producing closer segmentation boundaries by focusing on a reduced number of samples. The best performing model obtained an $F_1$-score of 99.38\% and a delineation error of $2.19 \pm 17.73$ ms and $4.45 \pm 18.32$ ms for all wave's fiducials (onsets and offsets, respectively), as averaged across the P, QRS and T waves for three distinct freely available databases. The excellent results were obtained despite the heterogeneous characteristics of the tested databases, in terms of lead configurations (Holter, 12-lead), sampling frequencies ($250$, $500$ and $2,000$ Hz) and represented pathophysiologies (e.g., different types of arrhythmias, sinus rhythm with structural heart disease), hinting at its generalization capabilities, while outperforming current state-of-the-art delineation approaches.
On the Linguistic Capacity of Real-Time Counter Automata
Apr 15, 2020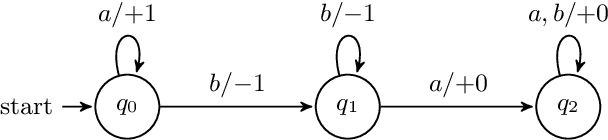
Counter machines have achieved a newfound relevance to the field of natural language processing (NLP): recent work suggests some strong-performing recurrent neural networks utilize their memory as counters. Thus, one potential way to understand the success of these networks is to revisit the theory of counter computation. Therefore, we study the abilities of real-time counter machines as formal grammars, focusing on formal properties that are relevant for NLP models. We first show that several variants of the counter machine converge to express the same class of formal languages. We also prove that counter languages are closed under complement, union, intersection, and many other common set operations. Next, we show that counter machines cannot evaluate boolean expressions, even though they can weakly validate their syntax. This has implications for the interpretability and evaluation of neural network systems: successfully matching syntactic patterns does not guarantee that counter memory accurately encodes compositional semantics. Finally, we consider whether counter languages are semilinear. This work makes general contributions to the theory of formal languages that are of potential interest for understanding recurrent neural networks.
Improved Feature Importance Computations for Tree Models: Shapley vs. Banzhaf
Aug 09, 2021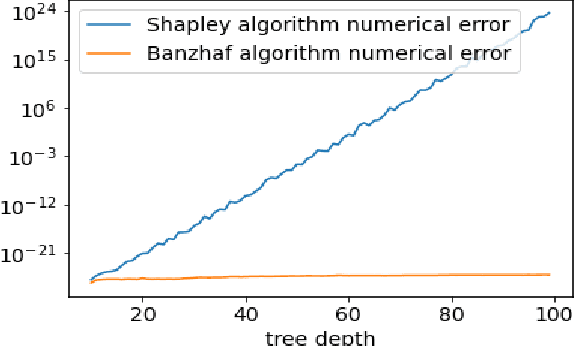
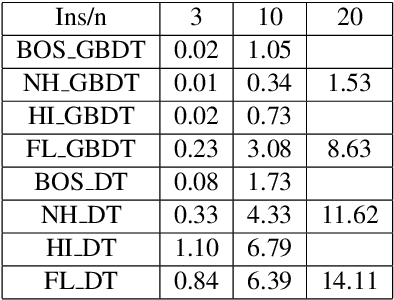
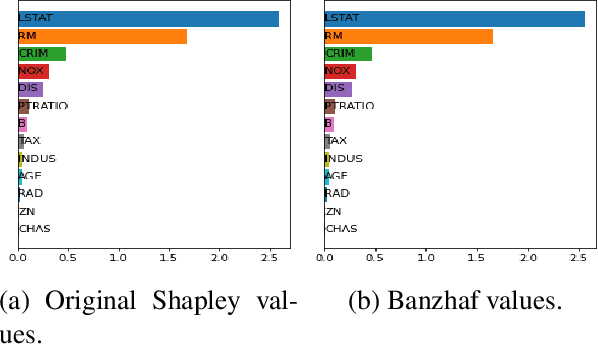
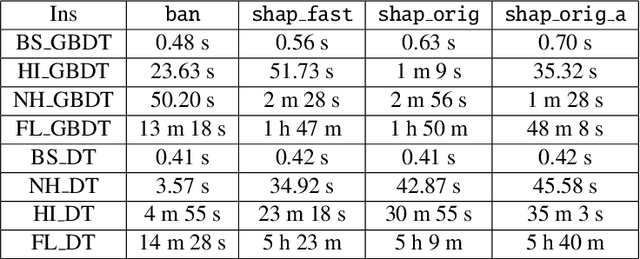
Shapley values are one of the main tools used to explain predictions of tree ensemble models. The main alternative to Shapley values are Banzhaf values that have not been understood equally well. In this paper we make a step towards filling this gap, providing both experimental and theoretical comparison of these model explanation methods. Surprisingly, we show that Banzhaf values offer several advantages over Shapley values while providing essentially the same explanations. We verify that Banzhaf values: (1) have a more intuitive interpretation, (2) allow for more efficient algorithms, and (3) are much more numerically robust. We provide an experimental evaluation of these theses. In particular, we show that on real world instances. Additionally, from a theoretical perspective we provide new and improved algorithm computing the same Shapley value based explanations as the algorithm of Lundberg et al. [Nat. Mach. Intell. 2020]. Our algorithm runs in $O(TLD+n)$ time, whereas the previous algorithm had $O(TLD^2+n)$ running time bound. Here, $T$ is the number of trees, $L$ is the maximum number of leaves in a tree, and $D$ denotes the maximum depth of a tree in the ensemble. Using the computational techniques developed for Shapley values we deliver an optimal $O(TL+n)$ time algorithm for computing Banzhaf values based explanations. In our experiments these algorithms give running times smaller even by an order of magnitude.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge