"Time": models, code, and papers
Towards a Comprehensive Solution for a Vision-based Digitized Neurological Examination
May 15, 2022
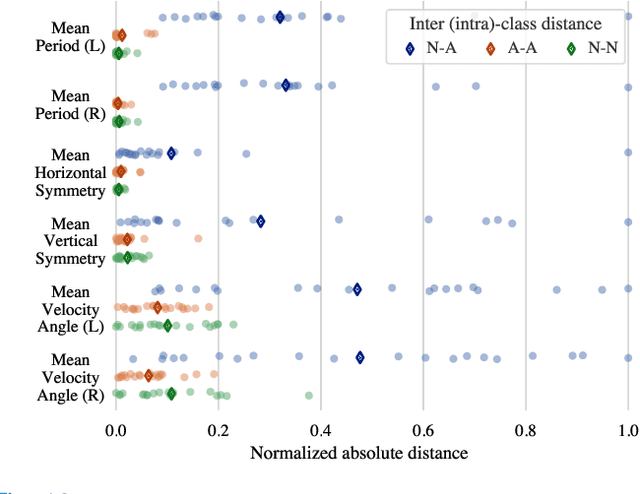
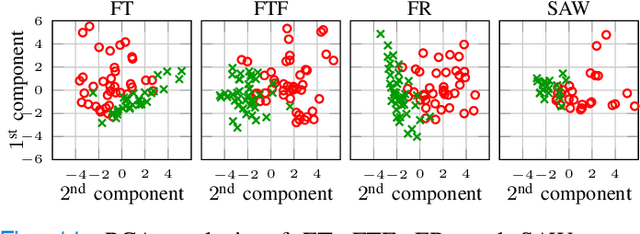
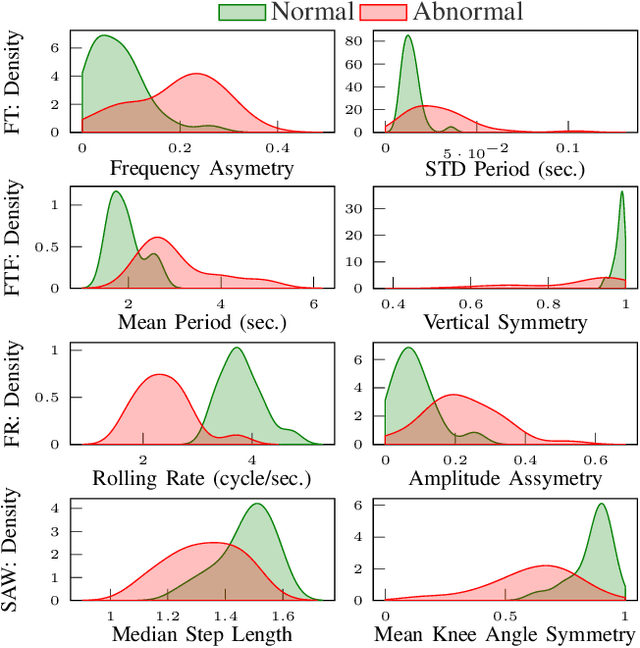
The ability to use digitally recorded and quantified neurological exam information is important to help healthcare systems deliver better care, in-person and via telehealth, as they compensate for a growing shortage of neurologists. Current neurological digital biomarker pipelines, however, are narrowed down to a specific neurological exam component or applied for assessing specific conditions. In this paper, we propose an accessible vision-based exam and documentation solution called Digitized Neurological Examination (DNE) to expand exam biomarker recording options and clinical applications using a smartphone/tablet. Through our DNE software, healthcare providers in clinical settings and people at home are enabled to video capture an examination while performing instructed neurological tests, including finger tapping, finger to finger, forearm roll, and stand-up and walk. Our modular design of the DNE software supports integrations of additional tests. The DNE extracts from the recorded examinations the 2D/3D human-body pose and quantifies kinematic and spatio-temporal features. The features are clinically relevant and allow clinicians to document and observe the quantified movements and the changes of these metrics over time. A web server and a user interface for recordings viewing and feature visualizations are available. DNE was evaluated on a collected dataset of 21 subjects containing normal and simulated-impaired movements. The overall accuracy of DNE is demonstrated by classifying the recorded movements using various machine learning models. Our tests show an accuracy beyond 90% for upper-limb tests and 80% for the stand-up and walk tests.
Understanding A Class of Decentralized and Federated Optimization Algorithms: A Multi-Rate Feedback Control Perspective
Apr 27, 2022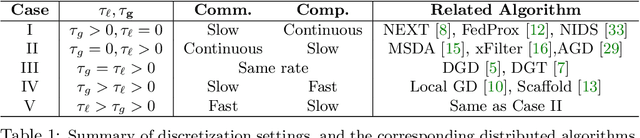
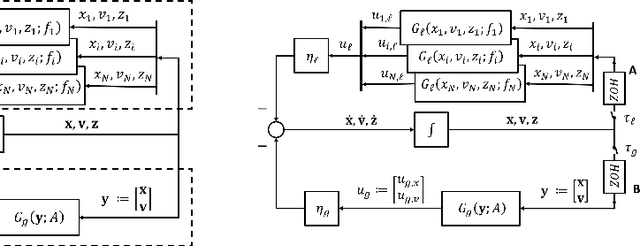
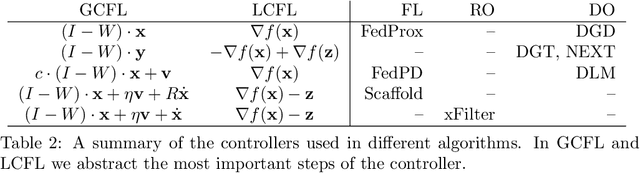
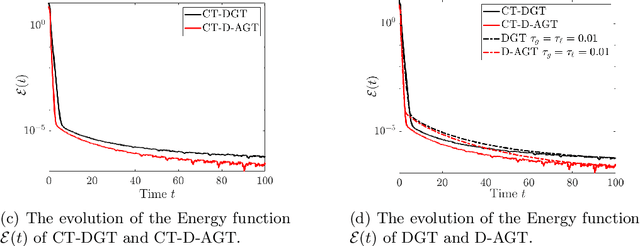
Distributed algorithms have been playing an increasingly important role in many applications such as machine learning, signal processing, and control. Significant research efforts have been devoted to developing and analyzing new algorithms for various applications. In this work, we provide a fresh perspective to understand, analyze, and design distributed optimization algorithms. Through the lens of multi-rate feedback control, we show that a wide class of distributed algorithms, including popular decentralized/federated schemes, can be viewed as discretizing a certain continuous-time feedback control system, possibly with multiple sampling rates, such as decentralized gradient descent, gradient tracking, and federated averaging. This key observation not only allows us to develop a generic framework to analyze the convergence of the entire algorithm class. More importantly, it also leads to an interesting way of designing new distributed algorithms. We develop the theory behind our framework and provide examples to highlight how the framework can be used in practice.
Path sampling of recurrent neural networks by incorporating known physics
Mar 01, 2022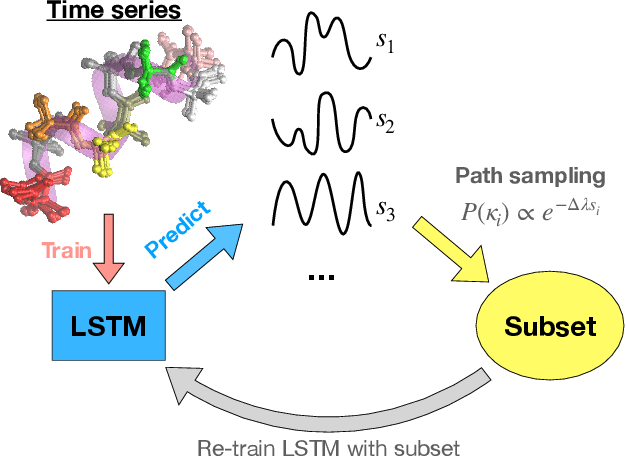
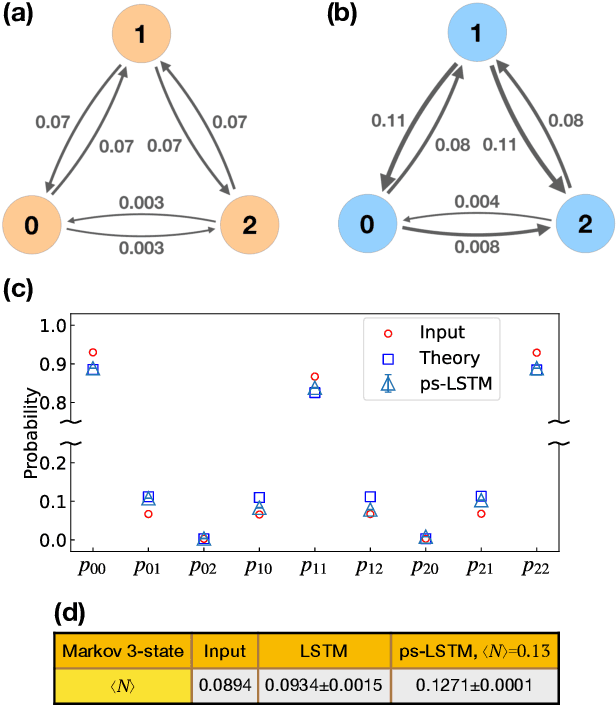
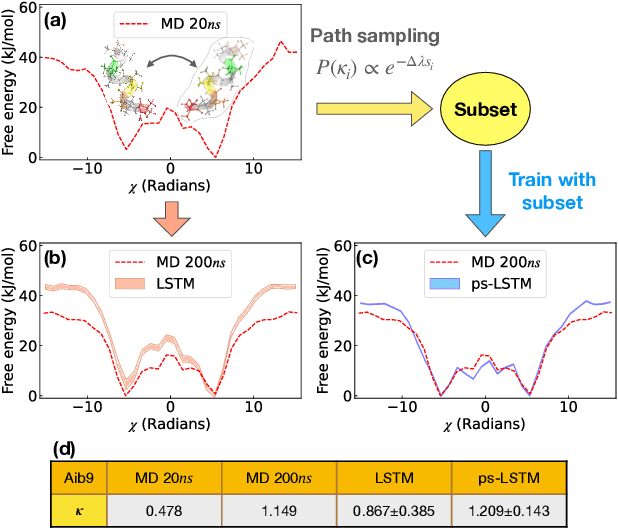
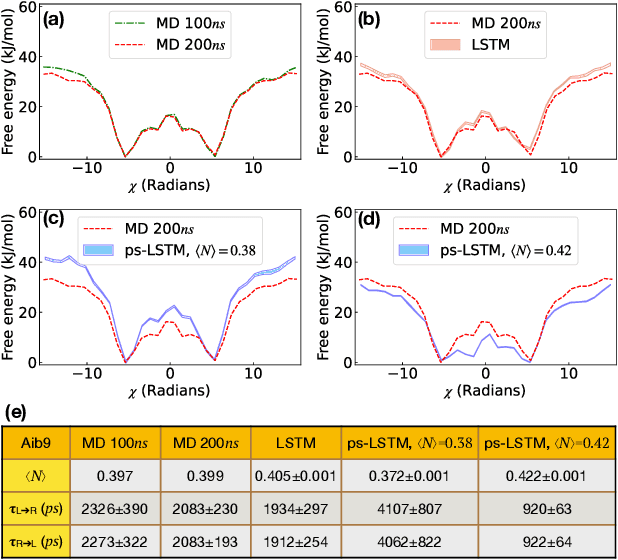
Recurrent neural networks have seen widespread use in modeling dynamical systems in varied domains such as weather prediction, text prediction and several others. Often one wishes to supplement the experimentally observed dynamics with prior knowledge or intuition about the system. While the recurrent nature of these networks allows them to model arbitrarily long memories in the time series used in training, it makes it harder to impose prior knowledge or intuition through generic constraints. In this work, we present a path sampling approach based on principle of Maximum Caliber that allows us to include generic thermodynamic or kinetic constraints into recurrent neural networks. We show the method here for a widely used type of recurrent neural network known as long short-term memory network in the context of supplementing time series collecting from all-atom molecular dynamics. We demonstrate the power of the formalism for different applications. Our method can be easily generalized to other generative artificial intelligence models and to generic time series in different areas of physical and social sciences, where one wishes to supplement limited data with intuition or theory based corrections.
TomoSAR-ALISTA: Efficient TomoSAR Imaging via Deep Unfolded Network
May 05, 2022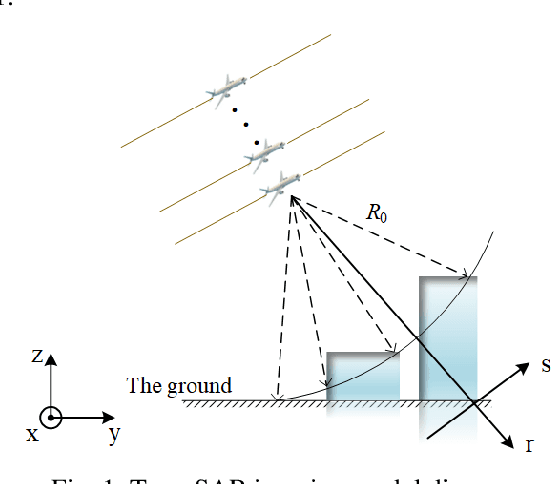
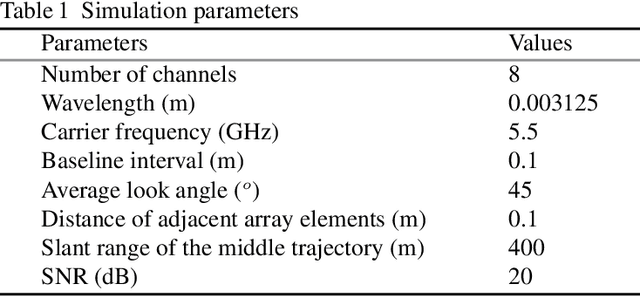

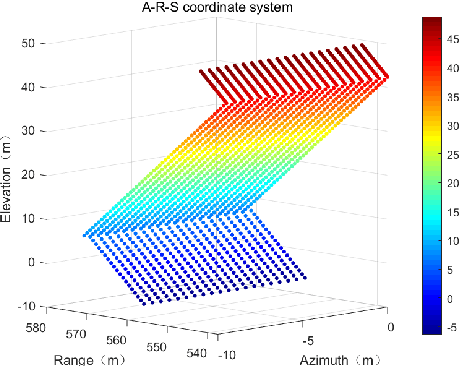
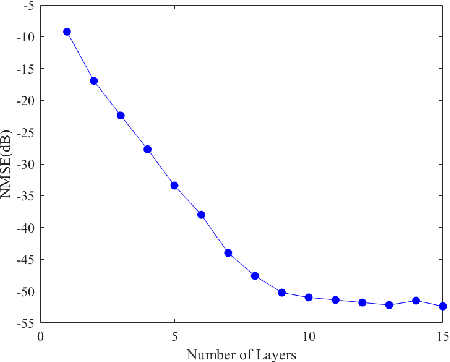
Synthetic aperture radar (SAR) tomography (TomoSAR) has attracted remarkable interest for its ability in achieving three-dimensional reconstruction along the elevation direction from multiple observations. In recent years, compressed sensing (CS) technique has been introduced into TomoSAR considering for its super-resolution ability with limited samples. Whereas, the CS-based methods suffer from several drawbacks, including weak noise resistance, high computational complexity and complex parameter fine-tuning. Among the different CS algorithms, iterative soft-thresholding algorithm (ISTA) is widely used as a robust reconstruction approach, however, the parameters in the ISTA algorithm are manually chosen, which usually requires a time-consuming fine-tuning process to achieve the best performance. Aiming at efficient TomoSAR imaging, a novel sparse unfolding network named analytic learned ISTA (ALISTA) is proposed towards the TomoSAR imaging problem in this paper, and the key parameters of ISTA are learned from training data via deep learning to avoid complex parameter fine-tuning and significantly relieves the training burden. In addition, experiments verify that it is feasible to use traditional CS algorithms as training labels, which provides a tangible supervised training method to achieve better 3D reconstruction performance even in the absence of labeled data in real applications.
Learning Model Predictive Control for Quadrotors
Feb 15, 2022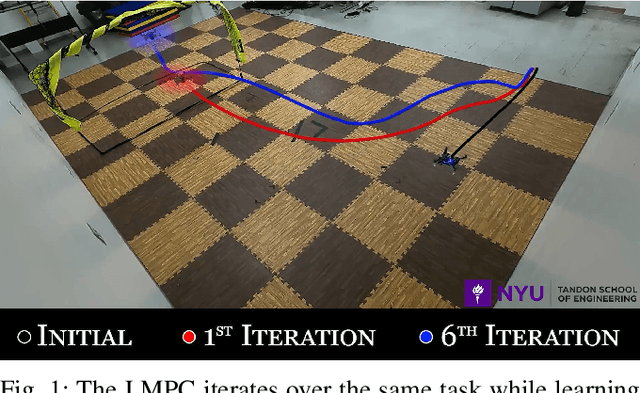
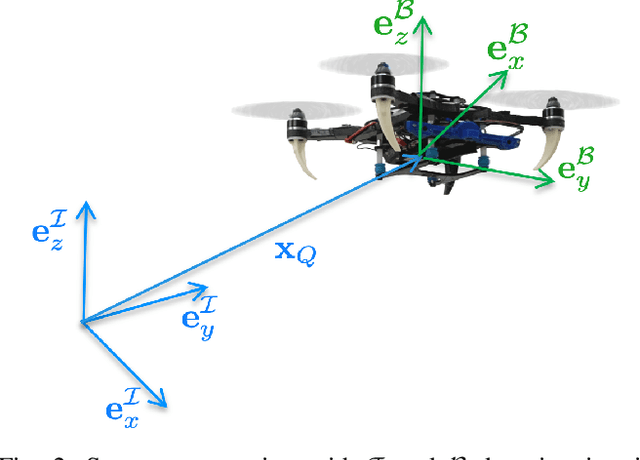
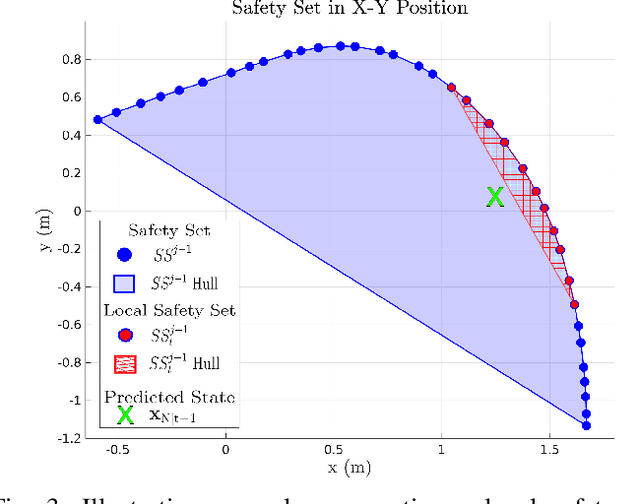
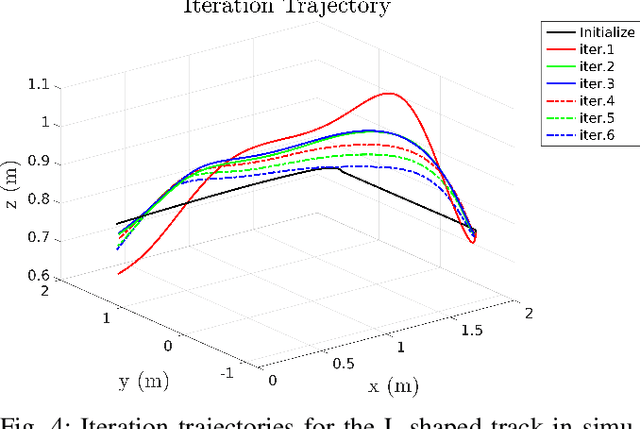
Aerial robots can enhance their safe and agile navigation in complex and cluttered environments by efficiently exploiting the information collected during a given task. In this paper, we address the learning model predictive control problem for quadrotors. We design a learning receding--horizon nonlinear control strategy directly formulated on the system nonlinear manifold configuration space SO(3)xR^3. The proposed approach exploits past successful task iterations to improve the system performance over time while respecting system dynamics and actuator constraints. We further relax its computational complexity making it compatible with real-time quadrotor control requirements. We show the effectiveness of the proposed approach in learning a minimum time control task, respecting dynamics, actuators, and environment constraints. Several experiments in simulation and real-world set-up validate the proposed approach.
Learning Trajectories of Hamiltonian Systems with Neural Networks
Apr 11, 2022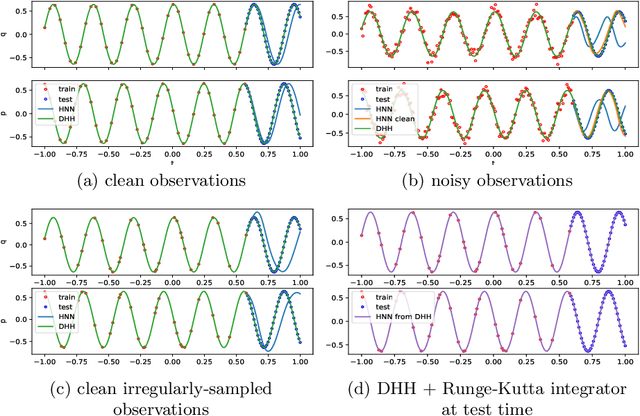
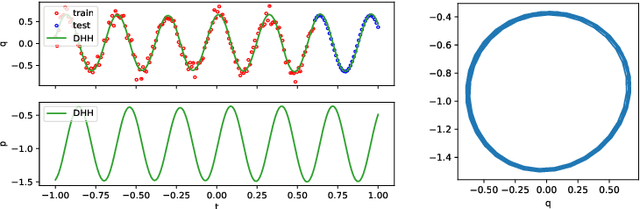
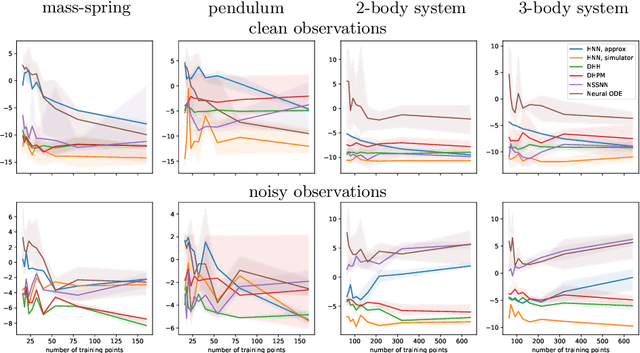
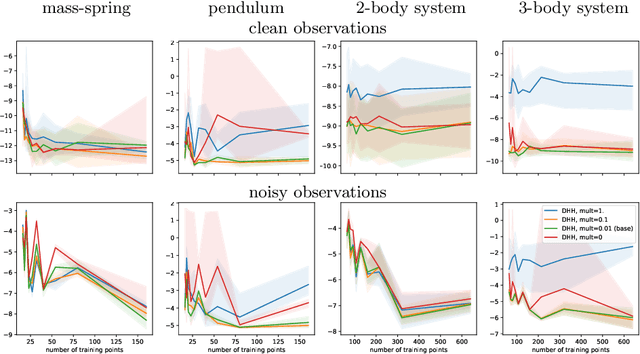
Modeling of conservative systems with neural networks is an area of active research. A popular approach is to use Hamiltonian neural networks (HNNs) which rely on the assumptions that a conservative system is described with Hamilton's equations of motion. Many recent works focus on improving the integration schemes used when training HNNs. In this work, we propose to enhance HNNs with an estimation of a continuous-time trajectory of the modeled system using an additional neural network, called a deep hidden physics model in the literature. We demonstrate that the proposed integration scheme works well for HNNs, especially with low sampling rates, noisy and irregular observations.
Promoting Saliency From Depth: Deep Unsupervised RGB-D Saliency Detection
May 15, 2022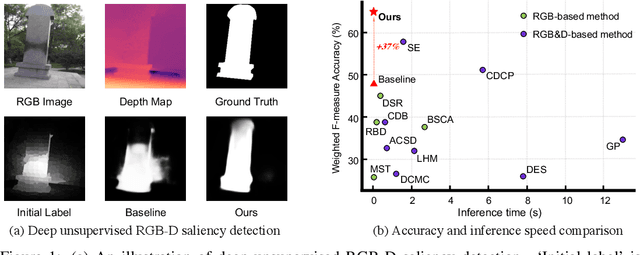
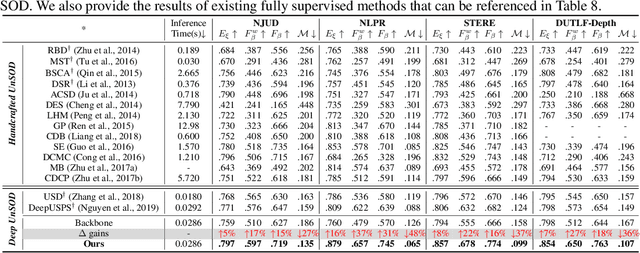
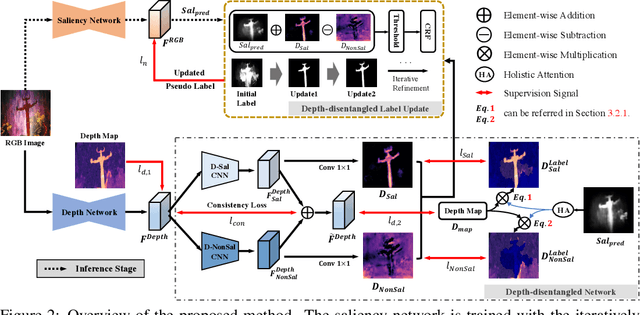
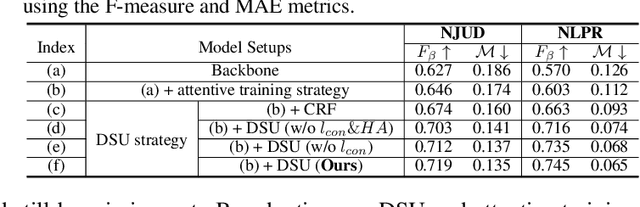
Growing interests in RGB-D salient object detection (RGB-D SOD) have been witnessed in recent years, owing partly to the popularity of depth sensors and the rapid progress of deep learning techniques. Unfortunately, existing RGB-D SOD methods typically demand large quantity of training images being thoroughly annotated at pixel-level. The laborious and time-consuming manual annotation has become a real bottleneck in various practical scenarios. On the other hand, current unsupervised RGB-D SOD methods still heavily rely on handcrafted feature representations. This inspires us to propose in this paper a deep unsupervised RGB-D saliency detection approach, which requires no manual pixel-level annotation during training. It is realized by two key ingredients in our training pipeline. First, a depth-disentangled saliency update (DSU) framework is designed to automatically produce pseudo-labels with iterative follow-up refinements, which provides more trustworthy supervision signals for training the saliency network. Second, an attentive training strategy is introduced to tackle the issue of noisy pseudo-labels, by properly re-weighting to highlight the more reliable pseudo-labels. Extensive experiments demonstrate the superior efficiency and effectiveness of our approach in tackling the challenging unsupervised RGB-D SOD scenarios. Moreover, our approach can also be adapted to work in fully-supervised situation. Empirical studies show the incorporation of our approach gives rise to notably performance improvement in existing supervised RGB-D SOD models.
RobotCore: An Open Architecture for Hardware Acceleration in ROS 2
May 08, 2022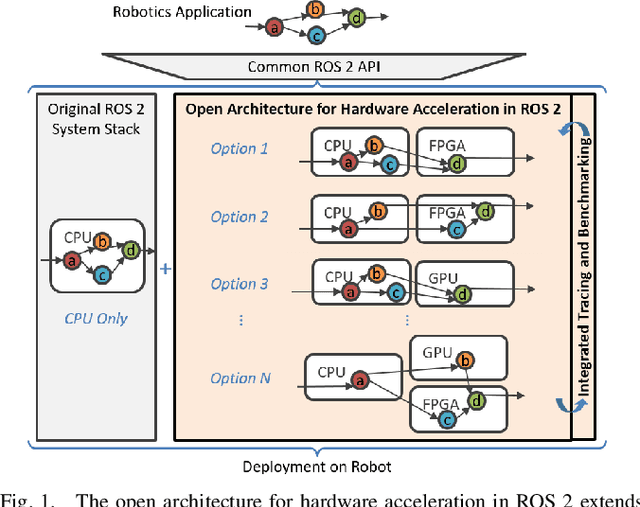
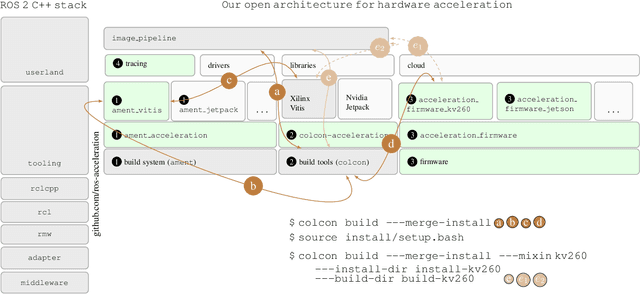
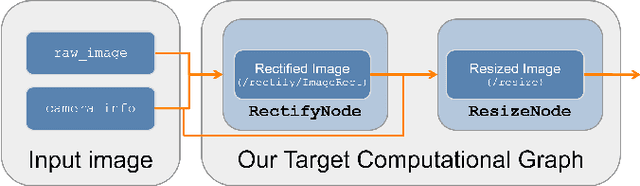
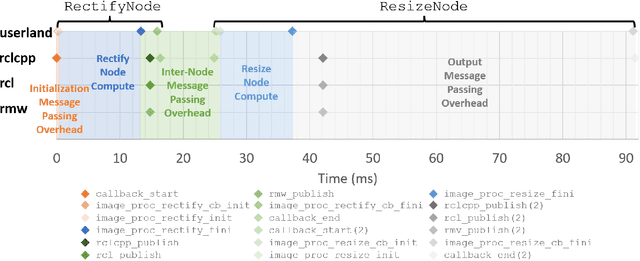
Hardware acceleration can revolutionize robotics, enabling new applications by speeding up robot response times while remaining power-efficient. However, the diversity of acceleration options makes it difficult for roboticists to easily deploy accelerated systems without expertise in each specific hardware platform. In this work, we address this challenge with RobotCore, an architecture to integrate hardware acceleration in the widely-used ROS 2 robotics software framework. This architecture is target-agnostic (supports edge, workstation, data center, or cloud targets) and accelerator-agnostic (supports both FPGAs and GPUs). It builds on top of the common ROS 2 build system and tools and is easily portable across different research and commercial solutions through a new firmware layer. We also leverage the Linux Tracing Toolkit next generation (LTTng) for low-overhead real-time tracing and benchmarking. To demonstrate the acceleration enabled by this architecture, we use it to deploy a ROS 2 perception computational graph on a CPU and FPGA. We employ our integrated tracing and benchmarking to analyze bottlenecks, uncovering insights that guide us to improve FPGA communication efficiency. In particular, we design an intra-FPGA ROS 2 node communication queue to enable faster data flows, and use it in conjunction with FPGA-accelerated nodes to achieve a 24.42% speedup over a CPU.
Adaptive Gaussian Fuzzy Classifier for Real-Time Emotion Recognition in Computer Games
Mar 05, 2021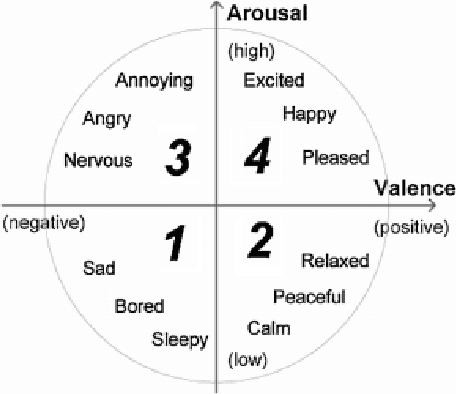
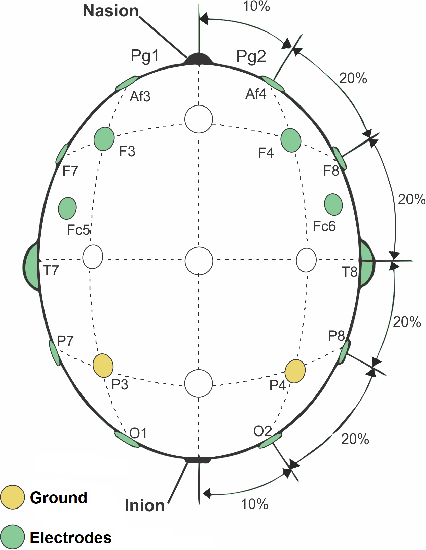
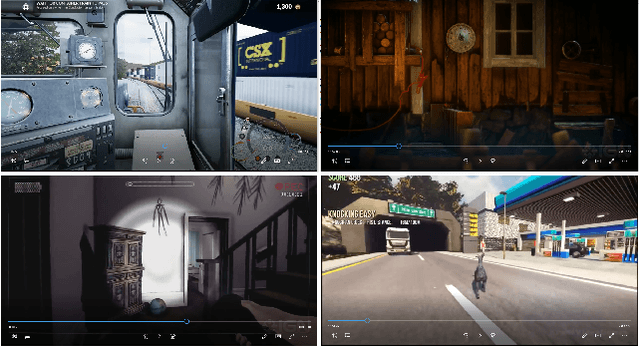
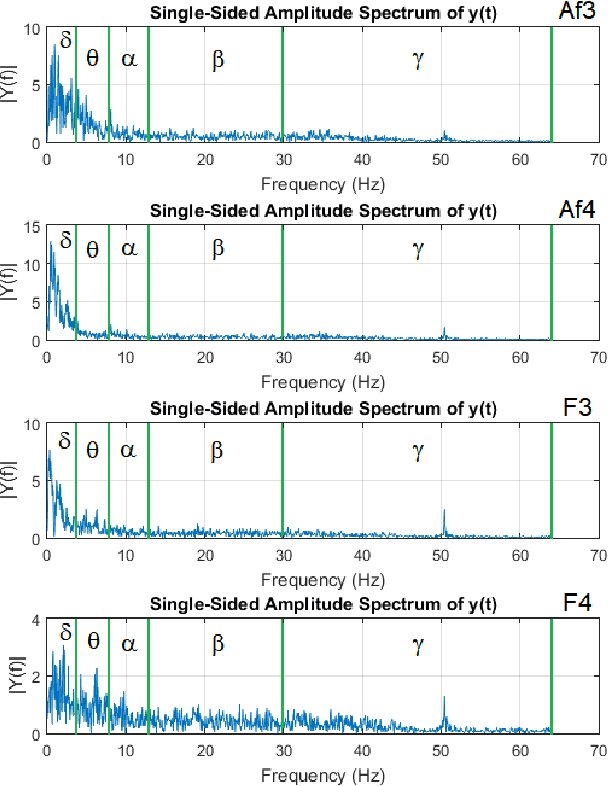
Human emotion recognition has become a need for more realistic and interactive machines and computer systems. The greatest challenge is the availability of high-performance algorithms to effectively manage individual differences and nonstationarities in physiological data streams, i.e., algorithms that self-customize to a user with no subject-specific calibration data. We describe an evolving Gaussian Fuzzy Classifier (eGFC), which is supported by an online semi-supervised learning algorithm to recognize emotion patterns from electroencephalogram (EEG) data streams. We extract features from the Fourier spectrum of EEG data. The data are provided by 28 individuals playing the games 'Train Sim World', 'Unravel', 'Slender The Arrival', and 'Goat Simulator' - a public dataset. Different emotions prevail, namely, boredom, calmness, horror and joy. We analyze the effect of individual electrodes, time window lengths, and frequency bands on the accuracy of user-independent eGFCs. We conclude that both brain hemispheres may assist classification, especially electrodes on the frontal (Af3-Af4), occipital (O1-O2), and temporal (T7-T8) areas. We observe that patterns may be eventually found in any frequency band; however, the Alpha (8-13Hz), Delta (1-4Hz), and Theta (4-8Hz) bands, in this order, are the highest correlated with emotion classes. eGFC has shown to be effective for real-time learning of EEG data. It reaches a 72.2% accuracy using a variable rule base, 10-second windows, and 1.8ms/sample processing time in a highly-stochastic time-varying 4-class classification problem.
A SSIM Guided cGAN Architecture For Clinically Driven Generative Image Synthesis of Multiplexed Spatial Proteomics Channels
May 20, 2022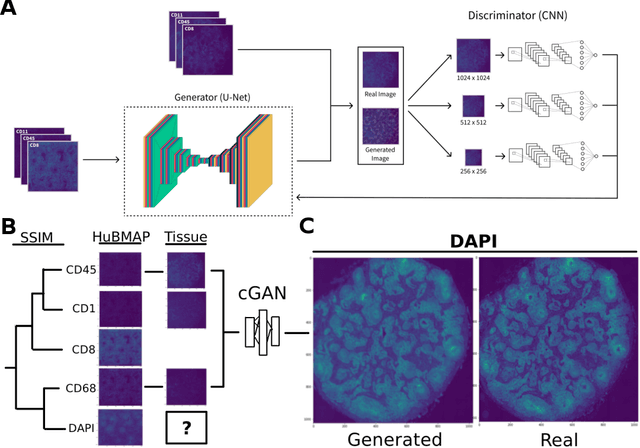
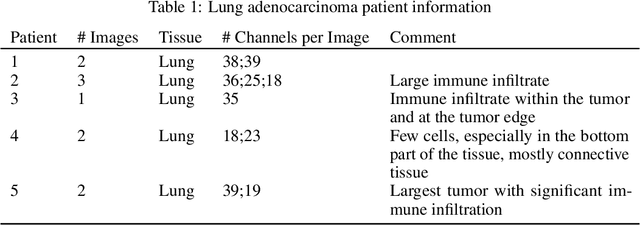


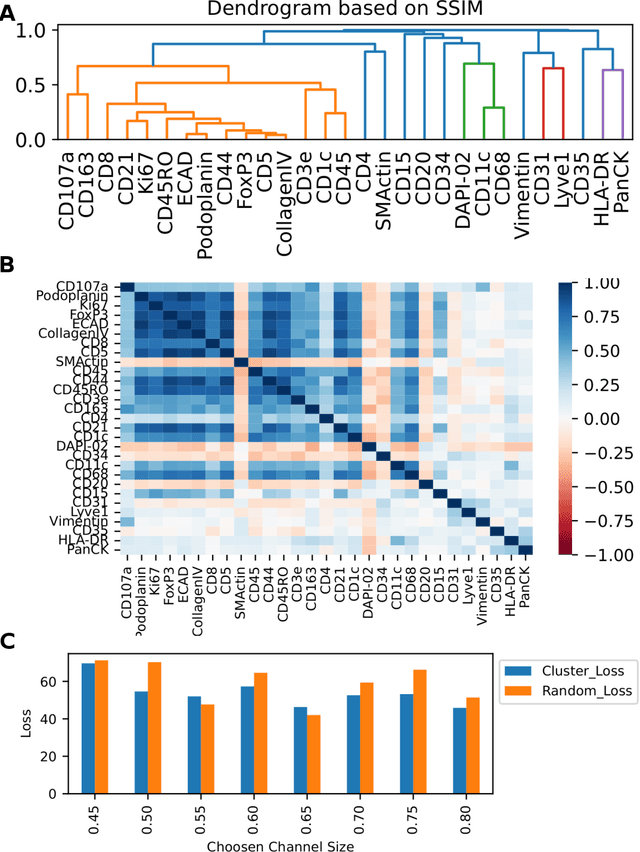
Here we present a structural similarity index measure (SSIM) guided conditional Generative Adversarial Network (cGAN) that generatively performs image-to-image (i2i) synthesis to generate photo-accurate protein channels in multiplexed spatial proteomics images. This approach can be utilized to accurately generate missing spatial proteomics channels that were not included during experimental data collection either at the bench or the clinic. Experimental spatial proteomic data from the Human BioMolecular Atlas Program (HuBMAP) was used to generate spatial representations of missing proteins through a U-Net based image synthesis pipeline. HuBMAP channels were hierarchically clustered by the (SSIM) as a heuristic to obtain the minimal set needed to recapitulate the underlying biology represented by the spatial landscape of proteins. We subsequently prove that our SSIM based architecture allows for scaling of generative image synthesis to slides with up to 100 channels, which is better than current state of the art algorithms which are limited to data with 11 channels. We validate these claims by generating a new experimental spatial proteomics data set from human lung adenocarcinoma tissue sections and show that a model trained on HuBMAP can accurately synthesize channels from our new data set. The ability to recapitulate experimental data from sparsely stained multiplexed histological slides containing spatial proteomic will have tremendous impact on medical diagnostics and drug development, and also raises important questions on the medical ethics of utilizing data produced by generative image synthesis in the clinical setting. The algorithm that we present in this paper will allow researchers and clinicians to save time and costs in proteomics based histological staining while also increasing the amount of data that they can generate through their experiments.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge