"Time": models, code, and papers
Optimizing Predictions for Very Small Data Sets: a case study on Open-Source Project Health Prediction
Jan 16, 2023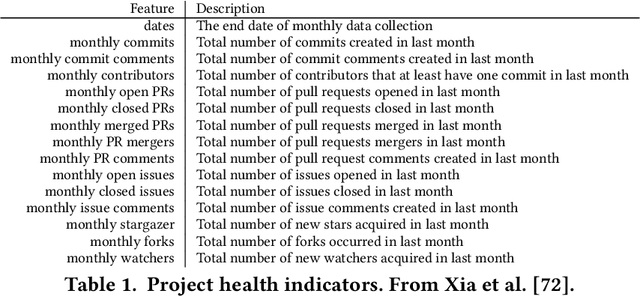
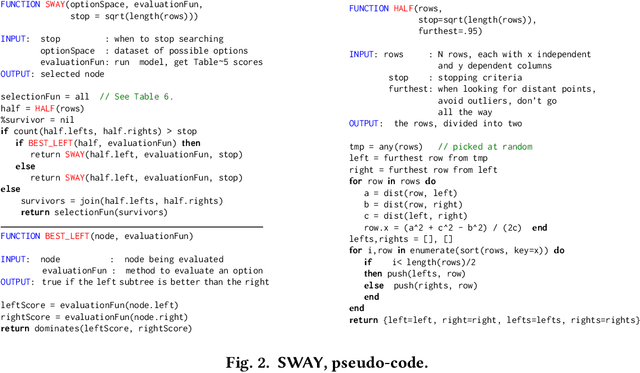



When learning from very small data sets, the resulting models can make many mistakes. For example, consider learning predictors for open source project health. The training data for this task may be very small (e.g. five years of data, collected every month means just 60 rows of training data). Using this data, prior work had unacceptably large errors in their learned predictors. We show that these high errors rates can be tamed by better configuration of the control parameters of the machine learners. For example, we present here a {\em landscape analytics} method (called SNEAK) that (a)~clusters the data to find the general landscape of the hyperparameters; then (b)~explores a few representatives from each part of that landscape. SNEAK is both faster and and more effective than prior state-of-the-art hyperparameter optimization algorithms (FLASH, HYPEROPT, OPTUNA, and differential evolution). More importantly, the configurations found by SNEAK had far less error that other methods. We conjecture that SNEAK works so well since it finds the most informative regions of the hyperparameters, then jumps to those regions. Other methods (that do not reflect over the landscape) can waste time exploring less informative options. From this, we make the following conclusions. Firstly, for predicting open source project health, we recommend landscape analytics (e.g.SNEAK). Secondly, and more generally, when learning from very small data sets, using hyperparameter optimization (e.g. SNEAK) to select learning control parameters. Due to its speed and implementation simplicity, we suggest SNEAK might also be useful in other ``data-light'' SE domains. To assist other researchers in repeating, improving, or even refuting our results, all our scripts and data are available on GitHub at https://github.com/zxcv123456qwe/niSneak
Fully Elman Neural Network: A Novel Deep Recurrent Neural Network Optimized by an Improved Harris Hawks Algorithm for Classification of Pulmonary Arterial Wedge Pressure
Jan 16, 2023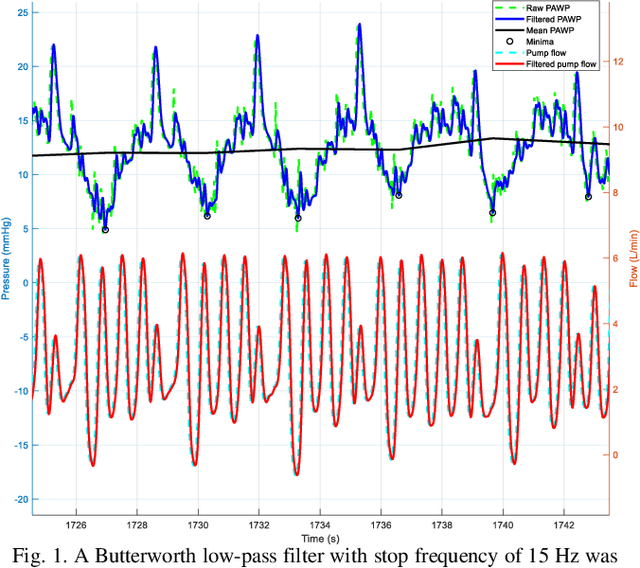
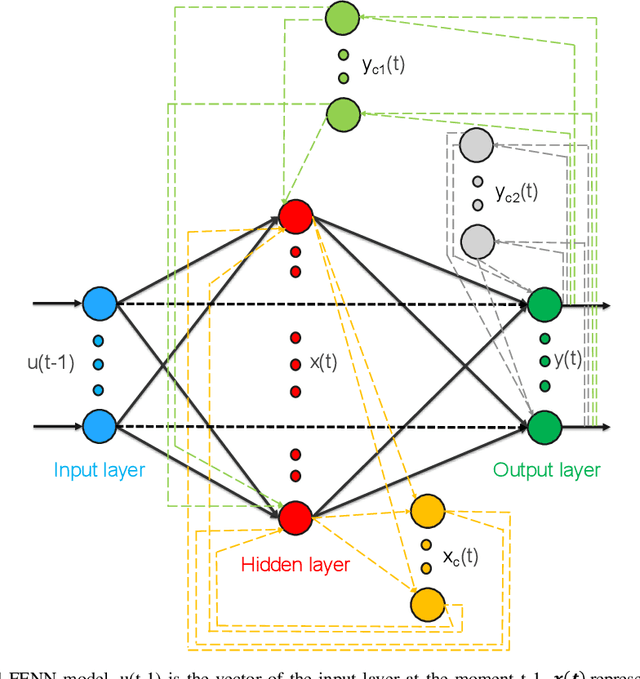
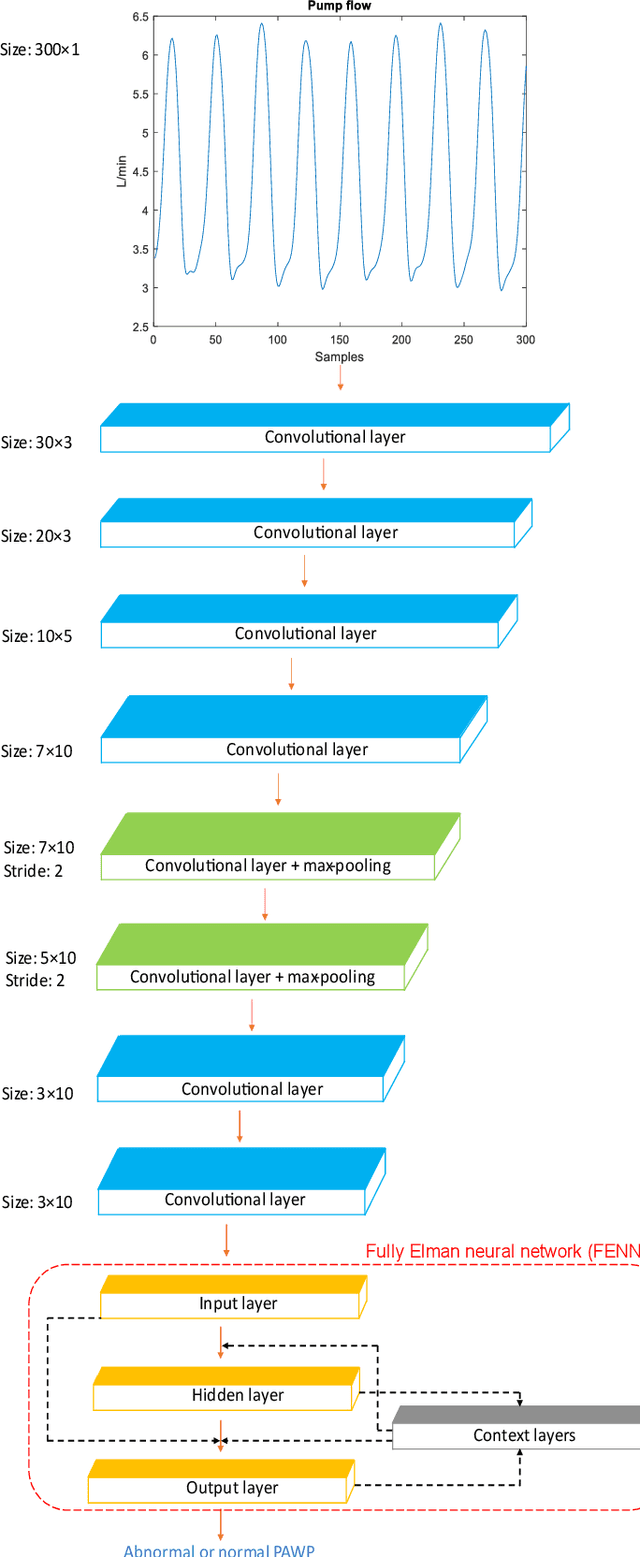
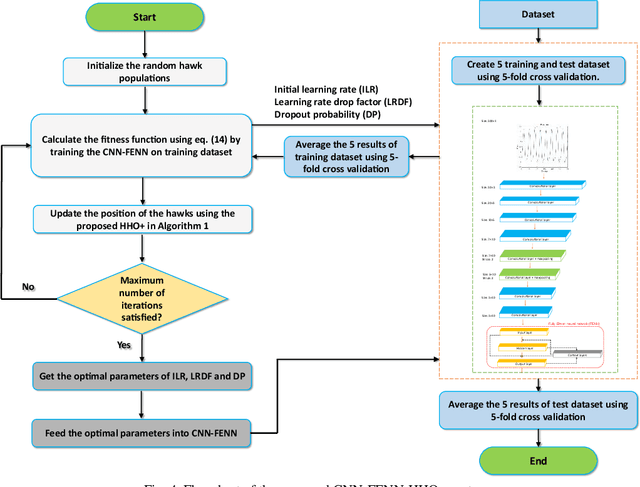
Heart failure (HF) is one of the most prevalent life-threatening cardiovascular diseases in which 6.5 million people are suffering in the USA and more than 23 million worldwide. Mechanical circulatory support of HF patients can be achieved by implanting a left ventricular assist device (LVAD) into HF patients as a bridge to transplant, recovery or destination therapy and can be controlled by measurement of normal and abnormal pulmonary arterial wedge pressure (PAWP). While there are no commercial long-term implantable pressure sensors to measure PAWP, real-time non-invasive estimation of abnormal and normal PAWP becomes vital. In this work, first an improved Harris Hawks optimizer algorithm called HHO+ is presented and tested on 24 unimodal and multimodal benchmark functions. Second, a novel fully Elman neural network (FENN) is proposed to improve the classification performance. Finally, four novel 18-layer deep learning methods of convolutional neural networks (CNNs) with multi-layer perceptron (CNN-MLP), CNN with Elman neural networks (CNN-ENN), CNN with fully Elman neural networks (CNN-FENN), and CNN with fully Elman neural networks optimized by HHO+ algorithm (CNN-FENN-HHO+) for classification of abnormal and normal PAWP using estimated HVAD pump flow were developed and compared. The estimated pump flow was derived by a non-invasive method embedded into the commercial HVAD controller. The proposed methods are evaluated on an imbalanced clinical dataset using 5-fold cross-validation. The proposed CNN-FENN-HHO+ method outperforms the proposed CNN-MLP, CNN-ENN and CNN-FENN methods and improved the classification performance metrics across 5-fold cross-validation. The proposed methods can reduce the likelihood of hazardous events like pulmonary congestion and ventricular suction for HF patients and notify identified abnormal cases to the hospital, clinician and cardiologist.
Data-aware customization of activation functions reduces neural network error
Jan 16, 2023
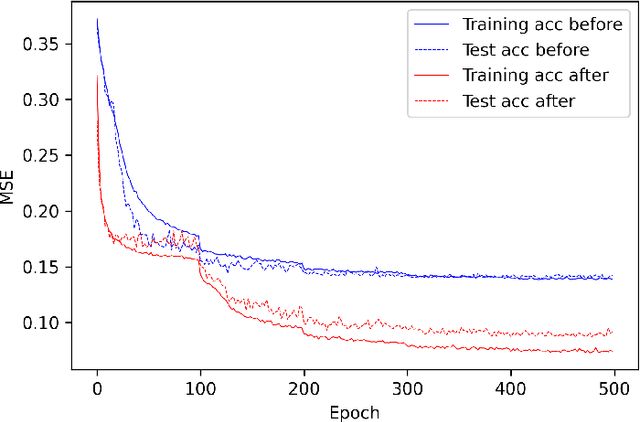
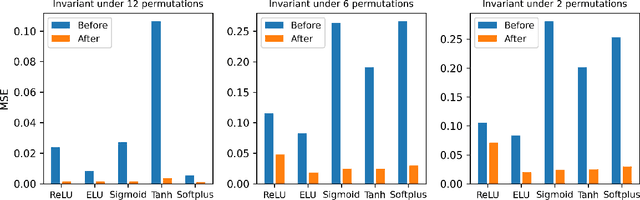
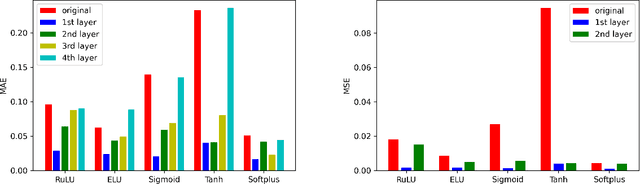
Activation functions play critical roles in neural networks, yet current off-the-shelf neural networks pay little attention to the specific choice of activation functions used. Here we show that data-aware customization of activation functions can result in striking reductions in neural network error. We first give a simple linear algebraic explanation of the role of activation functions in neural networks; then, through connection with the Diaconis-Shahshahani Approximation Theorem, we propose a set of criteria for good activation functions. As a case study, we consider regression tasks with a partially exchangeable target function, \emph{i.e.} $f(u,v,w)=f(v,u,w)$ for $u,v\in \mathbb{R}^d$ and $w\in \mathbb{R}^k$, and prove that for such a target function, using an even activation function in at least one of the layers guarantees that the prediction preserves partial exchangeability for best performance. Since even activation functions are seldom used in practice, we designed the ``seagull'' even activation function $\log(1+x^2)$ according to our criteria. Empirical testing on over two dozen 9-25 dimensional examples with different local smoothness, curvature, and degree of exchangeability revealed that a simple substitution with the ``seagull'' activation function in an already-refined neural network can lead to an order-of-magnitude reduction in error. This improvement was most pronounced when the activation function substitution was applied to the layer in which the exchangeable variables are connected for the first time. While the improvement is greatest for low-dimensional data, experiments on the CIFAR10 image classification dataset showed that use of ``seagull'' can reduce error even for high-dimensional cases. These results collectively highlight the potential of customizing activation functions as a general approach to improve neural network performance.
CAFA: Class-Aware Feature Alignment for Test-Time Adaptation
Jun 01, 2022
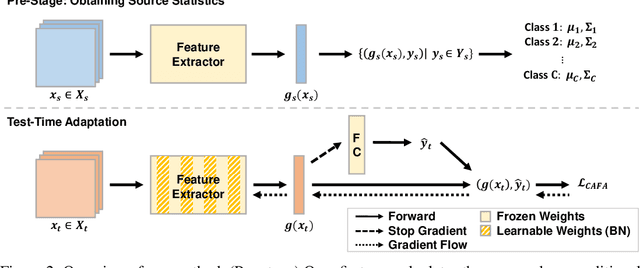
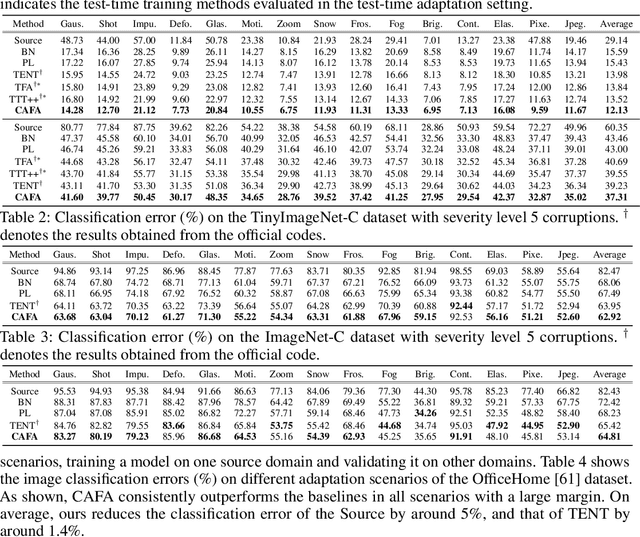
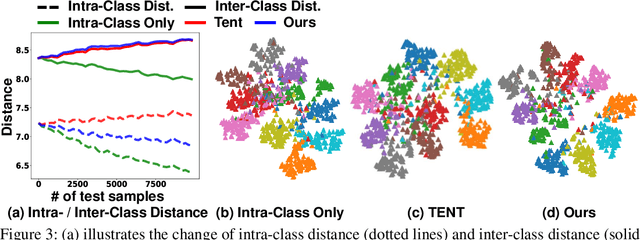
Despite recent advancements in deep learning, deep networks still suffer from performance degradation when they face new and different data from their training distributions. Addressing such a problem, test-time adaptation (TTA) aims to adapt a model to unlabeled test data on test time while making predictions simultaneously. TTA applies to pretrained networks without modifying their training procedures, which enables to utilize the already well-formed source distribution for adaptation. One possible approach is to align the representation space of test samples to the source distribution (\textit{i.e.,} feature alignment). However, performing feature alignments in TTA is especially challenging in that the access to labeled source data is restricted during adaptation. That is, a model does not have a chance to learn test data in a class-discriminative manner, which was feasible in other adaptation tasks (\textit{e.g.,} unsupervised domain adaptation) via supervised loss on the source data. Based on such an observation, this paper proposes \emph{a simple yet effective} feature alignment loss, termed as Class-Aware Feature Alignment (CAFA), which 1) encourages a model to learn target representations in a class-discriminative manner and 2) effectively mitigates the distribution shifts in test time, simultaneously. Our method does not require any hyper-parameters or additional losses, which are required in the previous approaches. We conduct extensive experiments and show our proposed method consistently outperforms existing baselines.
Test Time Transform Prediction for Open Set Histopathological Image Recognition
Jun 20, 2022
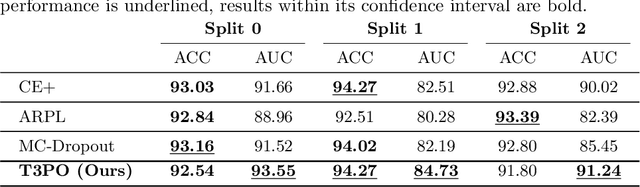
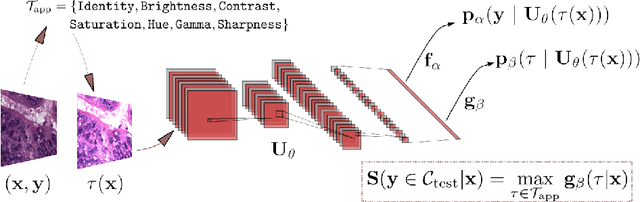
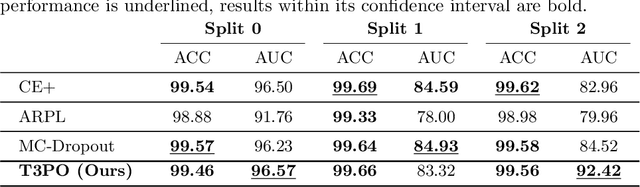
Tissue typology annotation in Whole Slide histological images is a complex and tedious, yet necessary task for the development of computational pathology models. We propose to address this problem by applying Open Set Recognition techniques to the task of jointly classifying tissue that belongs to a set of annotated classes, e.g. clinically relevant tissue categories, while rejecting in test time Open Set samples, i.e. images that belong to categories not present in the training set. To this end, we introduce a new approach for Open Set histopathological image recognition based on training a model to accurately identify image categories and simultaneously predict which data augmentation transform has been applied. In test time, we measure model confidence in predicting this transform, which we expect to be lower for images in the Open Set. We carry out comprehensive experiments in the context of colorectal cancer assessment from histological images, which provide evidence on the strengths of our approach to automatically identify samples from unknown categories. Code is released at https://github.com/agaldran/t3po .
Lessons Learned Applying Deep Learning Approaches to Forecasting Complex Seasonal Behavior
Jan 04, 2023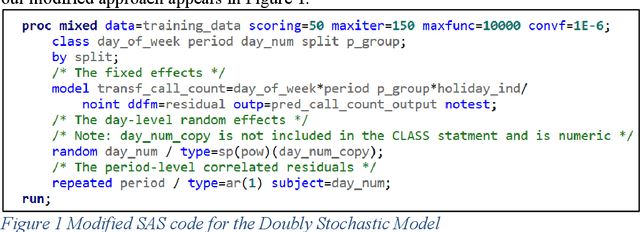

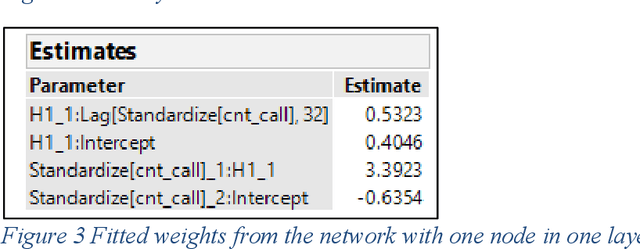
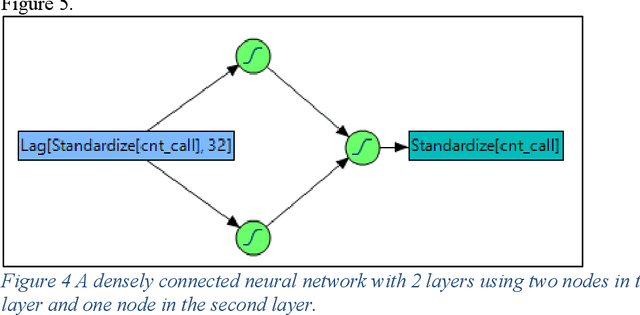
Deep learning methods have gained popularity in recent years through the media and the relative ease of implementation through open source packages such as Keras. We investigate the applicability of popular recurrent neural networks in forecasting call center volumes at a large financial services company. These series are highly complex with seasonal patterns - between hours of the day, day of the week, and time of the year - in addition to autocorrelation between individual observations. Though we investigate the financial services industry, the recommendations for modeling cyclical nonlinear behavior generalize across all sectors. We explore the optimization of parameter settings and convergence criteria for Elman (simple), Long Short-Term Memory (LTSM), and Gated Recurrent Unit (GRU) RNNs from a practical point of view. A designed experiment using actual call center data across many different "skills" (income call streams) compares performance measured by validation error rates of the best observed RNN configurations against other modern and classical forecasting techniques. We summarize the utility of and considerations required for using deep learning methods in forecasting.
* Published in 2019 Joint Statistical Meetings (JSM) proceedings
Semantic rule Web-based Diagnosis and Treatment of Vector-Borne Diseases using SWRL rules
Jan 08, 2023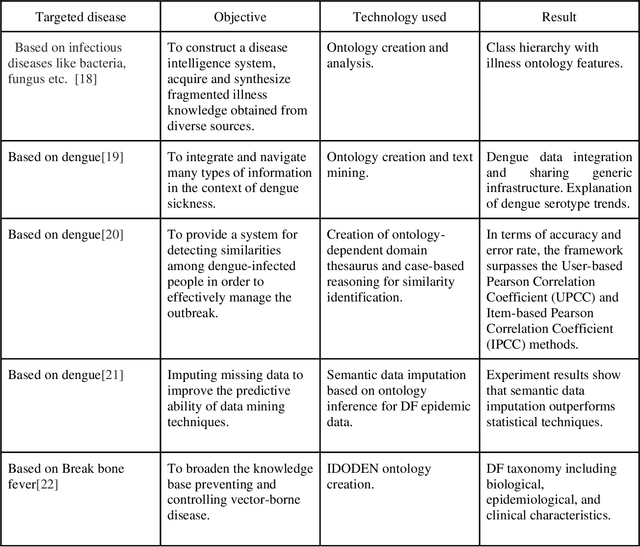
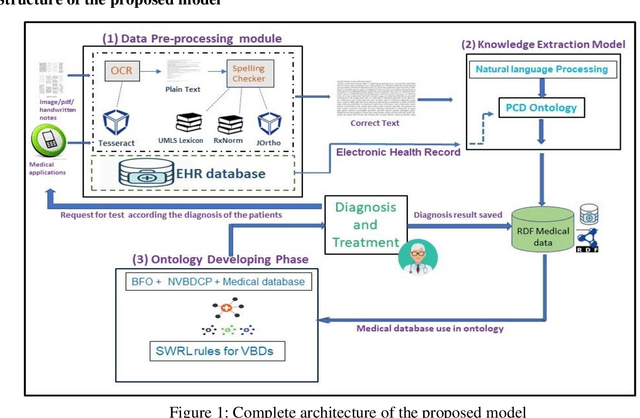
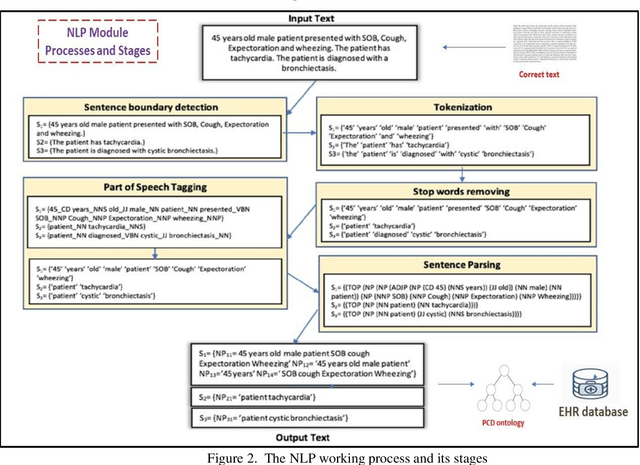

Vector-borne diseases (VBDs) are a kind of infection caused through the transmission of vectors generated by the bites of infected parasites, bacteria, and viruses, such as ticks, mosquitoes, triatomine bugs, blackflies, and sandflies. If these diseases are not properly treated within a reasonable time frame, the mortality rate may rise. In this work, we propose a set of ontologies that will help in the diagnosis and treatment of vector-borne diseases. For developing VBD's ontology, electronic health records taken from the Indian Health Records website, text data generated from Indian government medical mobile applications, and doctors' prescribed handwritten notes of patients are used as input. This data is then converted into correct text using Optical Character Recognition (OCR) and a spelling checker after pre-processing. Natural Language Processing (NLP) is applied for entity extraction from text data for making Resource Description Framework (RDF) medical data with the help of the Patient Clinical Data (PCD) ontology. Afterwards, Basic Formal Ontology (BFO), National Vector Borne Disease Control Program (NVBDCP) guidelines, and RDF medical data are used to develop ontologies for VBDs, and Semantic Web Rule Language (SWRL) rules are applied for diagnosis and treatment. The developed ontology helps in the construction of decision support systems (DSS) for the NVBDCP to control these diseases.
Deep Injective Prior for Inverse Scattering
Jan 08, 2023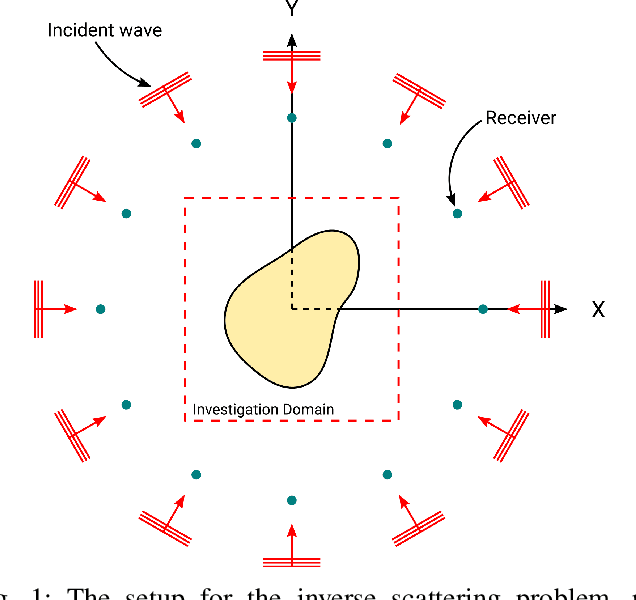
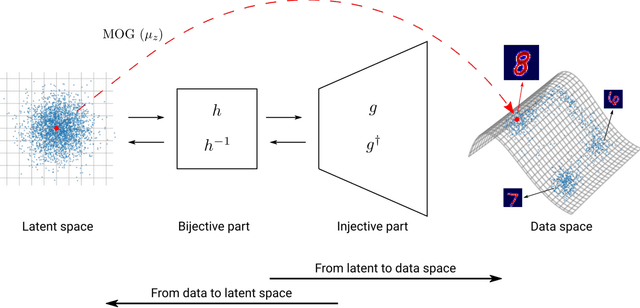
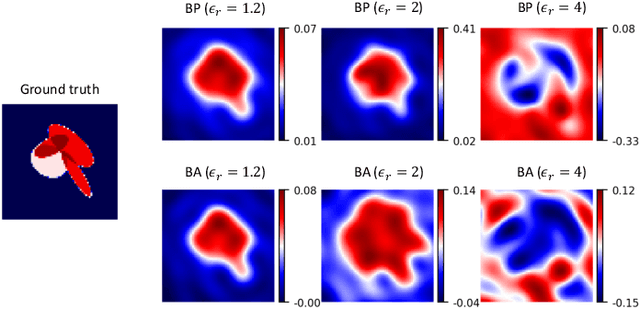
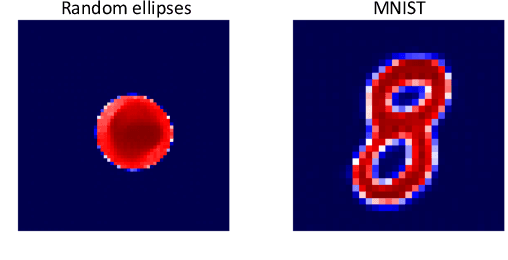
In electromagnetic inverse scattering, we aim to reconstruct object permittivity from scattered waves. Deep learning is a promising alternative to traditional iterative solvers, but it has been used mostly in a supervised framework to regress the permittivity patterns from scattered fields or back-projections. While such methods are fast at test-time and achieve good results for specific data distributions, they are sensitive to the distribution drift of the scattered fields, common in practice. If the distribution of the scattered fields changes due to changes in frequency, the number of transmitters and receivers, or any other real-world factor, an end-to-end neural network must be re-trained or fine-tuned on a new dataset. In this paper, we propose a new data-driven framework for inverse scattering based on deep generative models. We model the target permittivities by a low-dimensional manifold which acts as a regularizer and learned from data. Unlike supervised methods which require both scattered fields and target signals, we only need the target permittivities for training; it can then be used with any experimental setup. We show that the proposed framework significantly outperforms the traditional iterative methods especially for strong scatterers while having comparable reconstruction quality to state-of-the-art deep learning methods like U-Net.
Stochastic Langevin Monte Carlo for (weakly) log-concave posterior distributions
Jan 08, 2023In this paper, we investigate a continuous time version of the Stochastic Langevin Monte Carlo method, introduced in [WT11], that incorporates a stochastic sampling step inside the traditional over-damped Langevin diffusion. This method is popular in machine learning for sampling posterior distribution. We will pay specific attention in our work to the computational cost in terms of $n$ (the number of observations that produces the posterior distribution), and $d$ (the dimension of the ambient space where the parameter of interest is living). We derive our analysis in the weakly convex framework, which is parameterized with the help of the Kurdyka-\L ojasiewicz (KL) inequality, that permits to handle a vanishing curvature settings, which is far less restrictive when compared to the simple strongly convex case. We establish that the final horizon of simulation to obtain an $\varepsilon$ approximation (in terms of entropy) is of the order $( d \log(n)^2 )^{(1+r)^2} [\log^2(\varepsilon^{-1}) + n^2 d^{2(1+r)} \log^{4(1+r)}(n) ]$ with a Poissonian subsampling of parameter $\left(n ( d \log^2(n))^{1+r}\right)^{-1}$, where the parameter $r$ is involved in the KL inequality and varies between $0$ (strongly convex case) and $1$ (limiting Laplace situation).
Modeling Continuous Time Sequences with Intermittent Observations using Marked Temporal Point Processes
Jun 23, 2022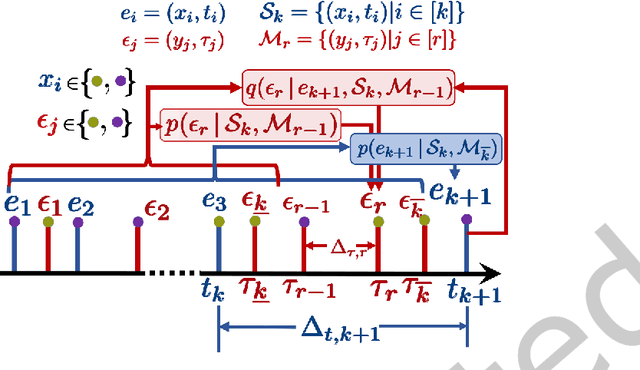
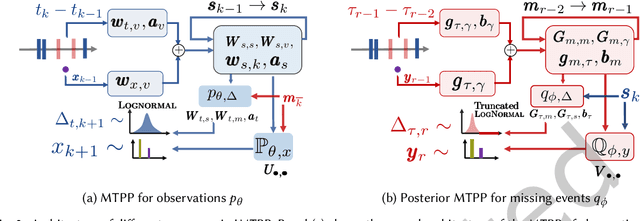
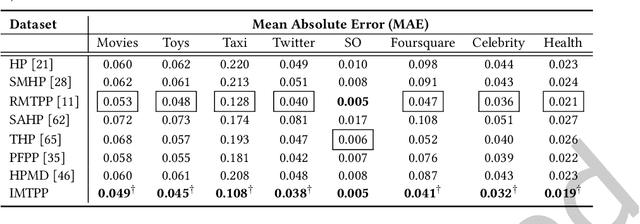
A large fraction of data generated via human activities such as online purchases, health records, spatial mobility etc. can be represented as a sequence of events over a continuous-time. Learning deep learning models over these continuous-time event sequences is a non-trivial task as it involves modeling the ever-increasing event timestamps, inter-event time gaps, event types, and the influences between different events within and across different sequences. In recent years neural enhancements to marked temporal point processes (MTPP) have emerged as a powerful framework to model the underlying generative mechanism of asynchronous events localized in continuous time. However, most existing models and inference methods in the MTPP framework consider only the complete observation scenario i.e. the event sequence being modeled is completely observed with no missing events -- an ideal setting that is rarely applicable in real-world applications. A recent line of work which considers missing events while training MTPP utilizes supervised learning techniques that require additional knowledge of missing or observed label for each event in a sequence, which further restricts its practicability as in several scenarios the details of missing events is not known apriori. In this work, we provide a novel unsupervised model and inference method for learning MTPP in presence of event sequences with missing events. Specifically, we first model the generative processes of observed events and missing events using two MTPP, where the missing events are represented as latent random variables. Then, we devise an unsupervised training method that jointly learns both the MTPP by means of variational inference. Such a formulation can effectively impute the missing data among the observed events and can identify the optimal position of missing events in a sequence.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge