"Time": models, code, and papers
Large Scale Radio Frequency Wideband Signal Detection & Recognition
Nov 04, 2022
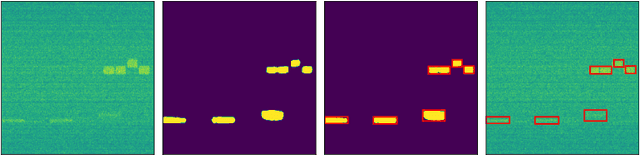
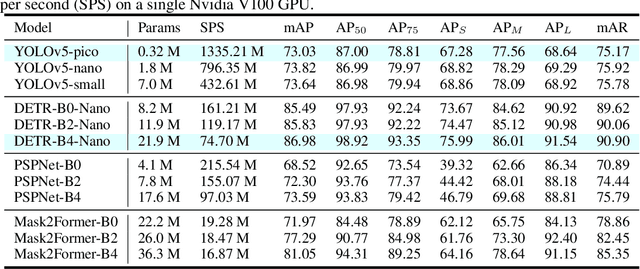

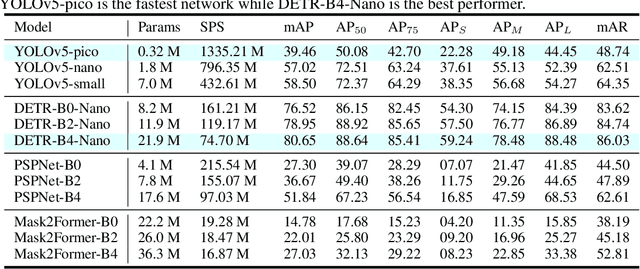
Applications of deep learning to the radio frequency (RF) domain have largely concentrated on the task of narrowband signal classification after the signals of interest have already been detected and extracted from a wideband capture. To encourage broader research with wideband operations, we introduce the WidebandSig53 (WBSig53) dataset which consists of 550 thousand synthetically-generated samples from 53 different signal classes containing approximately 2 million unique signals. We extend the TorchSig signal processing machine learning toolkit for open-source and customizable generation, augmentation, and processing of the WBSig53 dataset. We conduct experiments using state of the art (SoTA) convolutional neural networks and transformers with the WBSig53 dataset. We investigate the performance of signal detection tasks, i.e. detect the presence, time, and frequency of all signals present in the input data, as well as the performance of signal recognition tasks, where networks detect the presence, time, frequency, and modulation family of all signals present in the input data. Two main approaches to these tasks are evaluated with segmentation networks and object detection networks operating on complex input spectrograms. Finally, we conduct comparative analysis of the various approaches in terms of the networks' mean average precision, mean average recall, and the speed of inference.
Parameter Inference of Time Series by Delay Embeddings and Learning Differentiable Operators
Mar 11, 2022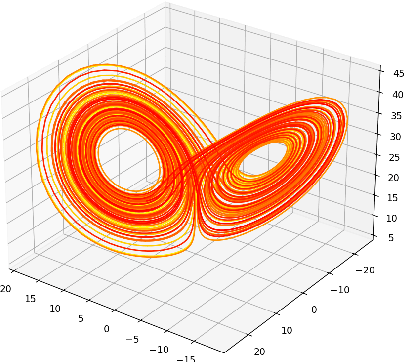
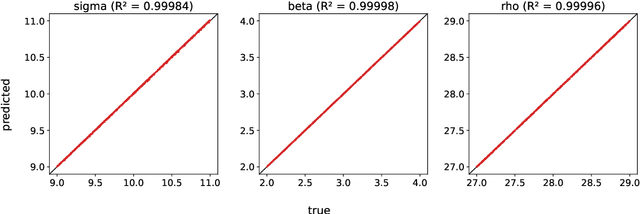
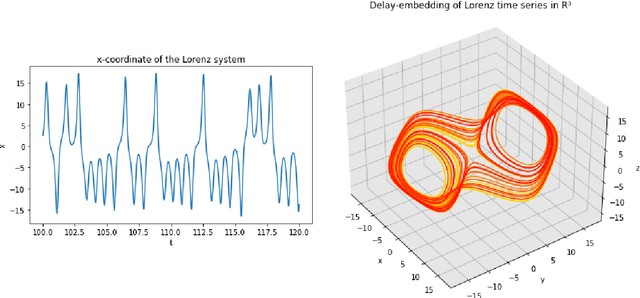
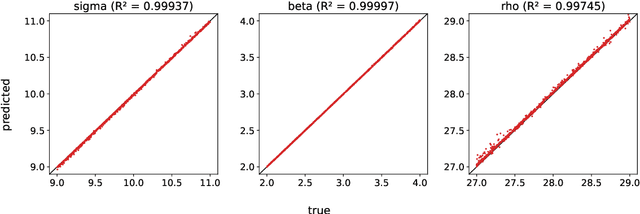
A common issue in dealing with real-world dynamical systems is identifying system parameters responsible for its behavior. A frequent scenario is that one has time series data, along with corresponding parameter labels, but there exists new time series with unknown parameter labels, which one seeks to identify. We tackle this problem by first delay-embedding the time series into a higher dimension to obtain a proper ordinary differential equation (ODE), and then having a neural network learn to predict future time-steps of the trajectory given the present time-step. We then use the learned neural network to backpropagate prediction errors through the parameter inputs of the neural network in order to obtain a gradient in parameter space. Using this gradient, we can approximately identify parameters of time series. We demonstrate the viability of our approach on the chaotic Lorenz system, as well as real-world data with the Hall-effect Thruster (HET).
SurvSet: An open-source time-to-event dataset repository
Mar 07, 2022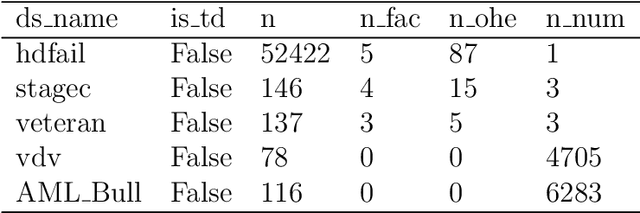
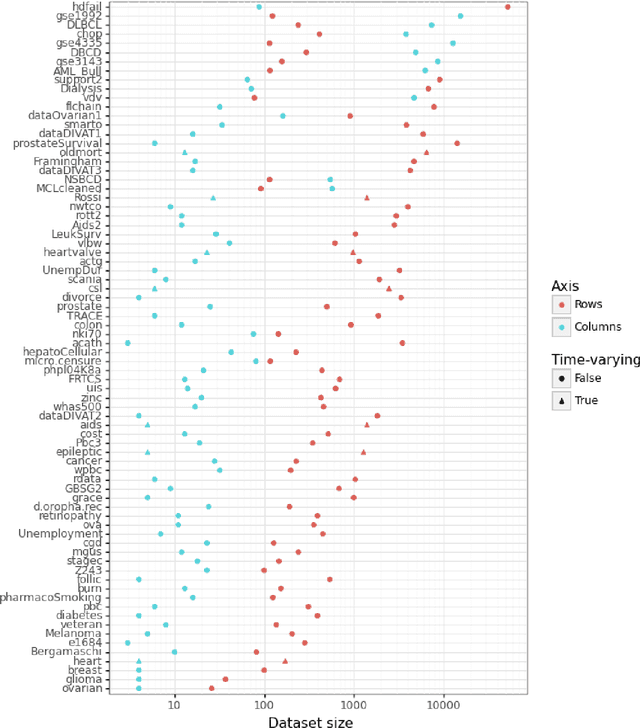
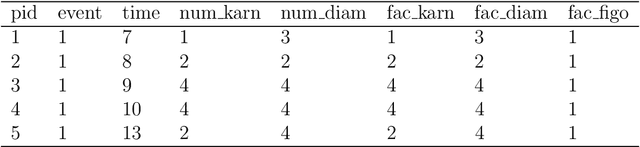
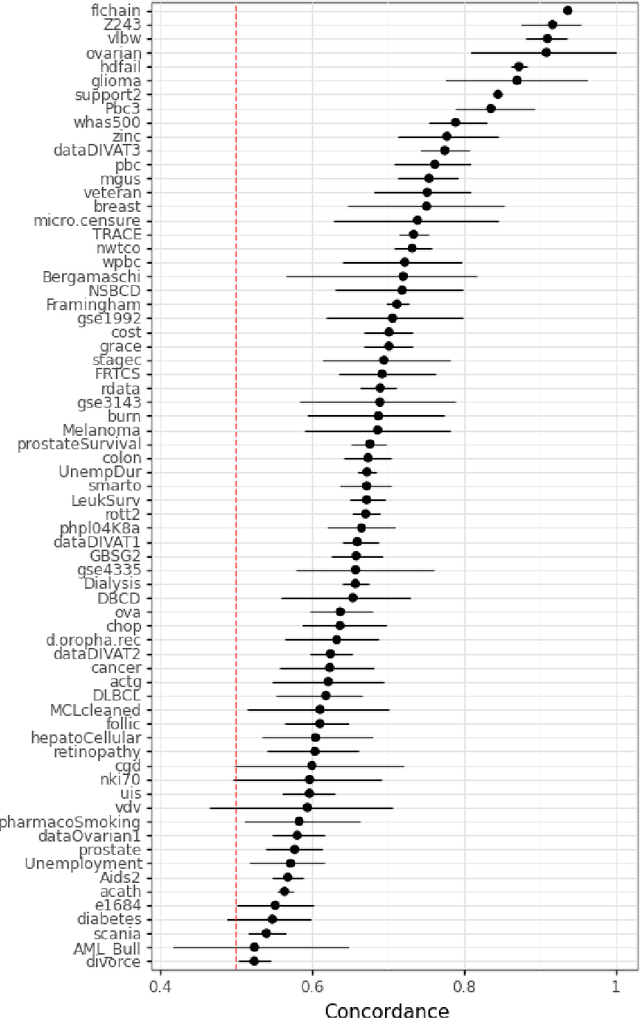
Time-to-event (T2E) analysis is a branch of statistics that models the duration of time it takes for an event to occur. Such events can include outcomes like death, unemployment, or product failure. Most modern machine learning (ML) algorithms, like decision trees and kernel methods, are supported for T2E modelling with data science software (python and R). To complement these developments, SurvSet is the first open-source T2E dataset repository designed for a rapid benchmarking of ML algorithms and statistical methods. The data in SurvSet have been consistently formatted so that a single preprocessing method will work for all datasets. SurvSet currently has 76 datasets which vary in dimensionality, time dependency, and background (the majority of which come from biomedicine). SurvSet is available on PyPI and can be installed with pip install SurvSet. R users can download the data directly from the corresponding git repository.
RSC: Accelerating Graph Neural Networks Training via Randomized Sparse Computations
Oct 19, 2022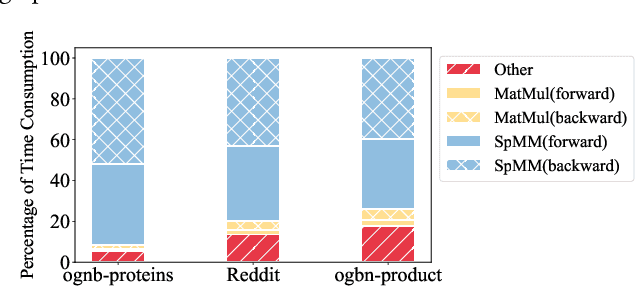
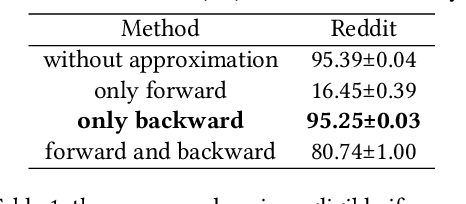

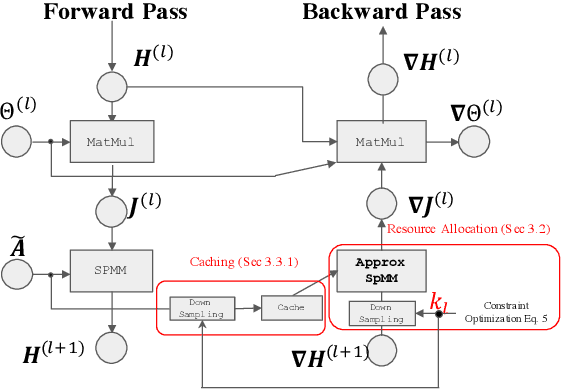
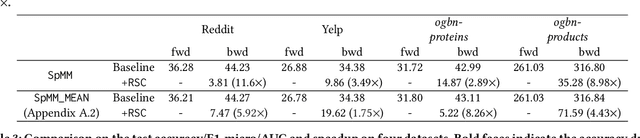
The training of graph neural networks (GNNs) is extremely time consuming because sparse graph-based operations are hard to be accelerated by hardware. Prior art explores trading off the computational precision to reduce the time complexity via sampling-based approximation. Based on the idea, previous works successfully accelerate the dense matrix based operations (e.g., convolution and linear) with negligible accuracy drop. However, unlike dense matrices, sparse matrices are stored in the irregular data format such that each row/column may have different number of non-zero entries. Thus, compared to the dense counterpart, approximating sparse operations has two unique challenges (1) we cannot directly control the efficiency of approximated sparse operation since the computation is only executed on non-zero entries; (2) sub-sampling sparse matrices is much more inefficient due to the irregular data format. To address the issues, our key idea is to control the accuracy-efficiency trade off by optimizing computation resource allocation layer-wisely and epoch-wisely. Specifically, for the first challenge, we customize the computation resource to different sparse operations, while limit the total used resource below a certain budget. For the second challenge, we cache previous sampled sparse matrices to reduce the epoch-wise sampling overhead. Finally, we propose a switching mechanisms to improve the generalization of GNNs trained with approximated operations. To this end, we propose Randomized Sparse Computation, which for the first time demonstrate the potential of training GNNs with approximated operations. In practice, rsc can achieve up to $11.6\times$ speedup for a single sparse operation and a $1.6\times$ end-to-end wall-clock time speedup with negligible accuracy drop.
Where to Pay Attention in Sparse Training for Feature Selection?
Nov 26, 2022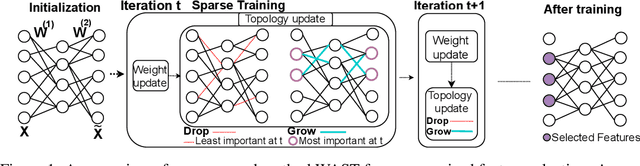
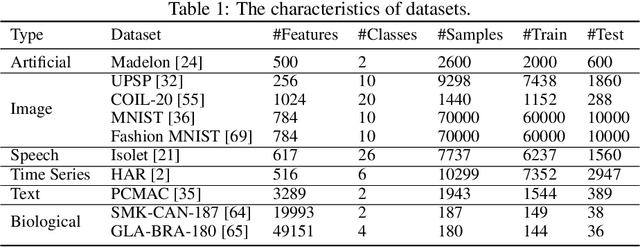
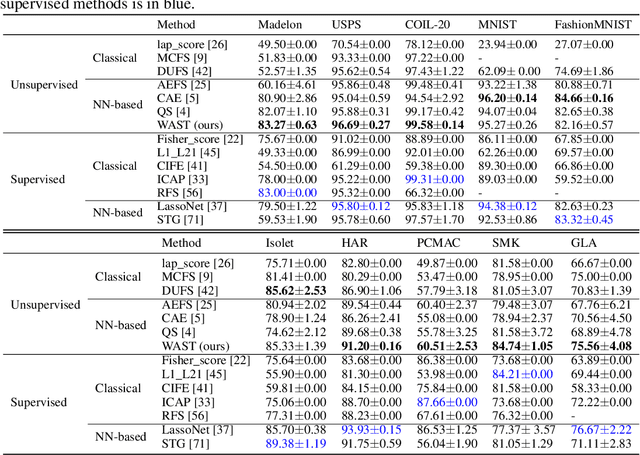
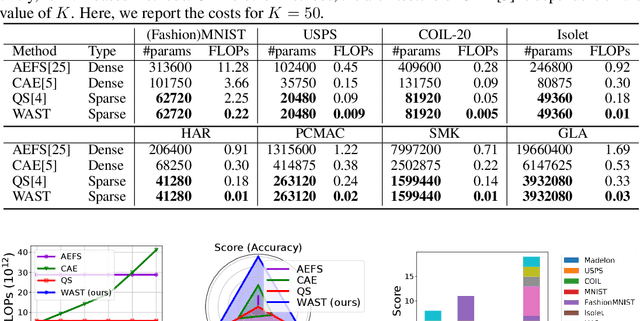
A new line of research for feature selection based on neural networks has recently emerged. Despite its superiority to classical methods, it requires many training iterations to converge and detect informative features. The computational time becomes prohibitively long for datasets with a large number of samples or a very high dimensional feature space. In this paper, we present a new efficient unsupervised method for feature selection based on sparse autoencoders. In particular, we propose a new sparse training algorithm that optimizes a model's sparse topology during training to pay attention to informative features quickly. The attention-based adaptation of the sparse topology enables fast detection of informative features after a few training iterations. We performed extensive experiments on 10 datasets of different types, including image, speech, text, artificial, and biological. They cover a wide range of characteristics, such as low and high-dimensional feature spaces, and few and large training samples. Our proposed approach outperforms the state-of-the-art methods in terms of selecting informative features while reducing training iterations and computational costs substantially. Moreover, the experiments show the robustness of our method in extremely noisy environments.
Sketch2FullStack: Generating Skeleton Code of Full Stack Website and Application from Sketch using Deep Learning and Computer Vision
Nov 26, 2022
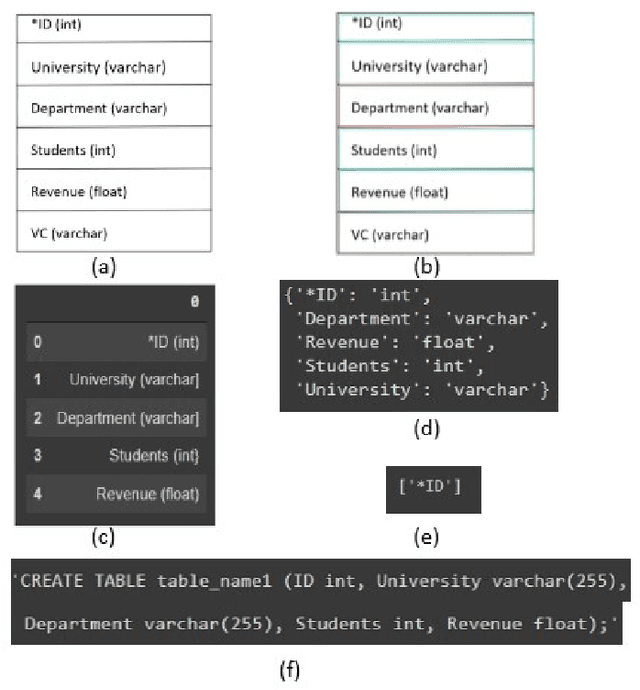

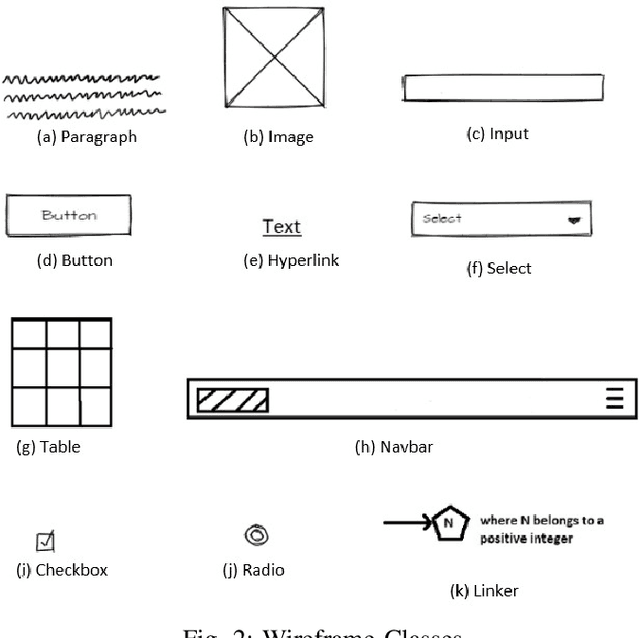
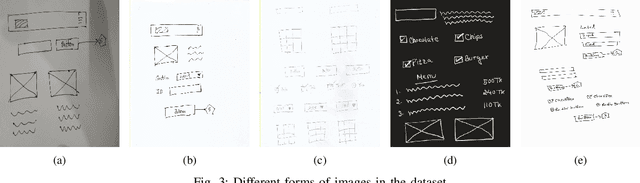
For a full-stack web or app development, it requires a software firm or more specifically a team of experienced developers to contribute a large portion of their time and resources to design the website and then convert it to code. As a result, the efficiency of the development team is significantly reduced when it comes to converting UI wireframes and database schemas into an actual working system. It would save valuable resources and fasten the overall workflow if the clients or developers can automate this process of converting the pre-made full-stack website design to get a partially working if not fully working code. In this paper, we present a novel approach of generating the skeleton code from sketched images using Deep Learning and Computer Vision approaches. The dataset for training are first-hand sketched images of low fidelity wireframes, database schemas and class diagrams. The approach consists of three parts. First, the front-end or UI elements detection and extraction from custom-made UI wireframes. Second, individual database table creation from schema designs and lastly, creating a class file from class diagrams.
High-Fidelity Simulation and Novel Data Analysis of the Bubble Creation and Sound Generation Processes in Breaking Waves
Nov 06, 2022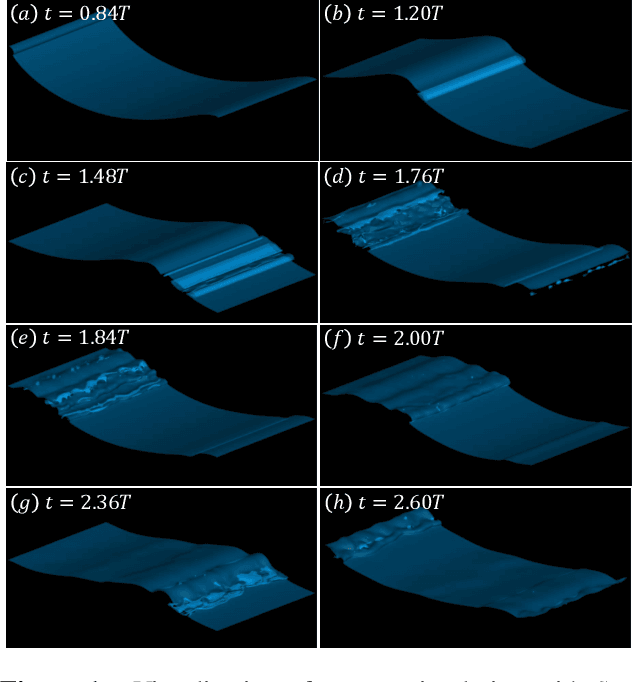
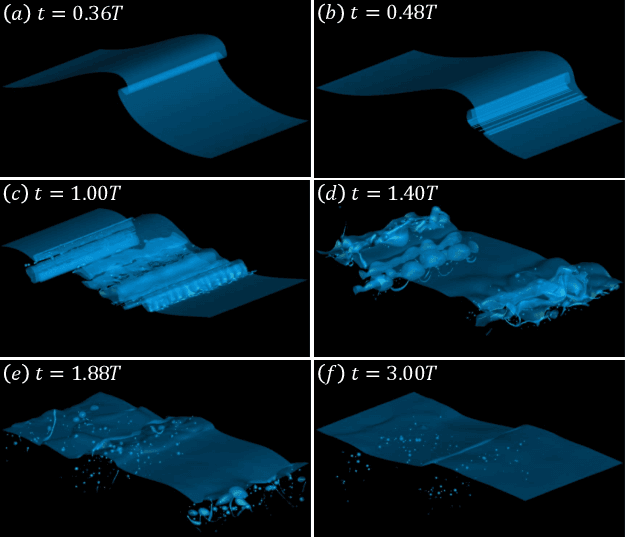
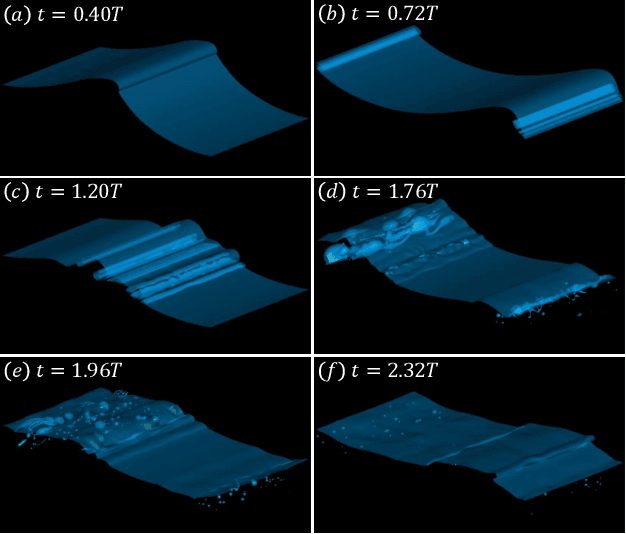

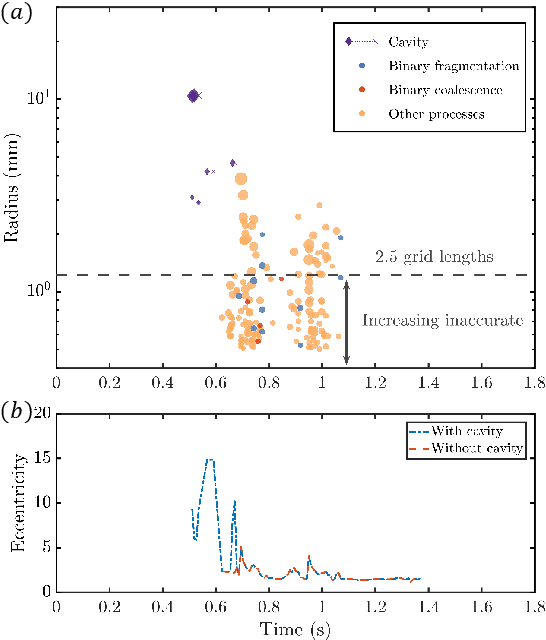
Recent increases in computing power have enabled the numerical simulation of many complex flow problems that are of practical and strategic interest for naval applications. A noticeable area of advancement is the computation of turbulent, two-phase flows resulting from wave breaking and other multiphase flow processes such as cavitation that can generate underwater sound and entrain bubbles in ship wakes, among other effects. Although advanced flow solvers are sophisticated and are capable of simulating high Reynolds number flows on large numbers of grid points, challenges in data analysis remain. Specifically, there is a critical need to transform highly resolved flow fields described on fine grids at discrete time steps into physically resolved features for which the flow dynamics can be understood and utilized in naval applications. This paper presents our recent efforts in this field. In previous works, we developed a novel algorithm to track bubbles in breaking wave simulations and to interpret their dynamical behavior over time (Gao et al., 2021a). We also discovered a new physical mechanism driving bubble production within breaking wave crests (Gao et al., 2021b) and developed a model to relate bubble behaviors to underwater sound generation (Gao et al., 2021c). In this work, we applied our bubble tracking algorithm to the breaking waves simulations and investigated the bubble trajectories, bubble creation mechanisms, and bubble acoustics based on our previous works.
* conference
Fully Automated Deep Learning-enabled Detection for Hepatic Steatosis on Computed Tomography: A Multicenter International Validation Study
Nov 06, 2022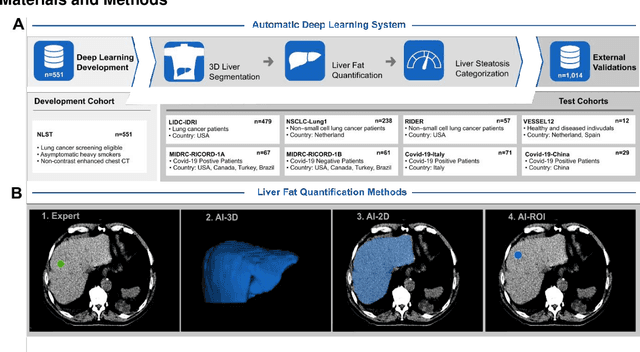
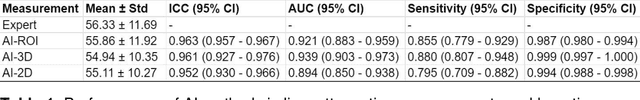
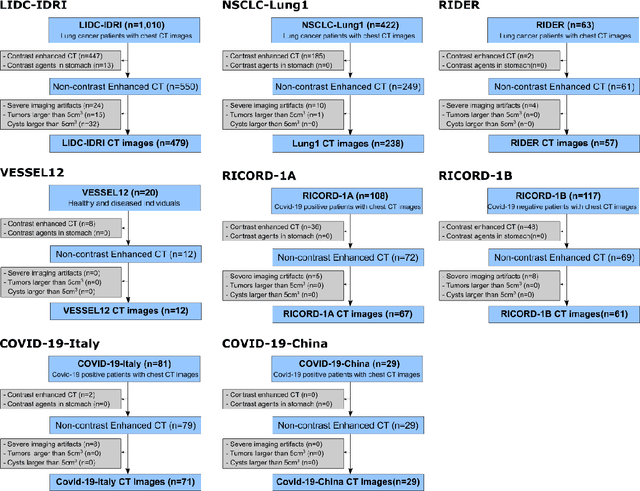
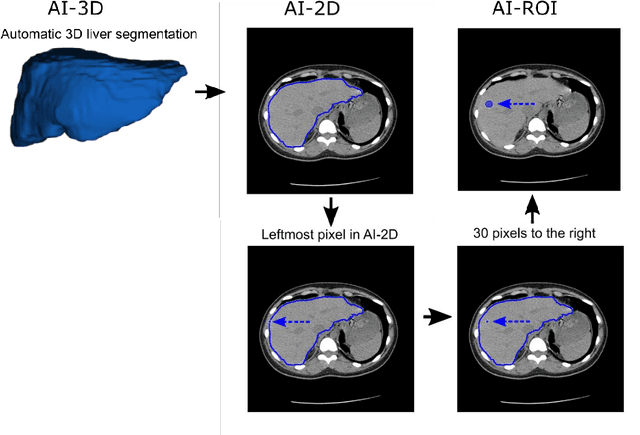
Despite high global prevalence of hepatic steatosis, no automated diagnostics demonstrated generalizability in detecting steatosis on multiple international datasets. Traditionally, hepatic steatosis detection relies on clinicians selecting the region of interest (ROI) on computed tomography (CT) to measure liver attenuation. ROI selection demands time and expertise, and therefore is not routinely performed in populations. To automate the process, we validated an existing artificial intelligence (AI) system for 3D liver segmentation and used it to purpose a novel method: AI-ROI, which could automatically select the ROI for attenuation measurements. AI segmentation and AI-ROI method were evaluated on 1,014 non-contrast enhanced chest CT images from eight international datasets: LIDC-IDRI, NSCLC-Lung1, RIDER, VESSEL12, RICORD-1A, RICORD-1B, COVID-19-Italy, and COVID-19-China. AI segmentation achieved a mean dice coefficient of 0.957. Attenuations measured by AI-ROI showed no significant differences (p = 0.545) and a reduction of 71% time compared to expert measurements. The area under the curve (AUC) of the steatosis classification of AI-ROI is 0.921 (95% CI: 0.883 - 0.959). If performed as a routine screening method, our AI protocol could potentially allow early non-invasive, non-pharmacological preventative interventions for hepatic steatosis. 1,014 expert-annotated liver segmentations of patients with hepatic steatosis annotations can be downloaded here: https://drive.google.com/drive/folders/1-g_zJeAaZXYXGqL1OeF6pUjr6KB0igJX.
Adaptive Robust Model Predictive Control via Uncertainty Cancellation
Dec 02, 2022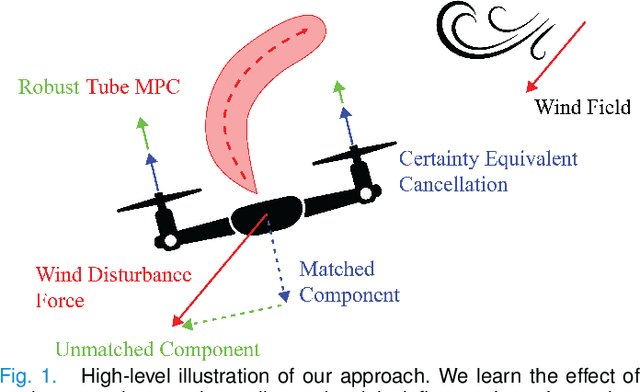
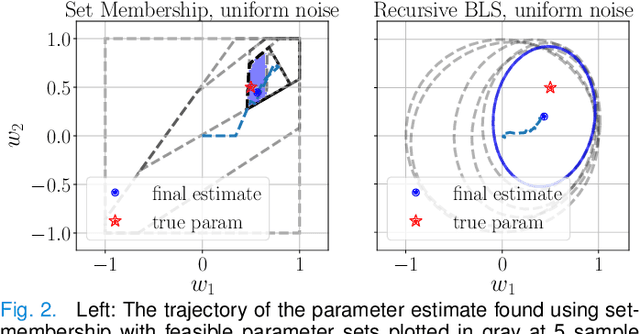
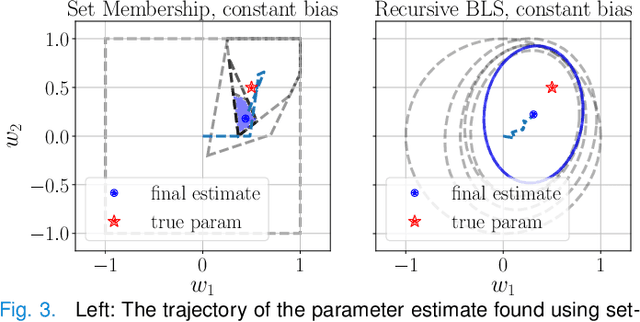
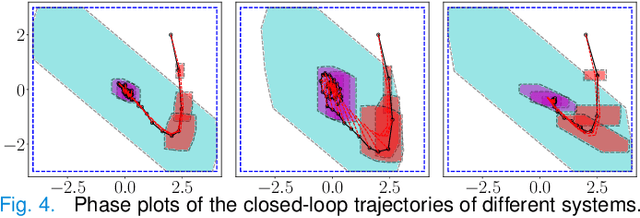
We propose a learning-based robust predictive control algorithm that compensates for significant uncertainty in the dynamics for a class of discrete-time systems that are nominally linear with an additive nonlinear component. Such systems commonly model the nonlinear effects of an unknown environment on a nominal system. We optimize over a class of nonlinear feedback policies inspired by certainty equivalent "estimate-and-cancel" control laws pioneered in classical adaptive control to achieve significant performance improvements in the presence of uncertainties of large magnitude, a setting in which existing learning-based predictive control algorithms often struggle to guarantee safety. In contrast to previous work in robust adaptive MPC, our approach allows us to take advantage of structure (i.e., the numerical predictions) in the a priori unknown dynamics learned online through function approximation. Our approach also extends typical nonlinear adaptive control methods to systems with state and input constraints even when we cannot directly cancel the additive uncertain function from the dynamics. We apply contemporary statistical estimation techniques to certify the system's safety through persistent constraint satisfaction with high probability. Moreover, we propose using Bayesian meta-learning algorithms that learn calibrated model priors to help satisfy the assumptions of the control design in challenging settings. Finally, we show in simulation that our method can accommodate more significant unknown dynamics terms than existing methods and that the use of Bayesian meta-learning allows us to adapt to the test environments more rapidly.
Mixing Up Contrastive Learning: Self-Supervised Representation Learning for Time Series
Mar 17, 2022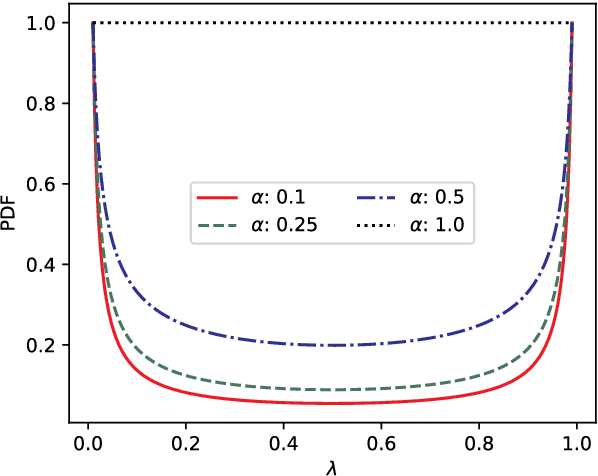
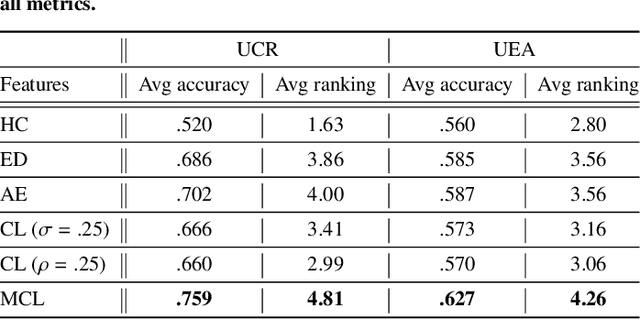
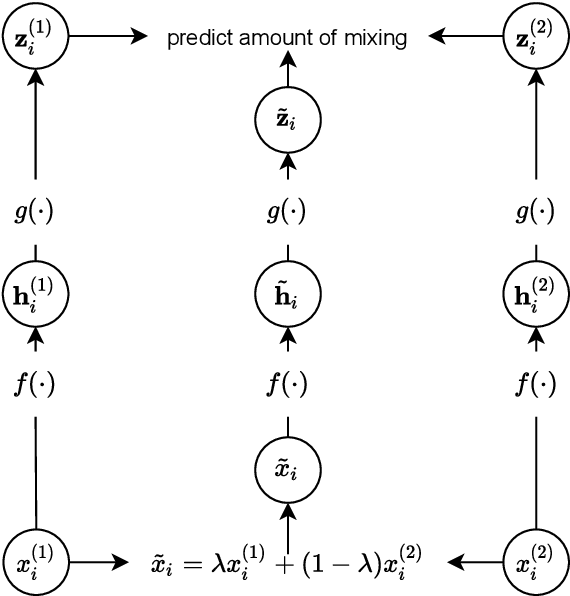
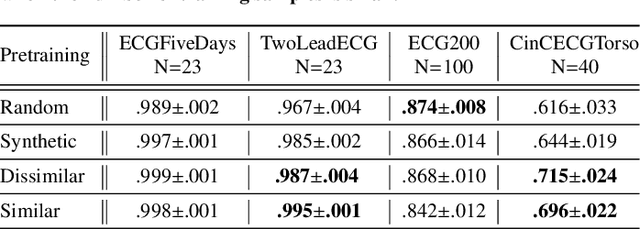
The lack of labeled data is a key challenge for learning useful representation from time series data. However, an unsupervised representation framework that is capable of producing high quality representations could be of great value. It is key to enabling transfer learning, which is especially beneficial for medical applications, where there is an abundance of data but labeling is costly and time consuming. We propose an unsupervised contrastive learning framework that is motivated from the perspective of label smoothing. The proposed approach uses a novel contrastive loss that naturally exploits a data augmentation scheme in which new samples are generated by mixing two data samples with a mixing component. The task in the proposed framework is to predict the mixing component, which is utilized as soft targets in the loss function. Experiments demonstrate the framework's superior performance compared to other representation learning approaches on both univariate and multivariate time series and illustrate its benefits for transfer learning for clinical time series.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge