"Time": models, code, and papers
Weakly Supervised Learning Significantly Reduces the Number of Labels Required for Intracranial Hemorrhage Detection on Head CT
Nov 29, 2022
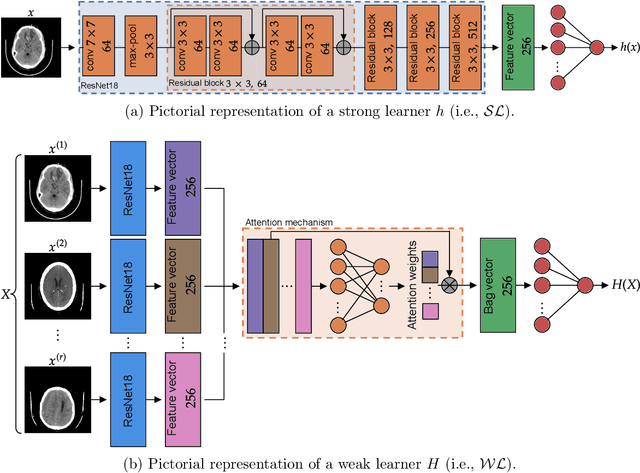

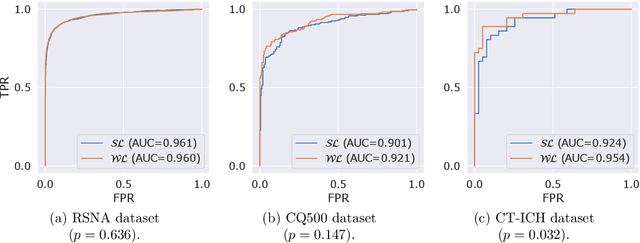
Modern machine learning pipelines, in particular those based on deep learning (DL) models, require large amounts of labeled data. For classification problems, the most common learning paradigm consists of presenting labeled examples during training, thus providing strong supervision on what constitutes positive and negative samples. This constitutes a major obstacle for the development of DL models in radiology--in particular for cross-sectional imaging (e.g., computed tomography [CT] scans)--where labels must come from manual annotations by expert radiologists at the image or slice-level. These differ from examination-level annotations, which are coarser but cheaper, and could be extracted from radiology reports using natural language processing techniques. This work studies the question of what kind of labels should be collected for the problem of intracranial hemorrhage detection in brain CT. We investigate whether image-level annotations should be preferred to examination-level ones. By framing this task as a multiple instance learning problem, and employing modern attention-based DL architectures, we analyze the degree to which different levels of supervision improve detection performance. We find that strong supervision (i.e., learning with local image-level annotations) and weak supervision (i.e., learning with only global examination-level labels) achieve comparable performance in examination-level hemorrhage detection (the task of selecting the images in an examination that show signs of hemorrhage) as well as in image-level hemorrhage detection (highlighting those signs within the selected images). Furthermore, we study this behavior as a function of the number of labels available during training. Our results suggest that local labels may not be necessary at all for these tasks, drastically reducing the time and cost involved in collecting and curating datasets.
BatmanNet: Bi-branch Masked Graph Transformer Autoencoder for Molecular Representation
Nov 29, 2022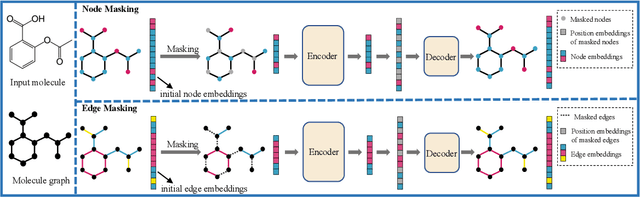
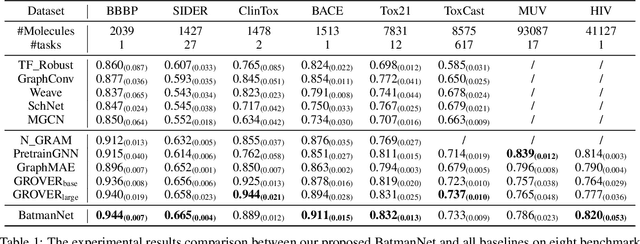
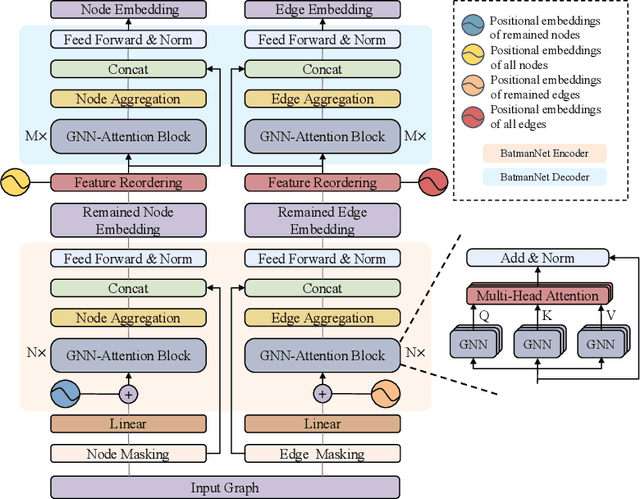
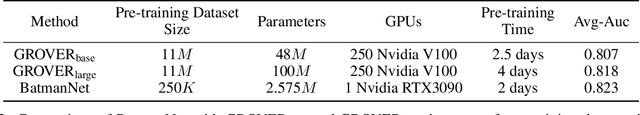
Although substantial efforts have been made using graph neural networks (GNNs) for AI-driven drug discovery (AIDD), effective molecular representation learning remains an open challenge, especially in the case of insufficient labeled molecules. Recent studies suggest that big GNN models pre-trained by self-supervised learning on unlabeled datasets enable better transfer performance in downstream molecular property prediction tasks. However, they often require large-scale datasets and considerable computational resources, which is time-consuming, computationally expensive, and environmentally unfriendly. To alleviate these limitations, we propose a novel pre-training model for molecular representation learning, Bi-branch Masked Graph Transformer Autoencoder (BatmanNet). BatmanNet features two tailored and complementary graph autoencoders to reconstruct the missing nodes and edges from a masked molecular graph. To our surprise, BatmanNet discovered that the highly masked proportion (60%) of the atoms and bonds achieved the best performance. We further propose an asymmetric graph-based encoder-decoder architecture for either nodes and edges, where a transformer-based encoder only takes the visible subset of nodes or edges, and a lightweight decoder reconstructs the original molecule from the latent representation and mask tokens. With this simple yet effective asymmetrical design, our BatmanNet can learn efficiently even from a much smaller-scale unlabeled molecular dataset to capture the underlying structural and semantic information, overcoming a major limitation of current deep neural networks for molecular representation learning. For instance, using only 250K unlabelled molecules as pre-training data, our BatmanNet with 2.575M parameters achieves a 0.5% improvement on the average AUC compared with the current state-of-the-art method with 100M parameters pre-trained on 11M molecules.
Predicting Survival Outcomes in the Presence of Unlabeled Data
Oct 25, 2022Many clinical studies require the follow-up of patients over time. This is challenging: apart from frequently observed drop-out, there are often also organizational and financial challenges, which can lead to reduced data collection and, in turn, can complicate subsequent analyses. In contrast, there is often plenty of baseline data available of patients with similar characteristics and background information, e.g., from patients that fall outside the study time window. In this article, we investigate whether we can benefit from the inclusion of such unlabeled data instances to predict accurate survival times. In other words, we introduce a third level of supervision in the context of survival analysis, apart from fully observed and censored instances, we also include unlabeled instances. We propose three approaches to deal with this novel setting and provide an empirical comparison over fifteen real-life clinical and gene expression survival datasets. Our results demonstrate that all approaches are able to increase the predictive performance over independent test data. We also show that integrating the partial supervision provided by censored data in a semi-supervised wrapper approach generally provides the best results, often achieving high improvements, compared to not using unlabeled data.
Early Discovery of Disappearing Entities in Microblogs
Oct 13, 2022
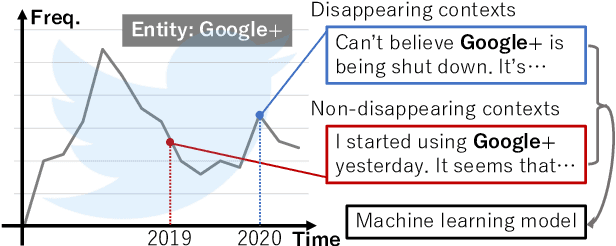
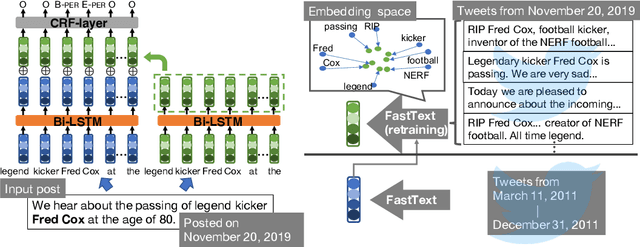
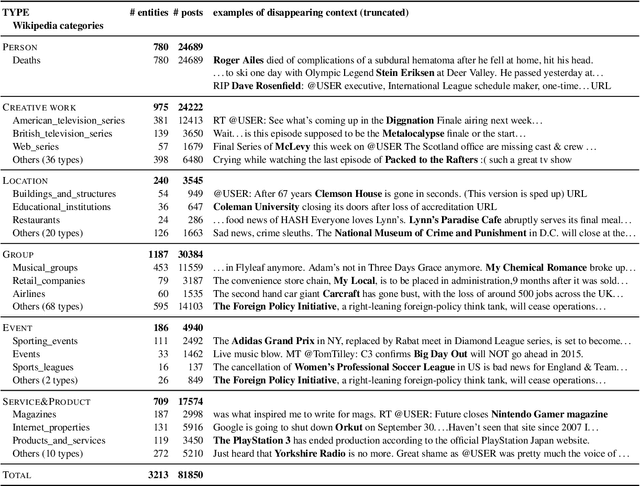
We make decisions by reacting to changes in the real world, in particular, the emergence and disappearance of impermanent entities such as events, restaurants, and services. Because we want to avoid missing out on opportunities or making fruitless actions after they have disappeared, it is important to know when entities disappear as early as possible. We thus tackle the task of detecting disappearing entities from microblogs, whose posts mention various entities, in a timely manner. The major challenge is detecting uncertain contexts of disappearing entities from noisy microblog posts. To collect these disappearing contexts, we design time-sensitive distant supervision, which utilizes entities from the knowledge base and time-series posts, for this task to build large-scale Twitter datasets\footnote{We will release the datasets (tweet IDs) used in the experiments to promote reproducibility.} for English and Japanese. To ensure robust detection in noisy environments, we refine pretrained word embeddings of the detection model on microblog streams of the target day. Experimental results on the Twitter datasets confirmed the effectiveness of the collected labeled data and refined word embeddings; more than 70\% of the detected disappearing entities in Wikipedia are discovered earlier than the update on Wikipedia, and the average lead-time is over one month.
Marginalized particle Gibbs for multiple state-space models coupled through shared parameters
Oct 13, 2022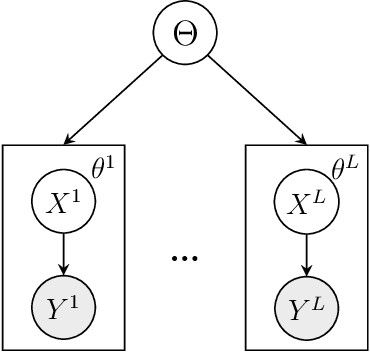
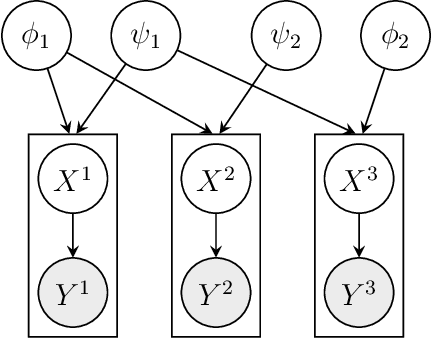
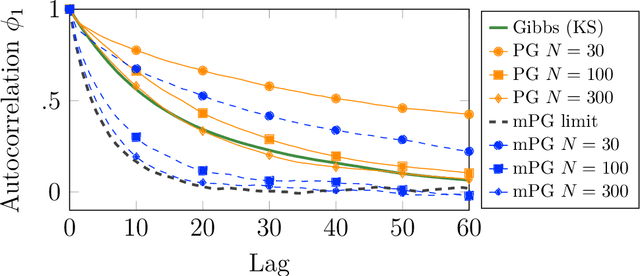
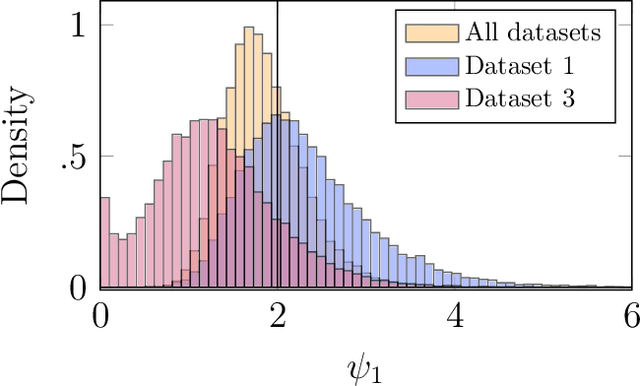
We consider Bayesian inference from multiple time series described by a common state-space model (SSM) structure, but where different subsets of parameters are shared between different submodels. An important example is disease-dynamics, where parameters can be either disease or location specific. Parameter inference in these models can be improved by systematically aggregating information from the different time series, most notably for short series. Particle Gibbs (PG) samplers are an efficient class of algorithms for inference in SSMs, in particular when conjugacy can be exploited to marginalize out model parameters from the state update. We present two different PG samplers that marginalize static model parameters on-the-fly: one that updates one model at a time conditioned on the datasets for the other models, and one that concurrently updates all models by stacking them into a high-dimensional SSM. The distinctive features of each sampler make them suitable for different modelling contexts. We provide insights on when each sampler should be used and show that they can be combined to form an efficient PG sampler for a model with strong dependencies between states and parameters. The performance is illustrated on two linear-Gaussian examples and on a real-world example on the spread of mosquito-borne diseases.
Experiments in Underwater Feature Tracking with Performance Guarantees Using a Small AUV
Oct 05, 2022
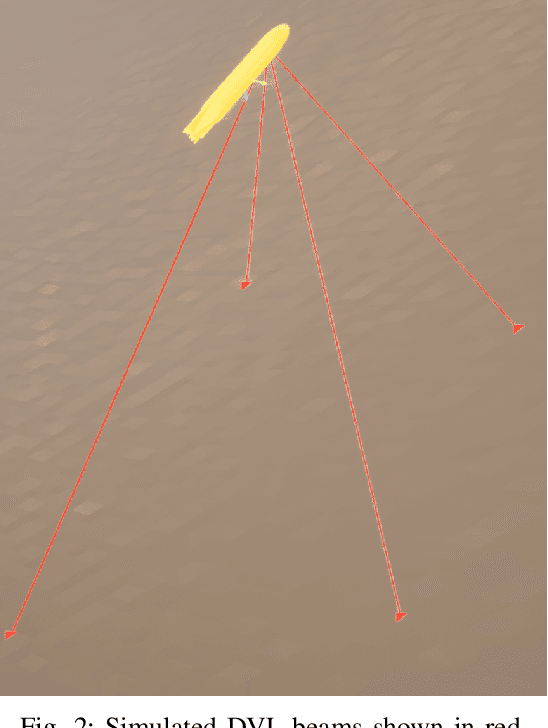

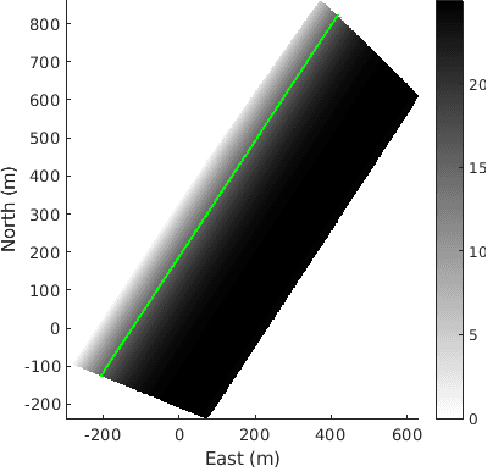
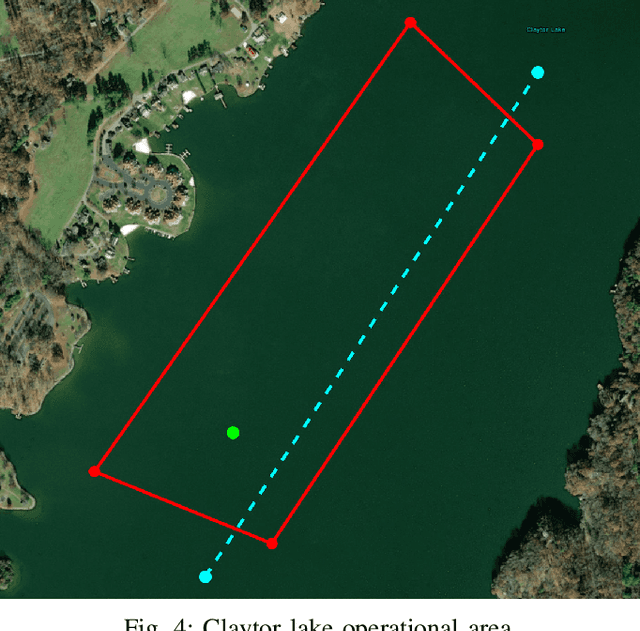
We present the results of experiments performed using a small autonomous underwater vehicle to determine the location of an isobath within a bounded area. The primary contribution of this work is to implement and integrate several recent developments real-time planning for environmental mapping, and to demonstrate their utility in a challenging practical example. We model the bathymetry within the operational area using a Gaussian process and propose a reward function that represents the task of mapping a desired isobath. As is common in applications where plans must be continually updated based on real-time sensor measurements, we adopt a receding horizon framework where the vehicle continually computes near-optimal paths. The sequence of paths does not, in general, inherit the optimality properties of each individual path. Our real-time planning implementation incorporates recent results that lead to performance guarantees for receding-horizon planning.
Search-Based Path Planning Algorithm for Autonomous Parking:Multi-Heuristic Hybrid A*
Oct 17, 2022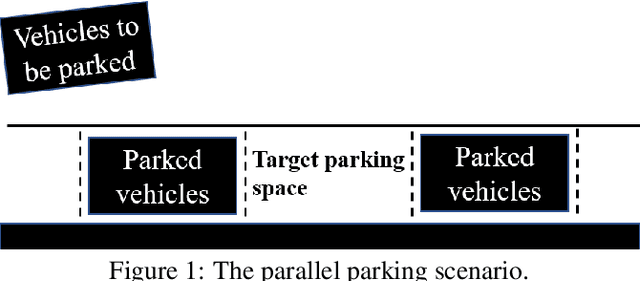
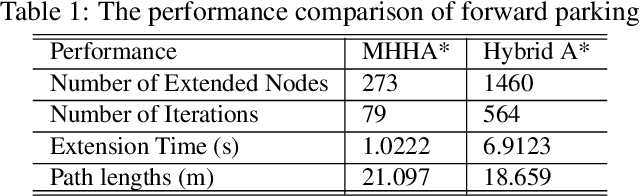
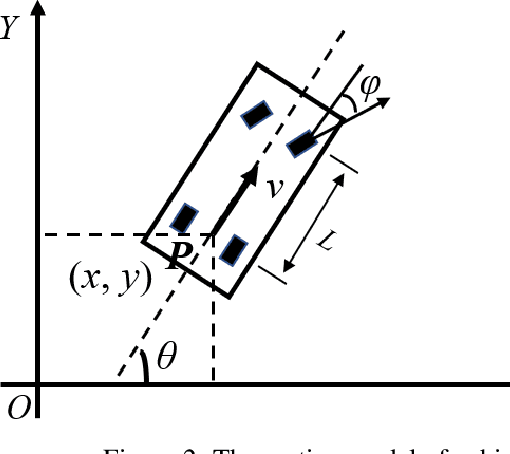
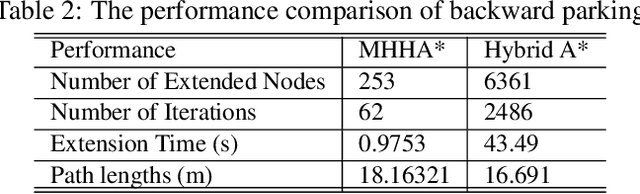
This paper proposed a novel method for autonomous parking. Autonomous parking has received a lot of attention because of its convenience, but due to the complex environment and the non-holonomic constraints of vehicle, it is difficult to get a collision-free and feasible path in a short time. To solve this problem, this paper introduced a novel algorithm called Multi-Heuristic Hybrid A* (MHHA*) which incorporates the characteristic of Multi-Heuristic A* and Hybrid A*. So it could provide the guarantee for completeness, the avoidance of local minimum and sub-optimality, and generate a feasible path in a short time. And this paper also proposed a new collision check method based on coordinate transformation which could improve the computational efficiency. The performance of the proposed method was compared with Hybrid A* in simulation experiments and its superiority has been proved.
Does an ensemble of GANs lead to better performance when training segmentation networks with synthetic images?
Nov 08, 2022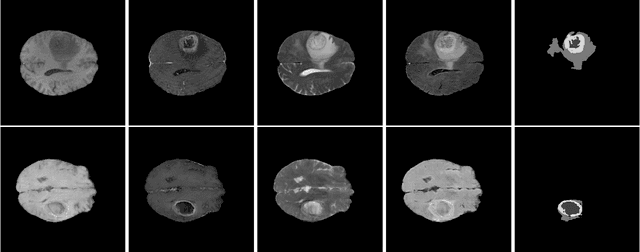
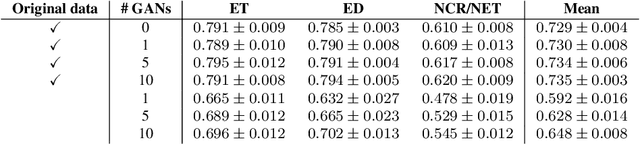
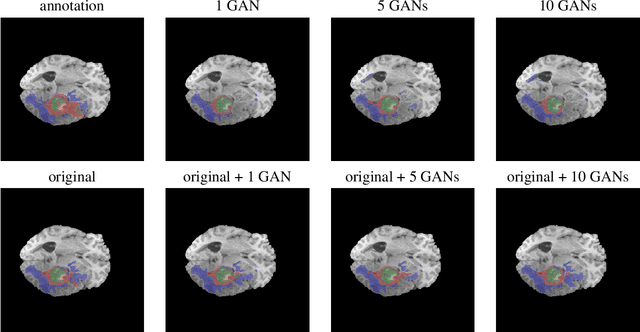
Large annotated datasets are required to train segmentation networks. In medical imaging, it is often difficult, time consuming and expensive to create such datasets, and it may also be difficult to share these datasets with other researchers. Different AI models can today generate very realistic synthetic images, which can potentially be openly shared as they do not belong to specific persons. However, recent work has shown that using synthetic images for training deep networks often leads to worse performance compared to using real images. Here we demonstrate that using synthetic images and annotations from an ensemble of 10 GANs, instead of from a single GAN, increases the Dice score on real test images with 4.7 % to 14.0 % on specific classes.
pyRDDLGym: From RDDL to Gym Environments
Nov 14, 2022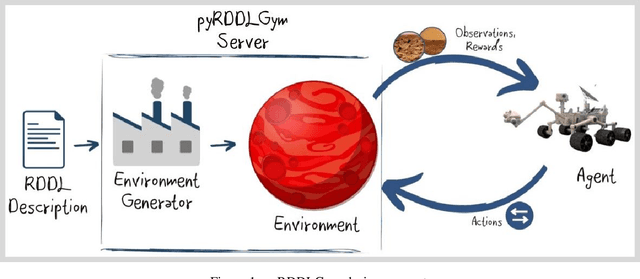
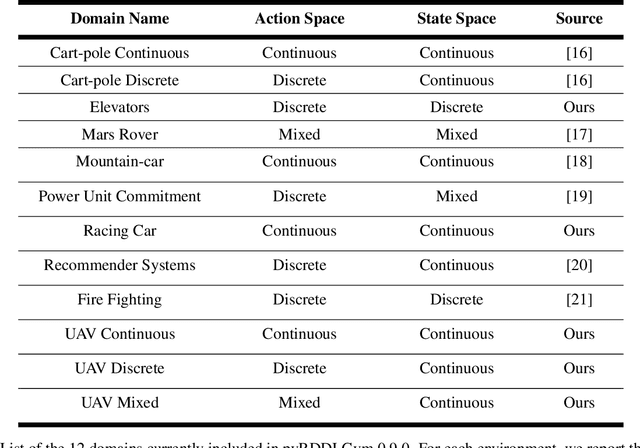


We present pyRDDLGym, a Python framework for auto-generation of OpenAI Gym environments from RDDL declerative description. The discrete time step evolution of variables in RDDL is described by conditional probability functions, which fits naturally into the Gym step scheme. Furthermore, since RDDL is a lifted description, the modification and scaling up of environments to support multiple entities and different configurations becomes trivial rather than a tedious process prone to errors. We hope that pyRDDLGym will serve as a new wind in the reinforcement learning community by enabling easy and rapid development of benchmarks due to the unique expressive power of RDDL. By providing explicit access to the model in the RDDL description, pyRDDLGym can also facilitate research on hybrid approaches for learning from interaction while leveraging model knowledge. We present the design and built-in examples of pyRDDLGym, and the additions made to the RDDL language that were incorporated into the framework.
Hypothesis Transfer in Bandits by Weighted Models
Nov 14, 2022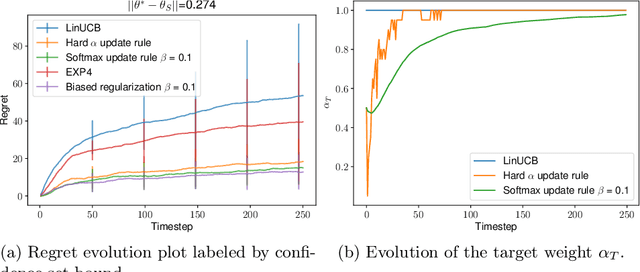
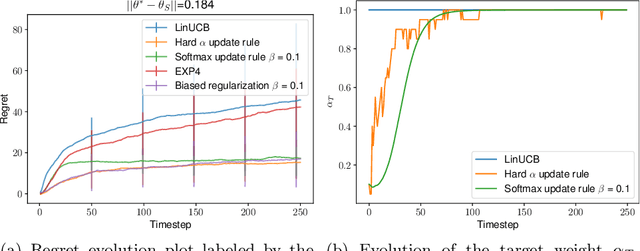
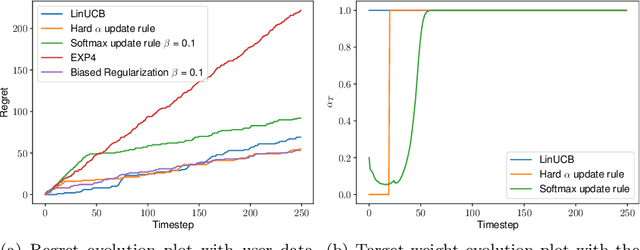
We consider the problem of contextual multi-armed bandits in the setting of hypothesis transfer learning. That is, we assume having access to a previously learned model on an unobserved set of contexts, and we leverage it in order to accelerate exploration on a new bandit problem. Our transfer strategy is based on a re-weighting scheme for which we show a reduction in the regret over the classic Linear UCB when transfer is desired, while recovering the classic regret rate when the two tasks are unrelated. We further extend this method to an arbitrary amount of source models, where the algorithm decides which model is preferred at each time step. Additionally we discuss an approach where a dynamic convex combination of source models is given in terms of a biased regularization term in the classic LinUCB algorithm. The algorithms and the theoretical analysis of our proposed methods substantiated by empirical evaluations on simulated and real-world data.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge