"Time": models, code, and papers
Weakly-supervised learning for image-based classification of primary melanomas into genomic immune subgroups
Feb 23, 2022

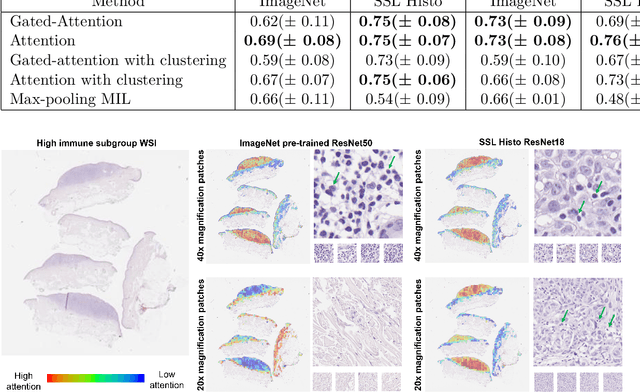
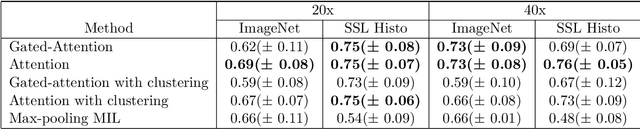
Determining early-stage prognostic markers and stratifying patients for effective treatment are two key challenges for improving outcomes for melanoma patients. Previous studies have used tumour transcriptome data to stratify patients into immune subgroups, which were associated with differential melanoma specific survival and potential treatment strategies. However, acquiring transcriptome data is a time-consuming and costly process. Moreover, it is not routinely used in the current clinical workflow. Here we attempt to overcome this by developing deep learning models to classify gigapixel H&E stained pathology slides, which are well established in clinical workflows, into these immune subgroups. Previous subtyping approaches have employed supervised learning which requires fully annotated data, or have only examined single genetic mutations in melanoma patients. We leverage a multiple-instance learning approach, which only requires slide-level labels and uses an attention mechanism to highlight regions of high importance to the classification. Moreover, we show that pathology-specific self-supervised models generate better representations compared to pathology-agnostic models for improving our model performance, achieving a mean AUC of 0.76 for classifying histopathology images as high or low immune subgroups. We anticipate that this method may allow us to find new biomarkers of high importance and could act as a tool for clinicians to infer the immune landscape of tumours and stratify patients, without needing to carry out additional expensive genetic tests.
A Lightweight Dual-Domain Attention Framework for Sparse-View CT Reconstruction
Feb 19, 2022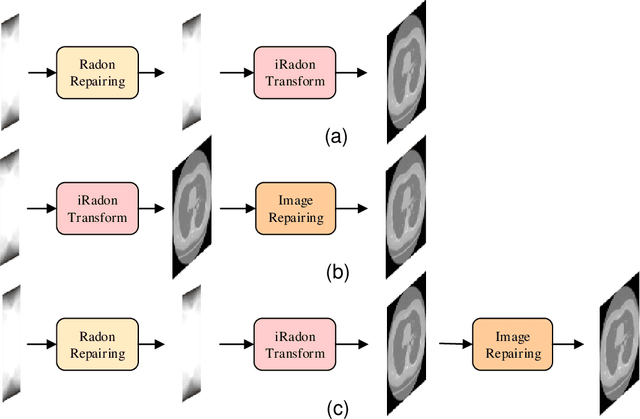
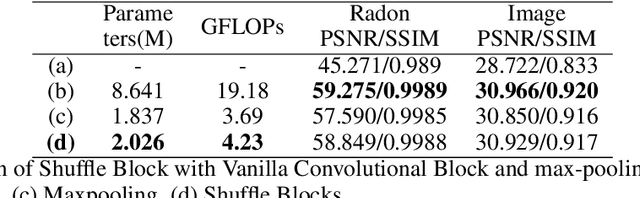

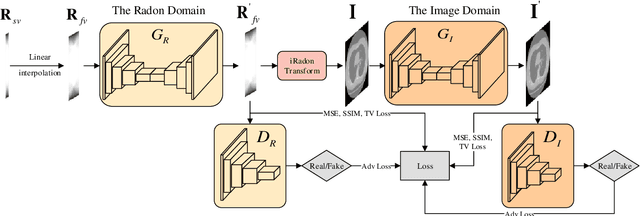

Computed Tomography (CT) plays an essential role in clinical diagnosis. Due to the adverse effects of radiation on patients, the radiation dose is expected to be reduced as low as possible. Sparse sampling is an effective way, but it will lead to severe artifacts on the reconstructed CT image, thus sparse-view CT image reconstruction has been a prevailing and challenging research area. With the popularity of mobile devices, the requirements for lightweight and real-time networks are increasing rapidly. In this paper, we design a novel lightweight network called CAGAN, and propose a dual-domain reconstruction pipeline for parallel beam sparse-view CT. CAGAN is an adversarial auto-encoder, combining the Coordinate Attention unit, which preserves the spatial information of features. Also, the application of Shuffle Blocks reduces the parameters by a quarter without sacrificing its performance. In the Radon domain, the CAGAN learns the mapping between the interpolated data and fringe-free projection data. After the restored Radon data is reconstructed to an image, the image is sent into the second CAGAN trained for recovering the details, so that a high-quality image is obtained. Experiments indicate that the CAGAN strikes an excellent balance between model complexity and performance, and our pipeline outperforms the DD-Net and the DuDoNet.
A Novel Self-Supervised Cross-Modal Image Retrieval Method In Remote Sensing
Feb 23, 2022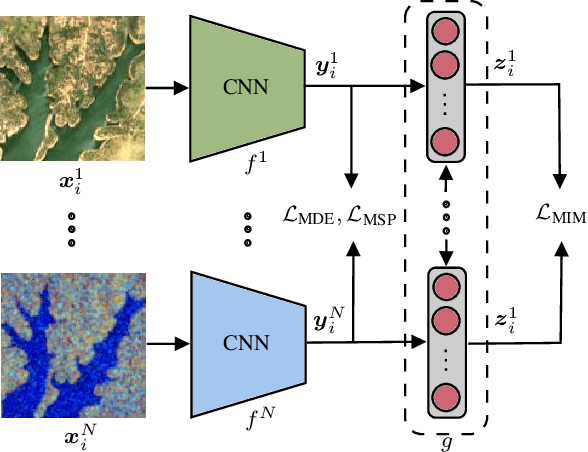
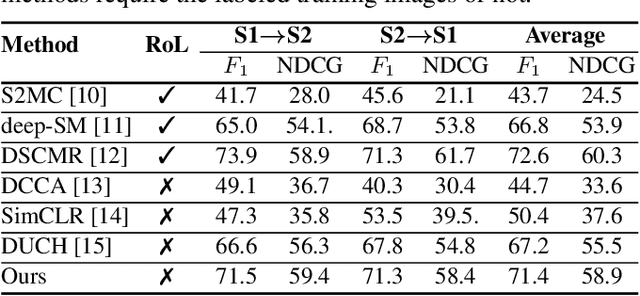

Due to the availability of multi-modal remote sensing (RS) image archives, one of the most important research topics is the development of cross-modal RS image retrieval (CM-RSIR) methods that search semantically similar images across different modalities. Existing CM-RSIR methods require annotated training images (which is time-consuming, costly and not feasible to gather in large-scale applications) and do not concurrently address intra- and inter-modal similarity preservation and inter-modal discrepancy elimination. In this paper, we introduce a novel self-supervised cross-modal image retrieval method that aims to: i) model mutual-information between different modalities in a self-supervised manner; ii) retain the distributions of modal-specific feature spaces similar; and iii) define most similar images within each modality without requiring any annotated training images. To this end, we propose a novel objective including three loss functions that simultaneously: i) maximize mutual information of different modalities for inter-modal similarity preservation; ii) minimize the angular distance of multi-modal image tuples for the elimination of inter-modal discrepancies; and iii) increase cosine similarity of most similar images within each modality for the characterization of intra-modal similarities. Experimental results show the effectiveness of the proposed method compared to state-of-the-art methods. The code of the proposed method is publicly available at https://git.tu-berlin.de/rsim/SS-CM-RSIR.
Time Series Data Augmentation for Deep Learning: A Survey
Feb 27, 2020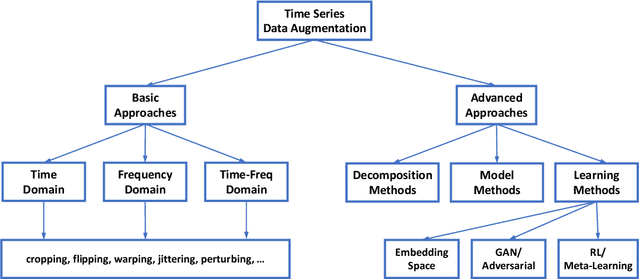
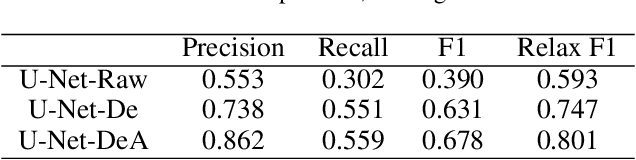
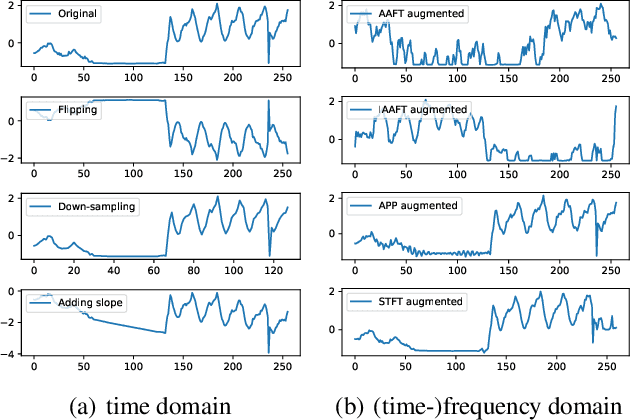
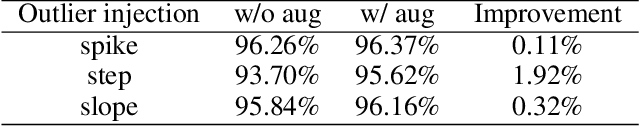
Deep learning performs remarkably well on many time series analysis tasks recently. The superior performance of deep neural networks relies heavily on a large number of training data to avoid overfitting. However, the labeled data of many real-world time series applications may be limited such as classification in medical time series and anomaly detection in AIOps. As an effective way to enhance the size and quality of the training data, data augmentation is crucial to the successful application of deep learning models on time series data. In this paper, we systematically review different data augmentation methods for time series. We propose a taxonomy for the reviewed methods, and then provide a structured review for these methods by highlighting their strengths and limitations. We also empirically compare different data augmentation methods for different tasks including time series anomaly detection, classification and forecasting. Finally, we discuss and highlight future research directions, including data augmentation in time-frequency domain, augmentation combination, and data augmentation and weighting for imbalanced class.
Robustly-reliable learners under poisoning attacks
Mar 08, 2022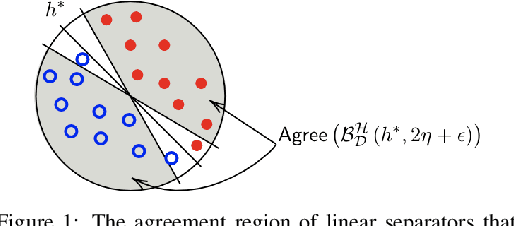
Data poisoning attacks, in which an adversary corrupts a training set with the goal of inducing specific desired mistakes, have raised substantial concern: even just the possibility of such an attack can make a user no longer trust the results of a learning system. In this work, we show how to achieve strong robustness guarantees in the face of such attacks across multiple axes. We provide robustly-reliable predictions, in which the predicted label is guaranteed to be correct so long as the adversary has not exceeded a given corruption budget, even in the presence of instance targeted attacks, where the adversary knows the test example in advance and aims to cause a specific failure on that example. Our guarantees are substantially stronger than those in prior approaches, which were only able to provide certificates that the prediction of the learning algorithm does not change, as opposed to certifying that the prediction is correct, as we are able to achieve in our work. Remarkably, we provide a complete characterization of learnability in this setting, in particular, nearly-tight matching upper and lower bounds on the region that can be certified, as well as efficient algorithms for computing this region given an ERM oracle. Moreover, for the case of linear separators over logconcave distributions, we provide efficient truly polynomial time algorithms (i.e., non-oracle algorithms) for such robustly-reliable predictions. We also extend these results to the active setting where the algorithm adaptively asks for labels of specific informative examples, and the difficulty is that the adversary might even be adaptive to this interaction, as well as to the agnostic learning setting where there is no perfect classifier even over the uncorrupted data.
Mixed-Block Neural Architecture Search for Medical Image Segmentation
Feb 23, 2022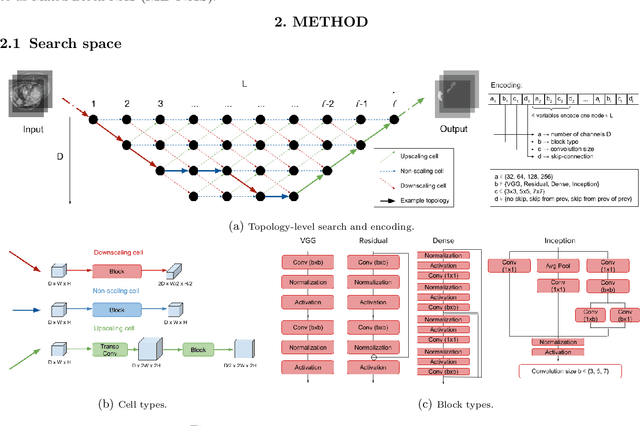
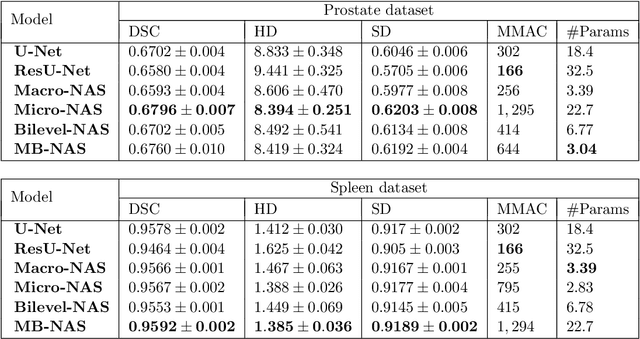
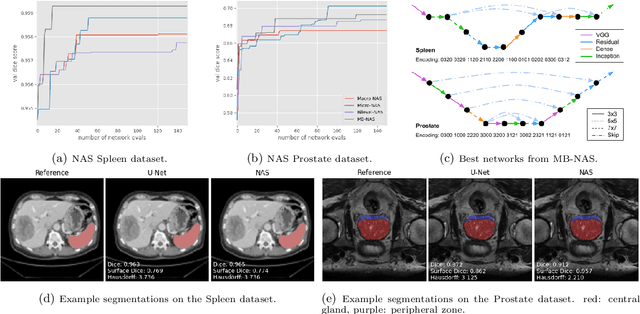
Deep Neural Networks (DNNs) have the potential for making various clinical procedures more time-efficient by automating medical image segmentation. Due to their strong, in some cases human-level, performance, they have become the standard approach in this field. The design of the best possible medical image segmentation DNNs, however, is task-specific. Neural Architecture Search (NAS), i.e., the automation of neural network design, has been shown to have the capability to outperform manually designed networks for various tasks. However, the existing NAS methods for medical image segmentation have explored a quite limited range of types of DNN architectures that can be discovered. In this work, we propose a novel NAS search space for medical image segmentation networks. This search space combines the strength of a generalised encoder-decoder structure, well known from U-Net, with network blocks that have proven to have a strong performance in image classification tasks. The search is performed by looking for the best topology of multiple cells simultaneously with the configuration of each cell within, allowing for interactions between topology and cell-level attributes. From experiments on two publicly available datasets, we find that the networks discovered by our proposed NAS method have better performance than well-known handcrafted segmentation networks, and outperform networks found with other NAS approaches that perform only topology search, and topology-level search followed by cell-level search.
COIN++: Data Agnostic Neural Compression
Jan 30, 2022
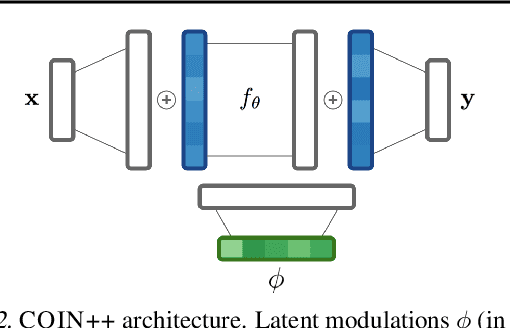
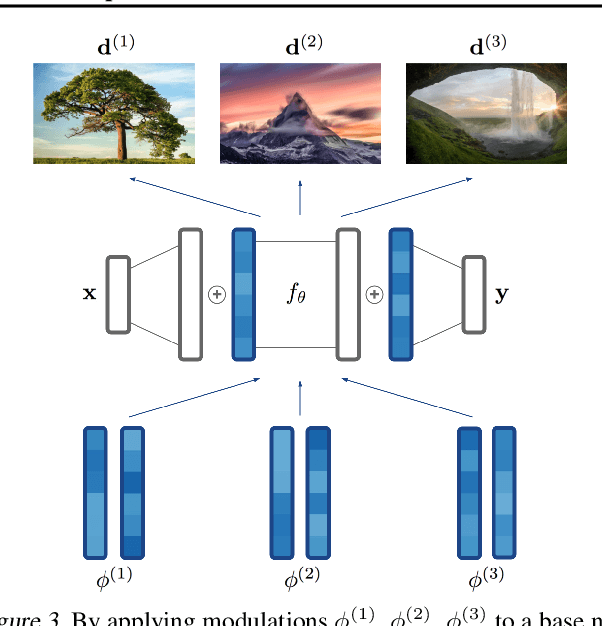
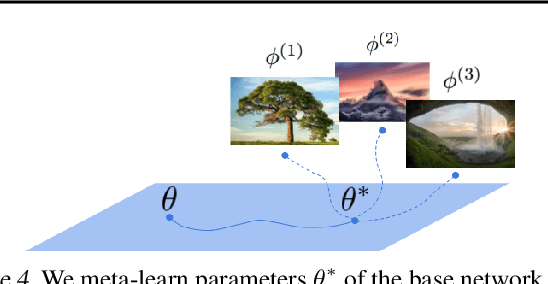
Neural compression algorithms are typically based on autoencoders that require specialized encoder and decoder architectures for different data modalities. In this paper, we propose COIN++, a neural compression framework that seamlessly handles a wide range of data modalities. Our approach is based on converting data to implicit neural representations, i.e. neural functions that map coordinates (such as pixel locations) to features (such as RGB values). Then, instead of storing the weights of the implicit neural representation directly, we store modulations applied to a meta-learned base network as a compressed code for the data. We further quantize and entropy code these modulations, leading to large compression gains while reducing encoding time by two orders of magnitude compared to baselines. We empirically demonstrate the effectiveness of our method by compressing various data modalities, from images to medical and climate data.
FamilySeer: Towards Optimized Tensor Codes by Exploiting Computation Subgraph Similarity
Jan 01, 2022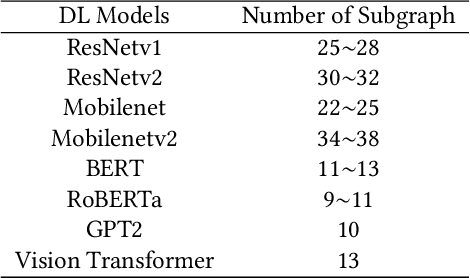
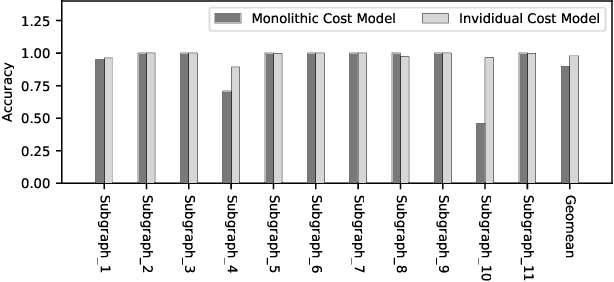
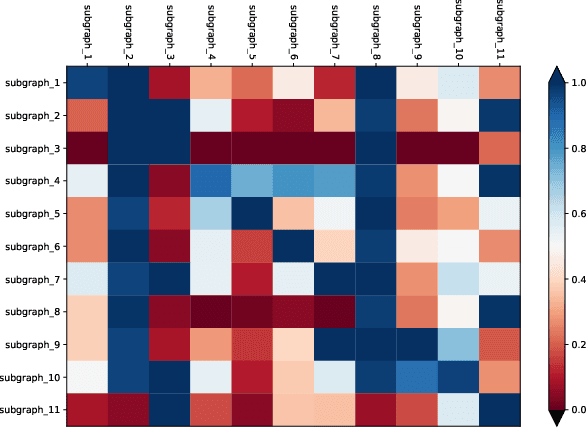

Deploying various deep learning (DL) models efficiently has boosted the research on DL compilers. The difficulty of generating optimized tensor codes drives DL compiler to ask for the auto-tuning approaches, and the increasing demands require increasing auto-tuning efficiency and quality. Currently, the DL compilers partition the input DL models into several subgraphs and leverage the auto-tuning to find the optimal tensor codes of these subgraphs. However, existing auto-tuning approaches usually regard subgraphs as individual ones and overlook the similarities across them, and thus fail to exploit better tensor codes under limited time budgets. We propose FamilySeer, an auto-tuning framework for DL compilers that can generate better tensor codes even with limited time budgets. FamilySeer exploits the similarities and differences among subgraphs can organize them into subgraph families, where the tuning of one subgraph can also improve other subgraphs within the same family. The cost model of each family gets more purified training samples generated by the family and becomes more accurate so that the costly measurements on real hardware can be replaced with the lightweight estimation through cost model. Our experiments show that FamilySeer can generate model codes with the same code performance more efficiently than state-of-the-art auto-tuning frameworks.
Robot Learning of Mobile Manipulation with Reachability Behavior Priors
Mar 08, 2022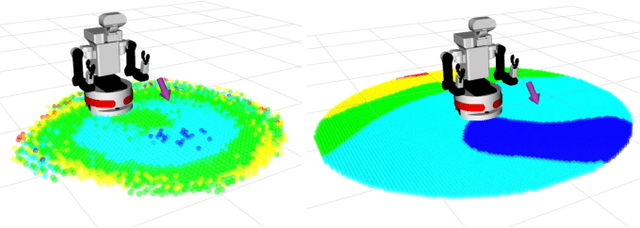
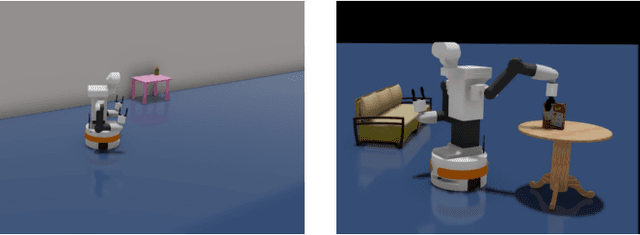
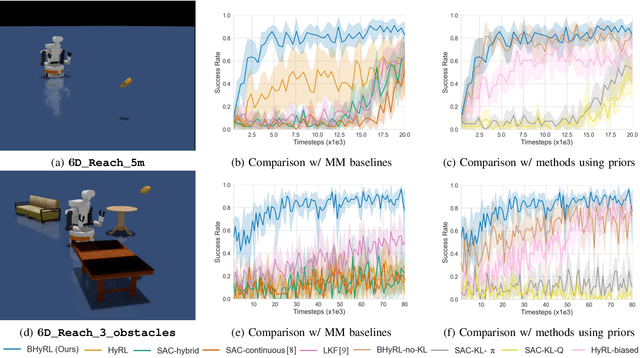

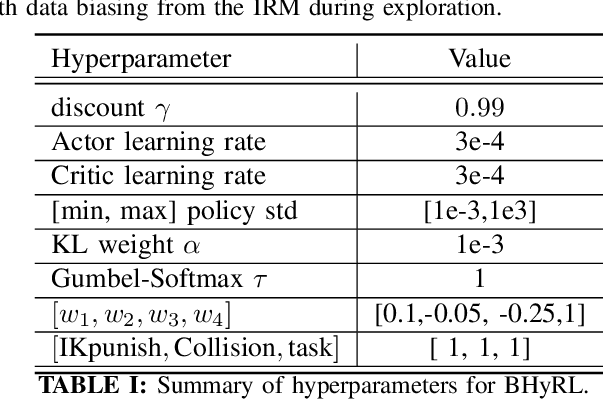
Mobile Manipulation (MM) systems are ideal candidates for taking up the role of a personal assistant in unstructured real-world environments. Among other challenges, MM requires effective coordination of the robot's embodiments for executing tasks that require both mobility and manipulation. Reinforcement Learning (RL) holds the promise of endowing robots with adaptive behaviors, but most methods require prohibitively large amounts of data for learning a useful control policy. In this work, we study the integration of robotic reachability priors in actor-critic RL methods for accelerating the learning of MM for reaching and fetching tasks. Namely, we consider the problem of optimal base placement and the subsequent decision of whether to activate the arm for reaching a 6D target. For this, we devise a novel Hybrid RL method that handles discrete and continuous actions jointly, resorting to the Gumbel-Softmax reparameterization. Next, we train a reachability prior using data from the operational robot workspace, inspired by classical methods. Subsequently, we derive Boosted Hybrid RL (BHyRL), a novel algorithm for learning Q-functions by modeling them as a sum of residual approximators. Every time a new task needs to be learned, we can transfer our learned residuals and learn the component of the Q-function that is task-specific, hence, maintaining the task structure from prior behaviors. Moreover, we find that regularizing the target policy with a prior policy yields more expressive behaviors. We evaluate our method in simulation in reaching and fetching tasks of increasing difficulty, and we show the superior performance of BHyRL against baseline methods. Finally, we zero-transfer our learned 6D fetching policy with BHyRL to our MM robot TIAGo++. For more details and code release, please refer to our project site: irosalab.com/rlmmbp
Parameter estimation for WMTI-Watson model of white matter using encoder-decoder recurrent neural network
Mar 02, 2022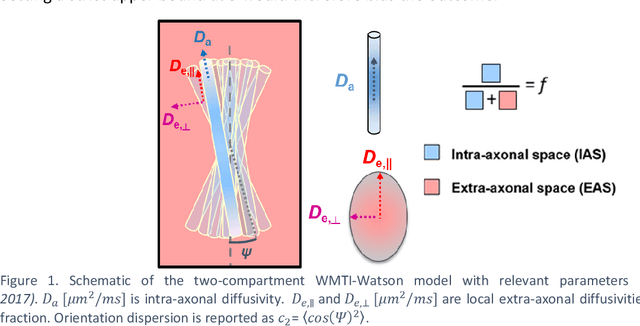

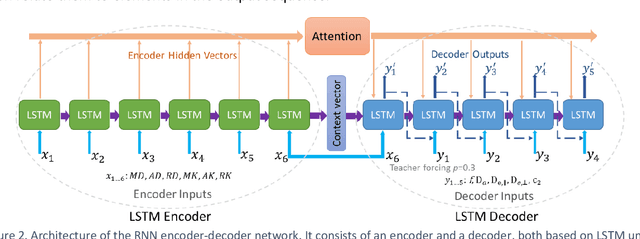
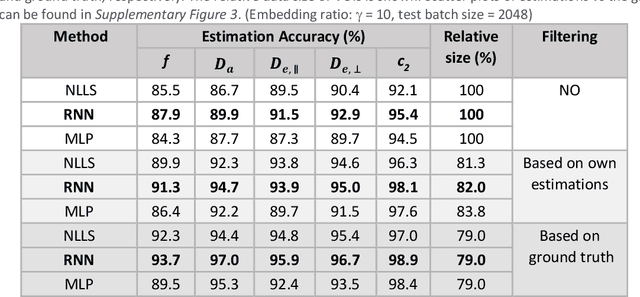
Biophysical modelling of the diffusion MRI signal provides estimates of specific microstructural tissue properties. Although nonlinear optimization such as non-linear least squares (NLLS) is the most widespread method for model estimation, it suffers from local minima and high computational cost. Deep Learning approaches are steadily replacing NL fitting, but come with the limitation that the model needs to be retrained for each acquisition protocol and noise level. The White Matter Tract Integrity (WMTI)-Watson model was proposed as an implementation of the Standard Model of diffusion in white matter that estimates model parameters from the diffusion and kurtosis tensors (DKI). Here we proposed a deep learning approach based on the encoder-decoder recurrent neural network (RNN) to increase the robustness and accelerate the parameter estimation of WMTI-Watson. We use an embedding approach to render the model insensitive to potential differences in distributions between training data and experimental data. This RNN-based solver thus has the advantage of being highly efficient in computation and more readily translatable to other datasets, irrespective of acquisition protocol and underlying parameter distributions as long as DKI was pre-computed from the data. In this study, we evaluated the performance of NLLS, the RNN-based method and a multilayer perceptron (MLP) on synthetic and in vivo datasets of rat and human brain. We showed that the proposed RNN-based fitting approach had the advantage of highly reduced computation time over NLLS (from hours to seconds), with similar accuracy and precision but improved robustness, and superior translatability to new datasets over MLP.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge